

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148289.16 + phase: 0
(200 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
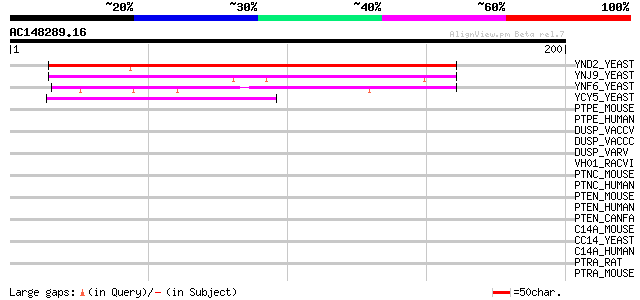
Score E
Sequences producing significant alignments: (bits) Value
YND2_YEAST (P53965) Hypothetical 32.8 kDa protein in NCE3-HHT2 i... 156 3e-38
YNJ9_YEAST (P50946) Hypothetical 27.2 kDa protein in POL1-RAS2 i... 101 1e-21
YNF6_YEAST (P53949) Hypothetical 22.5 kDa protein in ARP5-OMP2 i... 94 3e-19
YCY5_YEAST (P25366) Hypothetical 41.6 kDa protein in HMR 5'region 44 2e-04
PTPE_MOUSE (P49446) Protein-tyrosine phosphatase epsilon precurs... 39 0.010
PTPE_HUMAN (P23469) Protein-tyrosine phosphatase epsilon precurs... 38 0.018
DUSP_VACCV (P07239) Dual specificity protein phosphatase (EC 3.1... 37 0.040
DUSP_VACCC (P20495) Dual specificity protein phosphatase (EC 3.1... 37 0.040
DUSP_VARV (P33064) Dual specificity protein phosphatase (EC 3.1.... 36 0.067
VH01_RACVI (P80994) Dual specificity protein phosphatase (EC 3.1... 35 0.088
PTNC_MOUSE (P35831) Protein-tyrosine phosphatase, non-receptor t... 35 0.088
PTNC_HUMAN (Q05209) Protein-tyrosine phosphatase, non-receptor t... 35 0.088
PTEN_MOUSE (O08586) Phosphatidylinositol-3,4,5-trisphosphate 3-p... 35 0.11
PTEN_HUMAN (P60484) Phosphatidylinositol-3,4,5-trisphosphate 3-p... 35 0.11
PTEN_CANFA (P60483) Phosphatidylinositol-3,4,5-trisphosphate 3-p... 35 0.11
C14A_MOUSE (Q6GQT0) Dual specificity protein phosphatase CDC14A ... 34 0.20
CC14_YEAST (Q00684) Probable protein-tyrosine phosphatase CDC14 ... 34 0.26
C14A_HUMAN (Q9UNH5) Dual specificity protein phosphatase CDC14A ... 34 0.26
PTRA_RAT (Q03348) Protein-tyrosine phosphatase alpha precursor (... 33 0.44
PTRA_MOUSE (P18052) Protein-tyrosine phosphatase alpha precursor... 33 0.44
>YND2_YEAST (P53965) Hypothetical 32.8 kDa protein in NCE3-HHT2
intergenic region
Length = 281
Score = 156 bits (394), Expect = 3e-38
Identities = 76/148 (51%), Positives = 103/148 (69%), Gaps = 1/148 (0%)
Query: 15 LIPPPNFSMVEDCIYRSSLPKPSSFPFL-QTLNLRSIIYLCPEPYPEENLDFLKEQNIRL 73
+IPP NFS V IYRSS P+ +F FL + L L+SI+ L PE YP+ENL+FLK I+L
Sbjct: 116 VIPPENFSHVVGEIYRSSFPRQENFSFLHERLKLKSILVLIPEEYPQENLNFLKLTGIKL 175
Query: 74 FQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNWC 133
+Q G+ G E + + +AL+++++ N PIL+HC +GKHRTGCL+GC RKLQNW
Sbjct: 176 YQVGMSGNKEPFVNIPSHLLTKALEIVLNPANQPILIHCNRGKHRTGCLIGCIRKLQNWS 235
Query: 134 LSSAFEEYQRFAGVKSRAADLTFIERFD 161
L+ F+EY+RFA K+RA D FIE +D
Sbjct: 236 LTMIFDEYRRFAFPKARALDQQFIEMYD 263
>YNJ9_YEAST (P50946) Hypothetical 27.2 kDa protein in POL1-RAS2
intergenic region
Length = 238
Score = 101 bits (252), Expect = 1e-21
Identities = 59/152 (38%), Positives = 84/152 (54%), Gaps = 5/152 (3%)
Query: 15 LIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLF 74
++PP NF VE +YRS P P +FPFL L L++II+L E + L+F I L
