

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148345.17 + phase: 0
(528 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
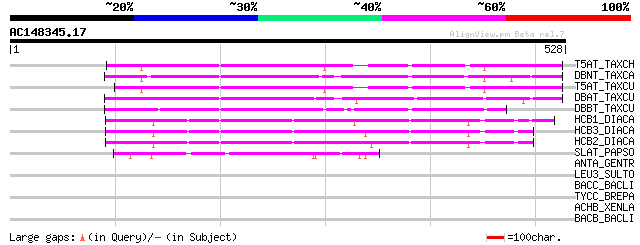
Sequences producing significant alignments: (bits) Value
T5AT_TAXCH (Q8S9G6) Taxadien-5-alpha-ol O-acetyltransferase (EC ... 213 8e-55
DBNT_TAXCA (Q8LL69) 3'-N-debenzoyl-2'-deoxytaxol N-benzoyltransf... 212 2e-54
T5AT_TAXCU (Q9M6F0) Taxadien-5-alpha-ol O-acetyltransferase (EC ... 209 1e-53
DBAT_TAXCU (Q9M6E2) 10-deacetylbaccatin III 10-O-acetyltransfera... 199 2e-50
DBBT_TAXCU (Q9FPW3) 2-alpha-hydroxytaxane 2-O-benzoyltransferase... 177 8e-44
HCB1_DIACA (O24645) Anthranilate N-benzoyltransferase protein 1 ... 146 1e-34
HCB3_DIACA (O23918) Anthranilate N-benzoyltransferase protein 3 ... 144 7e-34
HCB2_DIACA (O23917) Anthranilate N-benzoyltransferase protein 2 ... 142 2e-33
SLAT_PAPSO (Q94FT4) Salutaridinol 7-O-acetyltransferase (EC 2.3.... 76 2e-13
ANTA_GENTR (Q9ZWR8) Anthocyanin 5-aromatic acyltransferase (EC 2... 40 0.011
LEU3_SULTO (P50455) 3-isopropylmalate dehydrogenase (EC 1.1.1.85... 33 2.4
BACC_BACLI (O68008) Bacitracin synthetase 3 (BA3) [Includes: ATP... 32 3.1
TYCC_BREPA (O30409) Tyrocidine synthetase III [Includes: ATP-dep... 32 4.1
ACHB_XENLA (P49579) Acetylcholine receptor protein, beta chain p... 32 4.1
BACB_BACLI (O68007) Bacitracin synthetase 2 (BA2) [Includes: ATP... 32 5.3
>T5AT_TAXCH (Q8S9G6) Taxadien-5-alpha-ol O-acetyltransferase (EC
2.3.1.162)
(Taxa-4(20),11(12)-dien-5alpha-ol-O-acetyltransferase)
(Taxadienol acetyltransferase)
Length = 439
Score = 213 bits (543), Expect = 8e-55
Identities = 139/448 (31%), Positives = 221/448 (49%), Gaps = 35/448 (7%)
Query: 93 VRRSQPELVPPAAPTPREVKLLSDIDDQEGLR---FNIPMMFIYRHEPSMKEKDPVKVLR 149
V ++ +V P+ P P+ LS ID+ G+R FN +++ P+M DP K++R
Sbjct: 8 VNLNEKVMVGPSLPLPKTTLQLSSIDNLPGVRGSIFNALLIYNASPSPTMVSADPAKLIR 67
Query: 150 HALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFE 209
AL++ LVYY PFAGR+RE L V+CTGEG MF+EA AD L GD P F+
Sbjct: 68 EALAKILVYYPPFAGRLRETENGDLEVECTGEGAMFLEAMADNELSVLGD-FDDSNPSFQ 126
Query: 210 ELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGA 269
+LL+ +P D P+ ++QVTR CG F++ V +H + DG G F+ AEMARG
Sbjct: 127 QLLFSLPLDTNFKDLPLLVVQVTRFTCGGFVVGVSFHHGVCDGRGAAQFLKGLAEMARGE 186
Query: 270 HKPSIQPVWNREILMARDPPHITCNHHEY-------EQIFSSNTIKEEDTTTLVHQSFFF 322
K S++P+WNRE++ DP ++ H E+ E+I + I + +T + QS
Sbjct: 187 VKLSLEPIWNRELVKLDDPKYLQFFHFEFLRAPSIVEKIVQTYFIIDFETINYIKQSVME 246
Query: 323 RTSDIVVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNAN 382
+ C++F++ ++ W RT+A Q+ + ++++ ++ R+ F
Sbjct: 247 ECKEF-------------CSSFEVASAMTWIARTRAFQIPESEYVKILFGMDMRNSF--- 290
Query: 383 NSPL-VGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKE 441
N PL GYYGN AV L SL A+ +++K K + + + V+K
