

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148347.8 - phase: 0 /pseudo
(97 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
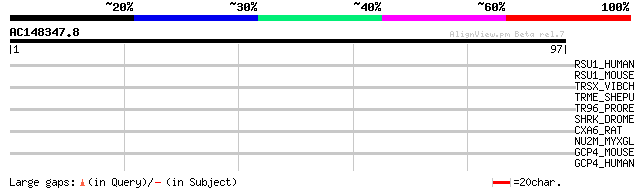
Score E
Sequences producing significant alignments: (bits) Value
RSU1_HUMAN (Q15404) Ras suppressor protein 1 (Rsu-1) (RSP-1) 30 0.80
RSU1_MOUSE (Q01730) Ras suppressor protein 1 (Rsu-1) (RSP-1) 28 3.1
TRSX_VIBCH (Q79S39) HTH-type transcriptional regulator for conju... 27 6.8
TRME_SHEPU (Q8GJK1) HTH-type transcriptional regulator for conju... 27 6.8
TR96_PRORE (Q79RI9) HTH-type transcriptional regulator for conju... 27 6.8
SHRK_DROME (Q24145) Tyrosine-protein kinase shark (EC 2.7.1.112) 27 6.8
CXA6_RAT (P28233) Gap junction alpha-6 protein (Connexin 33) (Cx33) 27 6.8
NU2M_MYXGL (O21078) NADH-ubiquinone oxidoreductase chain 2 (EC 1... 27 8.9
GCP4_MOUSE (Q9D4F8) Gamma-tubulin complex component 4 (GCP-4) 27 8.9
GCP4_HUMAN (Q9UGJ1) Gamma-tubulin complex component 4 (GCP-4) (h... 27 8.9
>RSU1_HUMAN (Q15404) Ras suppressor protein 1 (Rsu-1) (RSP-1)
Length = 276
Score = 30.4 bits (67), Expect = 0.80
Identities = 22/77 (28%), Positives = 42/77 (53%), Gaps = 5/77 (6%)
Query: 1 MQEQWVKDVMRVSGLQWLQHVSYLVIDQGLLSV----IAERWH-EVTSSFHLPIWEMIVT 55
M ++ + +++ V+GL L H++ LV+ L++ IAE + EV + F+ I E+
Sbjct: 21 MSDRGISNMLDVNGLFTLSHITQLVLSHNKLTMVPPNIAELKNLEVLNFFNNQIEELPTQ 80
Query: 56 LDDLSCLLHLSIESHML 72
+ L L HL++ + L
Sbjct: 81 ISSLQKLKHLNLGMNRL 97
>RSU1_MOUSE (Q01730) Ras suppressor protein 1 (Rsu-1) (RSP-1)
Length = 276
Score = 28.5 bits (62), Expect = 3.1
Identities = 21/77 (27%), Positives = 40/77 (51%), Gaps = 5/77 (6%)
Query: 1 MQEQWVKDVMRVSGLQWLQHVSYLVIDQGLLSV----IAERWH-EVTSSFHLPIWEMIVT 55
M ++ + ++ V+GL L H++ LV+ L+ +AE + EV + F+ I E+
Sbjct: 21 MSDRGISSMLDVNGLFSLAHITQLVLSHNKLTTVPPNVAELKNLEVLNFFNNQIEELPTQ 80
Query: 56 LDDLSCLLHLSIESHML 72
+ L L HL++ + L
Sbjct: 81 ISSLQKLKHLNLGMNRL 97
>TRSX_VIBCH (Q79S39) HTH-type transcriptional regulator for
conjugative element SXT
Length = 215
Score = 27.3 bits (59), Expect = 6.8
Identities = 12/36 (33%), Positives = 17/36 (46%)
Query: 11 RVSGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFH 46
R+ + W+Q + I +G AE W EVT H
Sbjct: 84 RIPLINWVQAGDWTEIAEGFAHEDAEEWREVTGKAH 119
>TRME_SHEPU (Q8GJK1) HTH-type transcriptional regulator for
conjugative element pMERPH
Length = 215
Score = 27.3 bits (59), Expect = 6.8
Identities = 12/36 (33%), Positives = 17/36 (46%)
Query: 11 RVSGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFH 46
R+ + W+Q + I +G AE W EVT H
Sbjct: 84 RIPLINWVQAGDWTEIAEGFAHEDAEEWREVTGKAH 119
>TR96_PRORE (Q79RI9) HTH-type transcriptional regulator for