Sbjct: 67 IVPPLNFCPVERYLYRSGQPSPVNFPFLLNLKLKTIIWLSNEEPQDTLLEFCDTHRINLQ 126
Query: 75 QFGIE---GKTEVSLPALRD-SIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQ 130
I G+ + L + SI+ L+ +V N+P+LV C G+HRTG ++GC R++
Sbjct: 127 FAAINPDAGEDDNPWDGLTEHSIINVLQTIVTQENYPLLVCCGMGRHRTGTVIGCLRRIM 186
Query: 131 NWCLSSAFEEYQRFAGVK-SRAADLTFIERFD 161
W L+S EEY+RF G + R IE FD
Sbjct: 187 GWNLASVSEEYRRFTGSRGGRILVELLIEAFD 218
>YNF6_YEAST (P53949) Hypothetical 22.5 kDa protein in ARP5-OMP2
intergenic region
Length = 197
Score = 93.6 bits (231), Expect = 3e-19
Identities = 55/152 (36%), Positives = 88/152 (57%), Gaps = 9/152 (5%)
Query: 16 IPPPNFSMV---EDCIYRSSLPKPSSFPFLQ-TLNLRSIIYLCPEPYP-EENLDFLKEQN 70
IPP NFS V + +YRS P P ++ F++ L+L++IIY+ + P EE FL+ +
Sbjct: 4 IPPLNFSPVVSTDVSLYRSGYPMPLNYSFIKHQLHLKTIIYIGDKDRPLEEYQSFLESEK 63
Query: 71 IRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRK-L 129
I+ + ++ + +++ + + L +++DVRN+PILVH +GKHR G +VG RK L
Sbjct: 64 IKYYHIFMDSSRD---EGIQERMNQVLHLVLDVRNYPILVHSNKGKHRVGVVVGIIRKLL 120
Query: 130 QNWCLSSAFEEYQRFAGVKSRAADLTFIERFD 161
Q W + +EY F+G DL FI F+
Sbjct: 121 QGWSTAGICQEYGLFSGGMKDGVDLEFITMFE 152
>YCY5_YEAST (P25366) Hypothetical 41.6 kDa protein in HMR 5'region
Length = 362
Score = 44.3 bits (103), Expect = 2e-04
Identities = 23/83 (27%), Positives = 43/83 (51%)
Query: 14 VLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRL 73
+L+PP NF + E+ IYR S + + FL+TLNL++ I++ + + DF +I+
Sbjct: 1 MLVPPANFGIAEEGIYRCSKVETLNLSFLETLNLKTAIFIGGQEPSKFFKDFFTRSSIKW 60
Query: 74 FQFGIEGKTEVSLPALRDSIMEA 96
+ + ++P S+ A
Sbjct: 61 IVLRMSDFSAAAVPVKSSSVSNA 83
Score = 37.4 bits (85), Expect = 0.023
Identities = 20/66 (30%), Positives = 34/66 (51%), Gaps = 5/66 (7%)
Query: 93 IMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNWCLSSAFEEYQRFAGVKSRAA 152
+ K L++V N+ +L+ K T ++G RK+Q W ++S EY+ F+G
Sbjct: 139 LKRTFKTLLNVDNYNVLLVDK-----TALVIGILRKIQKWNIASIINEYRLFSGKNRNYF 193
Query: 153 DLTFIE 158
TF+E
Sbjct: 194 AETFLE 199
>PTPE_MOUSE (P49446) Protein-tyrosine phosphatase epsilon precursor
(EC 3.1.3.48) (R-PTP-epsilon)
Length = 699
Score = 38.5 bits (88), Expect = 0.010
Identities = 23/62 (37%), Positives = 30/62 (48%), Gaps = 5/62 (8%)
Query: 67 KEQNIRLF-QFGIEGKTEVSLPA----LRDSIMEALKVLVDVRNHPILVHCKQGKHRTGC 121
+E+ +R+ QF G EV +PA + D I K NHPI VHC G RTG
Sbjct: 579 QEEQVRMVRQFHFHGWPEVGIPAEGKGMIDLIAAVQKQQQQTGNHPITVHCSAGAGRTGT 638
Query: 122 LV 123
+
Sbjct: 639 FI 640
>PTPE_HUMAN (P23469) Protein-tyrosine phosphatase epsilon precursor
(EC 3.1.3.48) (R-PTP-epsilon)
Length = 700
Score = 37.7 bits (86), Expect = 0.018
Identities = 22/62 (35%), Positives = 30/62 (47%), Gaps = 5/62 (8%)
Query: 67 KEQNIRLF-QFGIEGKTEVSLPA----LRDSIMEALKVLVDVRNHPILVHCKQGKHRTGC 121
+E+ +R+ QF G E+ +PA + D I K NHPI VHC G RTG
Sbjct: 580 QEEQVRVVRQFHFHGWPEIGIPAEGKGMIDLIAAVQKQQQQTGNHPITVHCSAGAGRTGT 639
Query: 122 LV 123
+
Sbjct: 640 FI 641