Sbjct: 291 NPPLPSGYYGNSIGTACAVDNVQDLLSGSLLRAIMIIKKSKVSLNDNFKSRA----VVKP 346
Query: 442 RCLFTTIRS---CVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPGAAYIIPHKNVEG 498
L + +D +R+ EVDFGWG AV + ++ +
Sbjct: 347 SELDVNMNHENVVAFADWSRLGFDEVDFGWGNAVSVSPVQQQCELAMQNYFLFLKPSKNK 406
Query: 499 EEGFILLVCLSFEAMKRFAKELDEMLGK 526
+G +L+ L MK F E++ M+ K
Sbjct: 407 PDGIKILMFLPLSKMKSFKIEMEAMMKK 434
>DBNT_TAXCA (Q8LL69) 3'-N-debenzoyl-2'-deoxytaxol
N-benzoyltransferase (EC 2.3.1.-) (DBTNBT)
Length = 441
Score = 212 bits (539), Expect = 2e-54
Identities = 145/446 (32%), Positives = 229/446 (50%), Gaps = 28/446 (6%)
Query: 91 FTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLR--FNIPMMFIYRHEPSMKEKDPVKVL 148
F V++ P +V P+ P+P+ LS +D R FN ++F + P DPVK++
Sbjct: 9 FHVKKFDPVMVAPSLPSPKATVQLSVVDSLTICRGIFNTLLVF---NAPDNISADPVKII 65
Query: 149 RHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCF 208
R ALS+ LVYY+P AGR+R +L V+CTG+G +FVEA + T+ D L P F
Sbjct: 66 REALSKVLVYYFPLAGRLRSKEIGELEVECTGDGALFVEAMVEDTISVLRD-LDDLNPSF 124
Query: 209 EELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARG 268
++L++ P I D + ++QVTR CG + V L H++ DG G F+ A AEMARG
Sbjct: 125 QQLVFWHPLDTAIEDLHLVIVQVTRFTCGGIAVGVTLPHSVCDGRGAAQFVTALAEMARG 184
Query: 269 AHKPSIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFFFRTSDIV 328
KPS++P+WNRE+L DP H+ N +++ I ++E L SF I
Sbjct: 185 EVKPSLEPIWNRELLNPEDPLHLQLN--QFDSICPPPMLEE-----LGQASFVINVDTIE 237
Query: 329 VLRLLVPFHLRH-CTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPLV 387
++ V C++F+++ + W RTKALQ+ + ++++ ++ R F N PL
Sbjct: 238 YMKQCVMEECNEFCSSFEVVAALVWIARTKALQIPHTENVKLLFAMDLRKLF---NPPLP 294
Query: 388 -GYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCLFT 446
GYYGN A+ L SL A+ +++K KA++ + Y S +V L
Sbjct: 295 NGYYGNAIGTAYAMDNVQDLLNGSLLRAIMIIKKAKADLKDNYSRS---RVVTNPYSLDV 351
Query: 447 TIRS---CVVSDVTRIKLREVDFGWGEAVYGGA---AKSGAGPFPGAAYIIPHKNVEGEE 500
+S +SD R+ E DFGWG + + ++G F Y++P KN + +
Sbjct: 352 NKKSDNILALSDWRRLGFYEADFGWGGPLNVSSLQRLENGLPMFSTFLYLLPAKN-KSDG 410
Query: 501 GFILLVCLSFEAMKRFAKELDEMLGK 526
+LL C+ +K F ++ M+ K
Sbjct: 411 IKLLLSCMPPTTLKSFKIVMEAMIEK 436
>T5AT_TAXCU (Q9M6F0) Taxadien-5-alpha-ol O-acetyltransferase (EC
2.3.1.162)
(Taxa-4(20),11(12)-dien-5alpha-ol-O-acetyltransferase)
(Taxadienol acetyltransferase)
Length = 439
Score = 209 bits (532), Expect = 1e-53
Identities = 137/441 (31%), Positives = 217/441 (49%), Gaps = 35/441 (7%)
Query: 100 LVPPAAPTPREVKLLSDIDDQEGLR---FNIPMMFIYRHEPSMKEKDPVKVLRHALSQAL 156
+V P+ P P+ LS ID+ G+R FN +++ P+M DP K +R AL++ L
Sbjct: 15 MVGPSPPLPKTTLQLSSIDNLPGVRGSIFNALLIYNASPSPTMISADPAKPIREALAKIL 74
Query: 157 VYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFEELLYDVP 216
VYY PFAGR+RE L V+CTGEG MF+EA AD L GD P F++LL+ +P
Sbjct: 75 VYYPPFAGRLRETENGDLEVECTGEGAMFLEAMADNELSVLGD-FDDSNPSFQQLLFSLP 133