conjugative element R391 (ORF-96 protein)
Length = 215
Score = 27.3 bits (59), Expect = 6.8
Identities = 12/36 (33%), Positives = 17/36 (46%)
Query: 11 RVSGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFH 46
R+ + W+Q + I +G AE W EVT H
Sbjct: 84 RIPLINWVQAGDWTEIAEGFAHEDAEEWREVTGKAH 119
>SHRK_DROME (Q24145) Tyrosine-protein kinase shark (EC 2.7.1.112)
Length = 939
Score = 27.3 bits (59), Expect = 6.8
Identities = 16/73 (21%), Positives = 34/73 (45%), Gaps = 8/73 (10%)
Query: 7 KDVMRVSGL---QWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLL 63
K ++R+ G+ + L V L +L I + HE+T++ L +W ++C +
Sbjct: 723 KCIVRLIGIAKGEMLMMVQELAPLGSMLQYILDHGHEITANAELKVW-----ASQIACGM 777
Query: 64 HLSIESHMLYHNI 76
H H ++ ++
Sbjct: 778 HYLESQHFVHRDL 790
>CXA6_RAT (P28233) Gap junction alpha-6 protein (Connexin 33) (Cx33)
Length = 286
Score = 27.3 bits (59), Expect = 6.8
Identities = 15/52 (28%), Positives = 25/52 (47%)
Query: 46 HLPIWEMIVTLDDLSCLLHLSIESHMLYHNIYLTRSEAVDLMVELLGSDSGE 97
H+ +W + V + LLHL+ +++ N L + E +L V SGE
Sbjct: 74 HVRLWVLQVIFVSVPTLLHLAHVYYVIRQNEKLKKQEEEELKVAHFNGASGE 125
>NU2M_MYXGL (O21078) NADH-ubiquinone oxidoreductase chain 2 (EC
1.6.5.3)
Length = 348
Score = 26.9 bits (58), Expect = 8.9
Identities = 12/33 (36%), Positives = 20/33 (60%), Gaps = 1/33 (3%)
Query: 5 WVKDVMRVSG-LQWLQHVSYLVIDQGLLSVIAE 36
W+ VM VS L W+ V YL+I+ + +++ E
Sbjct: 187 WIVGVMAVSASLSWVTTVIYLIINFSIFTILIE 219
>GCP4_MOUSE (Q9D4F8) Gamma-tubulin complex component 4 (GCP-4)
Length = 667
Score = 26.9 bits (58), Expect = 8.9
Identities = 15/46 (32%), Positives = 24/46 (51%)
Query: 39 HEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLYHNIYLTRSEAV 84
H+V +F ++++ D+L LLHL+IE H H T+ V
Sbjct: 390 HDVNVAFQQSAHKVLLDDDNLLPLLHLTIEYHGKDHKADATQPREV 435
>GCP4_HUMAN (Q9UGJ1) Gamma-tubulin complex component 4 (GCP-4)
(hGCP4) (h76p) (Hgrip76)
Length = 667
Score = 26.9 bits (58), Expect = 8.9
Identities = 14/43 (32%), Positives = 24/43 (55%)
Query: 39 HEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLYHNIYLTRS 81
H+V +F ++++ D+L LLHL+IE H H T++
Sbjct: 390 HDVNVAFQQSAHKVLLDDDNLLPLLHLTIEYHGKEHKADATQA 432
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,308,510
Number of Sequences: 164201
Number of extensions: 378520
Number of successful extensions: 768
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 762
Number of HSP's gapped (non-prelim): 10
length of query: 97
length of database: 59,974,054
effective HSP length: 73
effective length of query: 24
effective length of database: 47,987,381
effective search space: 1151697144
effective search space used: 1151697144
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148347.8