>DUSP_VACCV (P07239) Dual specificity protein phosphatase (EC
3.1.3.48) (EC 3.1.3.16) (Late protein H1)
Length = 171
Score = 36.6 bits (83), Expect = 0.040
Identities = 27/104 (25%), Positives = 42/104 (39%), Gaps = 14/104 (13%)
Query: 29 YRSSLPKPSS-FPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLP 87
Y++++ PSS F LNL Y P NI + + T +
Sbjct: 39 YKNAMDAPSSEVKFKYVLNLTMDKYTLPN------------SNINIIHIPLVDDTTTDIS 86
Query: 88 ALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQN 131
D + L D RN P+LVHC G +R+G ++ + +N
Sbjct: 87 KYFDDVTAFLSKC-DQRNEPVLVHCAAGVNRSGAMILAYLMSKN 129
>DUSP_VACCC (P20495) Dual specificity protein phosphatase (EC
3.1.3.48) (EC 3.1.3.16) (Late protein H1)
Length = 171
Score = 36.6 bits (83), Expect = 0.040
Identities = 27/104 (25%), Positives = 42/104 (39%), Gaps = 14/104 (13%)
Query: 29 YRSSLPKPSS-FPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLP 87
Y++++ PSS F LNL Y P NI + + T +
Sbjct: 39 YKNAMDAPSSEVKFKYVLNLTMDKYTLPN------------SNINIIHIPLVDDTTTDIS 86
Query: 88 ALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQN 131
D + L D RN P+LVHC G +R+G ++ + +N
Sbjct: 87 KYFDDVTAFLSKC-DQRNEPVLVHCAAGVNRSGAMILAYLMSKN 129
>DUSP_VARV (P33064) Dual specificity protein phosphatase (EC
3.1.3.48) (EC 3.1.3.16) (Late protein H1)
Length = 171
Score = 35.8 bits (81), Expect = 0.067
Identities = 27/104 (25%), Positives = 42/104 (39%), Gaps = 14/104 (13%)
Query: 29 YRSSLPKPSS-FPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLP 87
Y++++ PSS F LNL Y P NI + + T +
Sbjct: 39 YKNAMNAPSSEVKFKYVLNLTMDKYTLPN------------SNINIIHIPLVDDTTTDIS 86
Query: 88 ALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQN 131
D + L D RN P+LVHC G +R+G ++ + +N
Sbjct: 87 KYFDDVTAFLSKC-DQRNEPVLVHCVAGVNRSGAMILAYLMSKN 129
>VH01_RACVI (P80994) Dual specificity protein phosphatase (EC
3.1.3.48) (EC 3.1.3.16) (Late protein H1)
Length = 171
Score = 35.4 bits (80), Expect = 0.088
Identities = 26/103 (25%), Positives = 44/103 (42%), Gaps = 12/103 (11%)
Query: 29 YRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLPA 88
Y++++ PSS + + I+ L + Y N NI + + T +
Sbjct: 39 YKNAMEAPSS-----EVKFKYILNLTMDKYSFTN------SNINIIHVPMVDDTSTDISI 87
Query: 89 LRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQN 131
D I L D RN P+LVHC G +R+G ++ + +N
Sbjct: 88 YFDDITAFLSKC-DQRNEPVLVHCAAGVNRSGAMILAYLMSKN 129
>PTNC_MOUSE (P35831) Protein-tyrosine phosphatase, non-receptor type
12 (EC 3.1.3.48) (Protein-tyrosine phosphatase P19)
(P19-PTP) (MPTP-PEST)
Length = 775
Score = 35.4 bits (80), Expect = 0.088
Identities = 20/63 (31%), Positives = 33/63 (51%), Gaps = 4/63 (6%)
Query: 63 LDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNH---PILVHCKQGKHRT 119
L+F E RL+QF + +P+ DSI++ + ++ + H PI +HC G RT
Sbjct: 180 LEFQNESR-RLYQFHYVNWPDHDVPSSFDSILDMISLMRKYQEHEDVPICIHCSAGCGRT 238
Query: 120 GCL 122
G +
Sbjct: 239 GAI 241