Query: 217 GSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGAHKPSIQP 276
D + ++QVTR CG F++ V +H + DG G F+ AEMARG K S++P
Sbjct: 134 LDTNFKDLSLLVVQVTRFTCGGFVVGVSFHHGVCDGRGAAQFLKGLAEMARGEVKLSLEP 193
Query: 277 VWNREILMARDPPHITCNHHEY-------EQIFSSNTIKEEDTTTLVHQSFFFRTSDIVV 329
+WNRE++ DP ++ H E+ E+I + I + +T + QS +
Sbjct: 194 IWNRELVKLDDPKYLQFFHFEFLRAPSIVEKIVQTYFIIDFETINYIKQSVMEECKEF-- 251
Query: 330 LRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPL-VG 388
C++F++ ++ W RT+A Q+ + ++++ ++ R+ F N PL G
Sbjct: 252 -----------CSSFEVASAMTWIARTRAFQIPESEYVKILFGMDMRNSF---NPPLPSG 297
Query: 389 YYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCLFTTI 448
YYGN AV L SL A+ +++K K + + + V+K L +
Sbjct: 298 YYGNSIGTACAVDNVQDLLSGSLLRAIMIIKKSKVSLNDNFKSRA----VVKPSELDVNM 353
Query: 449 RS---CVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPGAAYIIPHKNVEGEEGFILL 505
+D +R+ EVDFGWG AV + + ++ + +G +L
Sbjct: 354 NHENVVAFADWSRLGFDEVDFGWGNAVSVSPVQQQSALAMQNYFLFLKPSKNKPDGIKIL 413
Query: 506 VCLSFEAMKRFAKELDEMLGK 526
+ L MK F E++ M+ K
Sbjct: 414 MFLPLSKMKSFKIEMEAMMKK 434
>DBAT_TAXCU (Q9M6E2) 10-deacetylbaccatin III 10-O-acetyltransferase
(EC 2.3.1.167) (DBAT)
Length = 440
Score = 199 bits (505), Expect = 2e-50
Identities = 139/443 (31%), Positives = 214/443 (47%), Gaps = 21/443 (4%)
Query: 91 FTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKVLRH 150
F VR + +V P+ P+P+ LS +D+ G+R NI + + DP KV+R
Sbjct: 7 FVVRSLERVMVAPSQPSPKAFLQLSTLDNLPGVRENIFNTLLVYNASDRVSVDPAKVIRQ 66
Query: 151 ALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFEE 210
ALS+ LVYY PFAGR+R+ L V+CTGEG +FVEA AD L GD L P E+
Sbjct: 67 ALSKVLVYYSPFAGRLRKKENGDLEVECTGEGALFVEAMADTDLSVLGD-LDDYSPSLEQ 125
Query: 211 LLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGAH 270
LL+ +P I D ++QVTR CG F++ V H + DG G F+ A EMARG
Sbjct: 126 LLFCLPPDTDIEDIHPLVVQVTRFTCGGFVVGVSFCHGICDGLGAGQFLIAMGEMARGEI 185
Query: 271 KPSIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFFFRTSDIV-- 328
KPS +P+W RE+L DP + ++ ++ I +T + + Q TS+ +
Sbjct: 186 KPSSEPIWKRELLKPEDPLY-RFQYYHFQLICPPSTFGK------IVQGSLVITSETINC 238
Query: 329 VLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPLVG 388
+ + L C+ F+++++ W RT+ALQ+ + ++++ ++ R FN S G
Sbjct: 239 IKQCLREESKEFCSAFEVVSALAWIARTRALQIPHSENVKLIFAMDMRKLFNPPLSK--G 296
Query: 389 YYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVADLMVIKERCL-FTT 447
YYGN A+ L SL V +++K K + E + ++ + + +
Sbjct: 297 YYGNFVGTVCAMDNVKDLLSGSLLRVVRIIKKAKVSLNEHFTSTIVTPRSGSDESINYEN 356
Query: 448 IRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPGA----AYIIPHKNVEGEEGFI 503
I D R+ EVDFGWG A + G +I P KN +G
Sbjct: 357 IVG--FGDRRRLGFDEVDFGWGHADNVSLVQHGLKDVSVVQSYFLFIRPPKN--NPDGIK 412
Query: 504 LLVCLSFEAMKRFAKELDEMLGK 526
+L + +K F E++ M K
Sbjct: 413 ILSFMPPSIVKSFKFEMETMTNK 435
>DBBT_TAXCU (Q9FPW3) 2-alpha-hydroxytaxane 2-O-benzoyltransferase
(EC 2.3.1.166) (TBT) (2-debenzoyl-7,13-diacetylbaccatin
III-2-O-benzoyl transferase) (DBBT)
Length = 440
Score = 177 bits (448), Expect = 8e-44
Identities = 116/384 (30%), Positives = 194/384 (50%), Gaps = 15/384 (3%)
Query: 91 FTVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVKVLRH 150
F V + +V P +P+ + LS ID++ NI ++ S+ DP K +R
Sbjct: 4 FNVDMIERVIVAPCLQSPKNILHLSPIDNKTRGLTNILSVYNASQRVSVSA-DPAKTIRE 62
Query: 151 ALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFEE 210
ALS+ LVYY PFAGR+R L V+CTGEG +FVEA AD L D + P F++
Sbjct: 63 ALSKVLVYYPPFAGRLRNTENGDLEVECTGEGAVFVEAMADNDLSVLQD-FNEYDPSFQQ 121
Query: 211 LLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGAH 270
L++++ I D + +QVTR CG F++ +H++SDG G+ + EMARG
Sbjct: 122 LVFNLREDVNIEDLHLLTVQVTRFTCGGFVVGTRFHHSVSDGKGIGQLLKGMGEMARGEF 181
Query: 271 KPSIQPVWNREILMARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFFFRTSDIVVL 330
KPS++P+WNRE++ D ++ +H ++ I +++ ++V F R + +
Sbjct: 182 KPSLEPIWNREMVKPEDIMYLQFDHFDF--IHPPLNLEKSIQASMVIS--FERIN--YIK 235
Query: 331 RLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSRFNANNSPL-VGY 389
R ++ + F+++ + W RTK+ ++ ++ ++++ ++ R+ F +SPL GY
Sbjct: 236 RCMMEECKEFFSAFEVVVALIWLARTKSFRIPPNEYVKIIFPIDMRNSF---DSPLPKGY 292
Query: 390 YGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSV-ADLMVIKERCLFTTI 448
YGN A+ L SL YA+ L++K K + E + + + +
Sbjct: 293 YGNAIGNACAMDNVKDLLNGSLLYALMLIKKSKFALNENFKSRILTKPSTLDANMKHENV 352
Query: 449 RSCVVSDVTRIKLREVDFGWGEAV 472
C D + E DFGWG AV
Sbjct: 353 VGC--GDWRNLGFYEADFGWGNAV 374
>HCB1_DIACA (O24645) Anthranilate N-benzoyltransferase protein 1 (EC
2.3.1.144) (Anthranilate
N-hydroxycinnamoyl/benzoyltransferase 1)
Length = 445
Score = 146 bits (369), Expect = 1e-34
Identities = 123/451 (27%), Positives = 210/451 (46%), Gaps = 36/451 (7%)
Query: 92 TVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFN-IPMMFIYRH--------EPSMKEK 142
+++ Q +V PA TP + LS+ID ++ + IY+ PS
Sbjct: 2 SIQIKQSTMVRPAEETPNKSLWLSNIDMILRTPYSHTGAVLIYKQPDNNEDNIHPSSSMY 61
Query: 143 DPVKVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALH 202
+L ALS+ALV +YP AGR++ G + +DC EG +FVEAE+ L++FGD
Sbjct: 62 FDANILIEALSKALVPFYPMAGRLKIN-GDRYEIDCNAEGALFVEAESSHVLEDFGD-FR 119
Query: 203 PPLPCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAW 262
P ++ S+ I P+ ++Q+TR +CG + +H + DG F N+W
Sbjct: 120 PNDELHRVMVPTCDYSKGISSFPLLMVQLTRFRCGGVSIGFAQHHHVCDGMAHFEFNNSW 179
Query: 263 AEMARGAHKPSIQPVWNREI-LMARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFF 321
A +A+G P+++PV +R + L R+PP I +H ++E S + D T Q+ F
Sbjct: 180 ARIAKGL-LPALEPVHDRYLHLRPRNPPQIKYSHSQFEPFVPSLPNELLDGKTNKSQTLF 238
Query: 322 FRTSD---IVVLRLLVPFHLRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSR 378