>PTNC_HUMAN (Q05209) Protein-tyrosine phosphatase, non-receptor type
12 (EC 3.1.3.48) (Protein-tyrosine phosphatase G1)
(PTPG1)
Length = 780
Score = 35.4 bits (80), Expect = 0.088
Identities = 20/63 (31%), Positives = 33/63 (51%), Gaps = 4/63 (6%)
Query: 63 LDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNH---PILVHCKQGKHRT 119
L+F E RL+QF + +P+ DSI++ + ++ + H PI +HC G RT
Sbjct: 180 LEFQNESR-RLYQFHYVNWPDHDVPSSFDSILDMISLMRKYQEHEDVPICIHCSAGCGRT 238
Query: 120 GCL 122
G +
Sbjct: 239 GAI 241
>PTEN_MOUSE (O08586) Phosphatidylinositol-3,4,5-trisphosphate
3-phosphatase PTEN (EC 3.1.3.67) (Mutated in multiple
advanced cancers 1)
Length = 403
Score = 35.0 bits (79), Expect = 0.12
Identities = 30/123 (24%), Positives = 51/123 (41%), Gaps = 6/123 (4%)
Query: 28 IYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLP 87
+YR+++ F + N I LC E + + + N R+ Q+ E L
Sbjct: 45 VYRNNIDDVVRFLDSKHKNHYKIYNLCAERHYDT-----AKFNCRVAQYPFEDHNPPQLE 99
Query: 88 ALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNWCLSSAFEEYQRFAGV 147
++ + + L + NH +HCK GK RTG ++ C L A E + V
Sbjct: 100 LIKPFCEDLDQWLSEDDNHVAAIHCKAGKGRTGVMI-CAYLLHRGKFLKAQEALDFYGEV 158
Query: 148 KSR 150
++R
Sbjct: 159 RTR 161
>PTEN_HUMAN (P60484) Phosphatidylinositol-3,4,5-trisphosphate
3-phosphatase PTEN (EC 3.1.3.67) (Mutated in multiple
advanced cancers 1)
Length = 403
Score = 35.0 bits (79), Expect = 0.12
Identities = 30/123 (24%), Positives = 51/123 (41%), Gaps = 6/123 (4%)
Query: 28 IYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLP 87
+YR+++ F + N I LC E + + + N R+ Q+ E L
Sbjct: 45 VYRNNIDDVVRFLDSKHKNHYKIYNLCAERHYDT-----AKFNCRVAQYPFEDHNPPQLE 99
Query: 88 ALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNWCLSSAFEEYQRFAGV 147
++ + + L + NH +HCK GK RTG ++ C L A E + V
Sbjct: 100 LIKPFCEDLDQWLSEDDNHVAAIHCKAGKGRTGVMI-CAYLLHRGKFLKAQEALDFYGEV 158
Query: 148 KSR 150
++R
Sbjct: 159 RTR 161
>PTEN_CANFA (P60483) Phosphatidylinositol-3,4,5-trisphosphate
3-phosphatase PTEN (EC 3.1.3.67) (Mutated in multiple
advanced cancers 1)
Length = 403
Score = 35.0 bits (79), Expect = 0.12
Identities = 30/123 (24%), Positives = 51/123 (41%), Gaps = 6/123 (4%)
Query: 28 IYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLP 87
+YR+++ F + N I LC E + + + N R+ Q+ E L
Sbjct: 45 VYRNNIDDVVRFLDSKHKNHYKIYNLCAERHYDT-----AKFNCRVAQYPFEDHNPPQLE 99
Query: 88 ALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQNWCLSSAFEEYQRFAGV 147
++ + + L + NH +HCK GK RTG ++ C L A E + V
Sbjct: 100 LIKPFCEDLDQWLSEDDNHVAAIHCKAGKGRTGVMI-CAYLLHRGKFLKAQEALDFYGEV 158
Query: 148 KSR 150
++R
Sbjct: 159 RTR 161
>C14A_MOUSE (Q6GQT0) Dual specificity protein phosphatase CDC14A (EC
3.1.3.48) (EC 3.1.3.16) (CDC14 cell division cycle 14
homolog A)
Length = 554
Score = 34.3 bits (77), Expect = 0.20
Identities = 24/88 (27%), Positives = 40/88 (45%), Gaps = 8/88 (9%)