+ + + +L + + +T++++ + W +KA L +EI+++ V+ RSR
Sbjct: 239 ILSREQINTLKQKLDLSNNTTRLSTYEVVAAHVWRSVSKARGLSDHEEIKLIMPVDGRSR 298
Query: 379 FNANNSPLVGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVAD--- 435
N N S GY GN T G L N L V++ + ++Y+ S D
Sbjct: 299 IN-NPSLPKGYCGNVVFLAVCTATVGDLSCNPLTDTAGKVQEALKGLDDDYLRSAIDHTE 357
Query: 436 --------LMVIKERCLFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPGA 487
M E+ L+ + +V+ RI + +DFGWG + G + + G
Sbjct: 358 SKPGLPVPYMGSPEKTLYPNV---LVNSWGRIPYQAMDFGWGSPTFFGISNIF---YDGQ 411
Query: 488 AYIIPHKNVEGEEGFILLVCLSFEAMKRFAK 518
++IP + +G+ L + L + RF K
Sbjct: 412 CFLIPSR--DGDGSMTLAINLFSSHLSRFKK 440
>HCB3_DIACA (O23918) Anthranilate N-benzoyltransferase protein 3 (EC
2.3.1.144) (Anthranilate
N-hydroxycinnamoyl/benzoyltransferase 3)
Length = 445
Score = 144 bits (362), Expect = 7e-34
Identities = 121/431 (28%), Positives = 198/431 (45%), Gaps = 34/431 (7%)
Query: 92 TVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFN-IPMMFIYRH--------EPSMKEK 142
++ Q +V PA TP + LS ID ++ + IY+ +PS
Sbjct: 2 SIHIKQSTMVRPAEETPNKSLWLSKIDMILRTPYSHTGAVLIYKQPDNNEDNIQPSSSMY 61
Query: 143 DPVKVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALH 202
+L ALS+ALV YYP AGR++ G + +DC GEG +FVEAE+ L++FGD
Sbjct: 62 FDANILIEALSKALVPYYPMAGRLKIN-GDRYEIDCNGEGALFVEAESSHVLEDFGD-FR 119
Query: 203 PPLPCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAW 262
P ++ S+ I P+ ++Q+TR +CG + +H + D F N+W
Sbjct: 120 PNDELHRVMVPTCDYSKGISSFPLLMVQLTRFRCGGVSIGFAQHHHVCDRMSHFEFNNSW 179
Query: 263 AEMARGAHKPSIQPVWNREI-LMARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFF 321
A +A+G P+++PV +R + L R+PP I H ++E S + D T Q+ F
Sbjct: 180 ARIAKGL-LPALEPVHDRYLHLCPRNPPQIKYTHSQFEPFVPSLPKELLDGKTSKSQTLF 238
Query: 322 -FRTSDIVVLRLLVPFH--LRHCTTFDLITSCFWCCRTKALQLEADDEIRMMCIVNARSR 378
I L+ + + +T++++ W +KA L +EI+++ V+ RSR
Sbjct: 239 KLSREQINTLKQKLDWSNTTTRLSTYEVVAGHVWRSVSKARGLSDHEEIKLIMPVDGRSR 298
Query: 379 FNANNSPLVGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVAD--- 435
N N S GY GN T G L N L V++ + ++Y+ S D
Sbjct: 299 IN-NPSLPKGYCGNVVFLAVCTATVGDLACNPLTDTAGKVQEALKGLDDDYLRSAIDHTE 357
Query: 436 --------LMVIKERCLFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPGA 487
M E+ L+ + +V+ RI + +DFGWG + G + + G
Sbjct: 358 SKPDLPVPYMGSPEKTLYPNV---LVNSWGRIPYQAMDFGWGNPTFFGISNIF---YDGQ 411
Query: 488 AYIIPHKNVEG 498
++IP +N +G
Sbjct: 412 CFLIPSQNGDG 422
>HCB2_DIACA (O23917) Anthranilate N-benzoyltransferase protein 2 (EC
2.3.1.144) (Anthranilate
N-hydroxycinnamoyl/benzoyltransferase 2)
Length = 446
Score = 142 bits (359), Expect = 2e-33
Identities = 122/432 (28%), Positives = 196/432 (45%), Gaps = 35/432 (8%)
Query: 92 TVRRSQPELVPPAAPTPREVKLLSDIDDQEGLRFN-IPMMFIYRH--------EPSMKEK 142
+++ Q +V PA TP + LS ID ++ + IY+ PS
Sbjct: 2 SIQIKQSTMVRPAEETPNKSLWLSKIDMILRTPYSHTGAVLIYKQPDNNEDNIHPSSSMY 61
Query: 143 DPVKVLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALH 202
+L ALS+ALV YYP AGR++ G + +DC EG +FVEAE+ L++FGD
Sbjct: 62 FDANILIEALSKALVPYYPMAGRLKIN-GDRYEIDCNAEGALFVEAESSHVLEDFGD-FR 119
Query: 203 PPLPCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAW 262
P ++ S+ I P+ ++Q+TR +CG + +H DG F N+W
Sbjct: 120 PNDELHRVMVPTCDYSKGISSFPLLMVQLTRFRCGGVSIGFAQHHHACDGMSHFEFNNSW 179
Query: 263 AEMARGAHKPSIQPVWNREI-LMARDPPHITCNHHEYEQIFSSNTIKEEDTTTLVHQSFF 321
A +A+G P+++PV +R + L R+PP I H ++E S + D T Q+ F
Sbjct: 180 ARIAKGL-LPALEPVHDRYLHLRLRNPPQIKYTHSQFEPFVPSLPNELLDGKTNKSQTLF 238
Query: 322 -FRTSDIVVLRLLVPFHLRHCT---TFDLITSCFWCCRTKALQLEADDEIRMMCIVNARS 377
I L+ + T T++++ W +KA L +EI+++ V+ RS
Sbjct: 239 KLSREQINTLKQKLDLSSNTTTRLSTYEVVAGHVWRSVSKARGLSDHEEIKLIMPVDGRS 298
Query: 378 RFNANNSPLVGYYGNCFAYPAAVTTAGKLCGNSLGYAVELVRKLKAEVTEEYMHSVAD-- 435
R N N S GY GN T G L N L V++ + ++Y+ S D
Sbjct: 299 RIN-NPSLPKGYCGNVVFLAVCTATVGDLSCNPLTDTAGKVQEALKGLDDDYLRSAIDHT 357
Query: 436 ---------LMVIKERCLFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGAAKSGAGPFPG 486
M E+ L+ + +V+ RI + +DFGWG + G + + G
Sbjct: 358 ESKPDLPVPYMGSPEKTLYPNV---LVNSWGRIPYQAMDFGWGSPTFFGISNIF---YDG 411
Query: 487 AAYIIPHKNVEG 498
++IP +N +G
Sbjct: 412 QCFLIPSQNGDG 423
>SLAT_PAPSO (Q94FT4) Salutaridinol 7-O-acetyltransferase (EC
2.3.1.150) (salAT)
Length = 474
Score = 75.9 bits (185), Expect = 2e-13
Identities = 72/275 (26%), Positives = 124/275 (44%), Gaps = 29/275 (10%)
Query: 99 ELVPPAAPTPREVKL--LSDIDDQEGLRFNIPMMFIY-----RHEPSMKEKDPVKVLRHA 151
E + P PTP ++K LS +D L + +P++ Y S D + +L+ +
Sbjct: 15 ETIKPTTPTPSQLKNFNLSLLDQCFPLYYYVPIILFYPATAANSTGSSNHHDDLDLLKSS 74
Query: 152 LSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFEEL 211
LS+ LV++YP AGR+ + ++VDC +G+ F + + + EF PL +L
Sbjct: 75 LSKTLVHFYPMAGRMID----NILVDCHDQGINFYKVKIRGKMCEFMSQPDVPL---SQL 127
Query: 212 LYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGAHK 271
L S + + ++QV CG + ++H ++D A + F+ +WA + +
Sbjct: 128 LPSEVVSASVPKEALVIVQVNMFDCGGTAICSSVSHKIADAATMSTFIRSWASTTKTSRS 187
Query: 272 -PSIQPVWNREILMARD-----PP--HITCNHHEYEQIFSSNTIKEEDTTTLVHQSFFFR 323
S V +++++ + D PP +T E FSS ED T V + F F
Sbjct: 188 GGSTAAVTDQKLIPSFDSASLFPPSERLTSPSGMSEIPFSSTPEDTEDDKT-VSKRFVFD 246
Query: 324 TSDIVVLR--LLVPFH----LRHCTTFDLITSCFW 352
+ I +R L V H R T +++TS W
Sbjct: 247 FAKITSVREKLQVLMHDNYKSRRQTRVEVVTSLIW 281
>ANTA_GENTR (Q9ZWR8) Anthocyanin 5-aromatic acyltransferase (EC
2.3.1.153) (5AT)
Length = 469
Score = 40.4 bits (93), Expect = 0.011
Identities = 96/466 (20%), Positives = 174/466 (36%), Gaps = 58/466 (12%)
Query: 99 ELVPPAAPTPREVKLLSDIDDQEGLRFNIPMMFIYRHEPSMKEK---DPVKVLRHALSQA 155
++ PP+ T E+ L D L N ++ P + + L+ +LS