Query: 39 FPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALK 98
FP+ + N+ +I+ L + Y + ++ LF I+G T D+I+
Sbjct: 164 FPYFKKNNVTTIVRLNKKIYEAKRFTDAGFEHYDLFF--IDGSTP------SDNIVRRFL 215
Query: 99 VLVDVRNHPILVHCKQGKHRTGCLVGCF 126
+ + I VHCK G RTG L+ C+
Sbjct: 216 NICENTEGAIAVHCKAGLGRTGTLIACY 243
>CC14_YEAST (Q00684) Probable protein-tyrosine phosphatase CDC14 (EC
3.1.3.48)
Length = 551
Score = 33.9 bits (76), Expect = 0.26
Identities = 25/91 (27%), Positives = 43/91 (46%), Gaps = 5/91 (5%)
Query: 34 PKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLPALRDSI 93
P S F N++ ++ L Y +++ + + Q++ L E T L +++ +
Sbjct: 210 PFKSVLNFFANNNVQLVVRLNSHLYNKKHFEDIGIQHLDLI---FEDGTCPDLSIVKNFV 266
Query: 94 MEALKVLVDVRNHPILVHCKQGKHRTGCLVG 124
A ++ R I VHCK G RTGCL+G
Sbjct: 267 GAAETIIK--RGGKIAVHCKAGLGRTGCLIG 295
>C14A_HUMAN (Q9UNH5) Dual specificity protein phosphatase CDC14A (EC
3.1.3.48) (EC 3.1.3.16) (CDC14 cell division cycle 14
homolog A)
Length = 594
Score = 33.9 bits (76), Expect = 0.26
Identities = 23/88 (26%), Positives = 40/88 (45%), Gaps = 8/88 (9%)
Query: 39 FPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALK 98
FP+ + N+ +++ L + Y + ++ LF I+G T D+I+
Sbjct: 213 FPYFKKHNVTAVVRLNKKIYEAKRFTDAGFEHYDLFF--IDGSTP------SDNIVRRFL 264
Query: 99 VLVDVRNHPILVHCKQGKHRTGCLVGCF 126
+ + I VHCK G RTG L+ C+
Sbjct: 265 NICENTEGAIAVHCKAGLGRTGTLIACY 292
>PTRA_RAT (Q03348) Protein-tyrosine phosphatase alpha precursor (EC
3.1.3.48) (R-PTP-alpha)
Length = 796
Score = 33.1 bits (74), Expect = 0.44
Identities = 22/58 (37%), Positives = 27/58 (45%), Gaps = 6/58 (10%)
Query: 67 KEQNIRLFQFGIEGKTEVSLPA----LRDSIMEALKVLVDVRNHPILVHCKQGKHRTG 120
K + IR F F G EV +P+ + + I K NHPI VHC G RTG
Sbjct: 679 KSRQIRQFHF--HGWPEVGIPSDGKGMINIIAAVQKQQQQSGNHPITVHCSAGAGRTG 734
>PTRA_MOUSE (P18052) Protein-tyrosine phosphatase alpha precursor
(EC 3.1.3.48) (R-PTP-alpha) (LCA-related phosphatase)
(PTPTY-28)
Length = 829
Score = 33.1 bits (74), Expect = 0.44
Identities = 22/58 (37%), Positives = 27/58 (45%), Gaps = 6/58 (10%)
Query: 67 KEQNIRLFQFGIEGKTEVSLPA----LRDSIMEALKVLVDVRNHPILVHCKQGKHRTG 120
K + IR F F G EV +P+ + + I K NHPI VHC G RTG
Sbjct: 712 KSRQIRQFHF--HGWPEVGIPSDGKGMINIIAAVQKQQQQSGNHPITVHCSAGAGRTG 767
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.141 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,336,249
Number of Sequences: 164201
Number of extensions: 935997
Number of successful extensions: 2277
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 35
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 2227
Number of HSP's gapped (non-prelim): 69
length of query: 200
length of database: 59,974,054
effective HSP length: 105
effective length of query: 95
effective length of database: 42,732,949
effective search space: 4059630155
effective search space used: 4059630155
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC148289.16