Sbjct: 14 QVTPPSDTTDVELSLPVTFFDIPWLHLNKMQSLLFYDFPYPRTHFLDTVIPNLKASLSLT 73
Query: 156 LVYYYPFAGR----IREGAGRKLMVDCT-GEGVMFVEAEADVTLDEF-------GDALHP 203
L +Y P +G I+ G K G+ + + AE+D D + LH
Sbjct: 74 LKHYVPLSGNLLMPIKSGEMPKFQYSRDEGDSITLIVAESDQDFDYLKGHQLVDSNDLHG 133
Query: 204 PLPCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWA 263
++ + ++I P+ +QVT + + +H+++D +F+NAWA
Sbjct: 134 LFYVMPRVIRTMQDYKVI---PLVAVQVTVFPNRGIAVALTAHHSIADAKSFVMFINAWA 190
Query: 264 EMARGAH-----KPSIQPVWNREILMARDPPHITCNHHEYEQIFS--SNTIKEEDTTTLV 316
+ + ++ P ++R I+ T +E + + S + V
Sbjct: 191 YINKFGKDADLLSANLLPSFDRSIIKDLYGLEETF-WNEMQDVLEMFSRFGSKPPRFNKV 249
Query: 317 HQSFFFRTSDIVVLRLLVPFHLR------HCTTFDLITSCFWCCRTKA-----LQLEADD 365
++ ++I L+ V +LR TTF + W C K+ + ++D
Sbjct: 250 RATYVLSLAEIQKLKNKV-LNLRGSEPTIRVTTFTMTCGYVWTCMVKSKDDVVSEESSND 308
Query: 366 EIRM---MCIVNARSRFNANNSPLVGYYGNCFAYPAAVTTAGKLCGN-SLGYAVELVRKL 421
E + + R P Y+GNC A A T +L G+ L AV +
Sbjct: 309 ENELEYFSFTADCRGLLTPPCPP--NYFGNCLASCVAKATHKELVGDKGLLVAVAAI--- 363
Query: 422 KAEVTEEYMHSVADLMV-----IKERCLFTTIRSCVVSDVTRIKLREVDFGWGEAVYGGA 476
E E+ +H+ ++ + E + R ++ + VDFGWG+
Sbjct: 364 -GEAIEKRLHNEKGVLADAKTWLSESNGIPSKRFLGITGSPKFDSYGVDFGWGK-----P 417
Query: 477 AKSGAGPFPGAAYIIPHKNVEGEEGFILLVCLSFEAMKRFAKELDE 522
AK A I ++ + E+G + V L M FAK +E
Sbjct: 418 AKFDITSVDYAELIYVIQSRDFEKGVEIGVSLPKIHMDAFAKIFEE 463
>LEU3_SULTO (P50455) 3-isopropylmalate dehydrogenase (EC 1.1.1.85)
(Beta-IPM dehydrogenase) (IMDH) (3-IPM-DH)
Length = 336
Score = 32.7 bits (73), Expect = 2.4
Identities = 26/108 (24%), Positives = 45/108 (41%), Gaps = 2/108 (1%)
Query: 414 AVELVRKLKAEVTEEYMHSVADLMVIKERCLFTTIRSCVVSDVTRIKLREV--DFGWGEA 471
A V K K E +E Y+ + A +V + + V D+ + ++ G +
Sbjct: 183 ACRSVLKGKVEYSEMYVDAAAANLVRNPQMFDVIVTENVYGDILSDEASQIAGSLGIAPS 242
Query: 472 VYGGAAKSGAGPFPGAAYIIPHKNVEGEEGFILLVCLSFEAMKRFAKE 519
G K+ P GAA+ I KN+ F+L V + +E M + +
Sbjct: 243 ANIGDKKALFEPVHGAAFDIAGKNIGNPTAFLLSVSMMYERMYELSND 290
>BACC_BACLI (O68008) Bacitracin synthetase 3 (BA3) [Includes:
ATP-dependent isoleucine adenylase (IleA) (Isoleucine
activase); ATP-dependent D-phenylalanine adenylase
(D-PheA) (D-phenylalanine activase); ATP-dependent
histidine adenylase (HisA) (Histidine
Length = 6359
Score = 32.3 bits (72), Expect = 3.1
Identities = 27/108 (25%), Positives = 46/108 (42%), Gaps = 4/108 (3%)
Query: 167 REGAGRKLMVDCTGEGVMFVEAEADVTLDEFGDALHPPLPCFEELLYDVPGSELIIDRPI 226
R A R V GE V +E E D + ++G PL EE + + P+
Sbjct: 3637 RHEALRTAFVMVDGEPVQKIEKEVDFKV-KYGRLGQDPL---EEKIKAFIKPFALEKAPL 3692
Query: 227 RLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEMARGAHKPSI 274
+V + +L++ ++H +SDG + +F AE+ G P +
Sbjct: 3693 LRAEVLKASGDEHVLMLDMHHIISDGVSMAIFTRELAELYEGKTLPPL 3740
>TYCC_BREPA (O30409) Tyrocidine synthetase III [Includes:
ATP-dependent asparagine adenylase (AsnA) (Asparagine
activase); ATP-dependent glutamine adenylase (GlnA)
(Glutamine activase); ATP-dependent tyrosine adenylase
(TyrA) (Tyrosine activase); ATP-depe
Length = 6486
Score = 32.0 bits (71), Expect = 4.1
Identities = 17/62 (27%), Positives = 33/62 (52%), Gaps = 8/62 (12%)
Query: 228 LIQVTRLKCGS--FILVVYLNHTMSDGAGLKLFMNAWAEMARGAHKPSIQ------PVWN 279
LI+V+ LK G ++L ++H++SDG + + W ++ +G P ++ VW
Sbjct: 5314 LIRVSLLKIGEDRYVLFTDMHHSISDGVSSGILLAEWVQLYQGDVLPELRIQYKDFAVWQ 5373
Query: 280 RE 281
+E
Sbjct: 5374 QE 5375
>ACHB_XENLA (P49579) Acetylcholine receptor protein, beta chain
precursor (Fragment)
Length = 489
Score = 32.0 bits (71), Expect = 4.1
Identities = 19/66 (28%), Positives = 29/66 (43%), Gaps = 4/66 (6%)
Query: 54 FCIFVVTTDFRFTYTSHLLTHTTLIFSYHIMTSAPLLFTVRRSQPELVPPAAPTPREVKL 113
F + V+ R T H+ IF +++ P +RR +PE P AP PR+V
Sbjct: 316 FSVVVLNLHHRSPNTHHMPQWVKQIFIHYL----PKYLCIRRPKPETPLPVAPPPRQVTS 371
Query: 114 LSDIDD 119
D+
Sbjct: 372 TRHADE 377
>BACB_BACLI (O68007) Bacitracin synthetase 2 (BA2) [Includes:
ATP-dependent lysine adenylase (LysA) (Lysine activase);
ATP-dependent D-ornithine adenylase (D-OrnA)
(D-ornithine activase); Ornithine racemase (EC
5.1.1.12)]
Length = 2607
Score = 31.6 bits (70), Expect = 5.3
Identities = 44/184 (23%), Positives = 80/184 (42%), Gaps = 30/184 (16%)
Query: 93 VRRSQPELVPPAA---PTPREVK---LLSDIDDQEGLRFNIPMMFIYRHEPSMKEKDPVK 146
+ RS P+ P + P RE K +L+ +DD + +N+P+ ++K V+
Sbjct: 61 IERSFPKAAPSKSGTYPLSREQKRMFILNQLDDSK-TAYNMPL--------AVKINGEVQ 111
Query: 147 VLRHALSQALVYYYPFAGRIREGAGRKLMVDCTGEGVMFVEAEADVTLD--EFGDALHPP 204
+ R L QA R + R V GE V +E EA+ L+ E GD
Sbjct: 112 ISR--LEQAWKALIK-----RHESLRTSFVMLDGEPVQKIEQEAEFRLEYSELGDQ---- 160
Query: 205 LPCFEELLYDVPGSELIIDRPIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAE 264
E++ + EL P+ ++ ++ +++V ++H +SDG + + M +A+
Sbjct: 161 -SIQEKISRFIKPFELE-KAPLLRAEIVKVDEAEHMMMVDMHHIISDGVSIGILMKEFAD 218
Query: 265 MARG 268
G
Sbjct: 219 CCEG 222
Score = 30.8 bits (68), Expect = 9.1
Identities = 11/41 (26%), Positives = 23/41 (55%)
Query: 225 PIRLIQVTRLKCGSFILVVYLNHTMSDGAGLKLFMNAWAEM 265
P+ ++ R+ IL++ ++H +SDG + + M WA +
Sbjct: 1210 PLLRAELVRVNAAEHILLLDMHHIISDGVSIGILMKEWAAL 1250
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.326 0.140 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 63,662,284
Number of Sequences: 164201
Number of extensions: 2723187
Number of successful extensions: 7200
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 7139
Number of HSP's gapped (non-prelim): 24
length of query: 528
length of database: 59,974,054
effective HSP length: 115
effective length of query: 413
effective length of database: 41,090,939
effective search space: 16970557807
effective search space used: 16970557807
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC148345.17


