

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148241.11 + phase: 0
(390 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
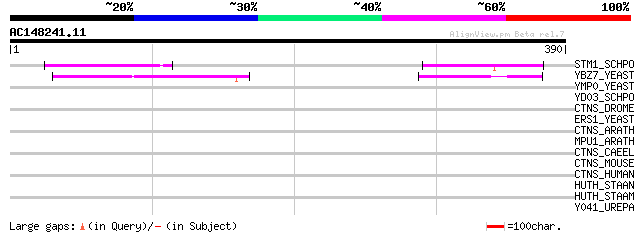
Score E
Sequences producing significant alignments: (bits) Value
STM1_SCHPO (Q10482) Seven transmembrane protein 1 70 9e-12
YBZ7_YEAST (P38279) Hypothetical 33.5 kDa protein in MRPS9-YSW1 ... 60 1e-08
YMP0_YEAST (Q03687) Hypothetical 46.9 kDa protein in PLB1-HXT2 i... 39 0.023
YD03_SCHPO (Q10227) Hypothetical protein C2E12.03c in chromosome I 39 0.030
CTNS_DROME (Q9VCR7) Cystinosin homolog 37 0.066
ERS1_YEAST (P17261) Transmembrane protein ERS1 (ERD suppressor) 37 0.086
CTNS_ARATH (P57758) Cystinosin homolog 35 0.33
MPU1_ARATH (Q9LTI3) Mannose-P-dolichol utilization defect 1 prot... 32 2.1
CTNS_CAEEL (Q09500) Cystinosin homolog 32 2.1
CTNS_MOUSE (P57757) Cystinosin 32 3.6
CTNS_HUMAN (O60931) Cystinosin 32 3.6
HUTH_STAAN (P64416) Histidine ammonia-lyase (EC 4.3.1.3) (Histid... 31 6.2
HUTH_STAAM (P64415) Histidine ammonia-lyase (EC 4.3.1.3) (Histid... 31 6.2
Y041_UREPA (Q9PRA4) Hypothetical protein UU041 30 8.1
>STM1_SCHPO (Q10482) Seven transmembrane protein 1
Length = 271
Score = 70.1 bits (170), Expect = 9e-12
Identities = 40/87 (45%), Positives = 50/87 (56%), Gaps = 2/87 (2%)
Query: 291 GWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILV--RTTEFESIK 348
G I + +Y C+RIPQI N K S EGL+ FV A + NTSY SILV +
Sbjct: 182 GCISSVLYFCARIPQIIKNHKAKSTEGLSIIFFVLASVGNTSYAFSILVFPASDYLNYTY 241
Query: 349 ANLPWLLDATVCVALDFFIISQYIYYR 375
ANLPW+L A + LD +I Q+I YR
Sbjct: 242 ANLPWILGAFSTIFLDIYIFYQFIKYR 268
Score = 51.6 bits (122), Expect = 3e-06
Identities = 27/90 (30%), Positives = 49/90 (54%), Gaps = 1/90 (1%)
Query: 25 LCNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCL 84
+ N+ ++S LG +SL W V IPQ++ ++N+S IS FL+ W+ GD N++G +
Sbjct: 10 MANILTELSSFLGALSLGCWVVLLIPQLLENYKNQSGESISDLFLIIWLIGDFFNVLGSI 69
Query: 85 LEPATLPTQFYTALTVINCSFMQALQSYYY 114
+ + +++ S + +Q YYY
Sbjct: 70 YGNVSSTVLVLSFYYIVSDSTL-LMQIYYY 98
Score = 33.9 bits (76), Expect = 0.73
Identities = 21/63 (33%), Positives = 34/63 (53%), Gaps = 3/63 (4%)
Query: 30 DDIS---FSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLE 86
DD++ F+ G +S V + A IPQII + KS+ G+S+ F + G+ L+
Sbjct: 172 DDLNAWPFTAGCISSVLYFCARIPQIIKNHKAKSTEGLSIIFFVLASVGNTSYAFSILVF 231
Query: 87 PAT 89
PA+
Sbjct: 232 PAS 234
>YBZ7_YEAST (P38279) Hypothetical 33.5 kDa protein in MRPS9-YSW1
intergenic region
Length = 296
Score = 60.1 bits (144), Expect = 1e-08
Identities = 32/87 (36%), Positives = 49/87 (55%), Gaps = 11/87 (12%)
Query: 288 QYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESI 347
Q LG++ A +Y SRIPQI LN KR S EG++ F+FA + NTS++ S+L
Sbjct: 209 QILGYLSAILYLGSRIPQIVLNFKRKSCEGVSFLFFLFACLGNTSFIISVL--------- 259
Query: 348 KANLPWLLDATVCVALDFFIISQYIYY 374
+ WL+ + + +DF + Q+ Y
Sbjct: 260 --SASWLIGSAGTLLMDFTVFIQFFLY 284
Score = 60.1 bits (144), Expect = 1e-08
Identities = 42/142 (29%), Positives = 67/142 (46%), Gaps = 5/142 (3%)
Query: 31 DISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCLLEPATL 90
++S G +S+ W V +PQI FR +S+ G+SL F++ W+ GDI N++G +++ L
Sbjct: 12 NLSGMAGSISICCWIVVFVPQIYENFRRQSAEGLSLLFIVLWLLGDIFNVMGAMMQ-NLL 70
Query: 91 PTQFYTALTVINCSFMQALQSYYYCKLYTMITSSDGANIVRMLRVNCYFYNCIILQDNEE 150
PT A + +Q +Y K I N+ + N +LQD
Sbjct: 71 PTMIILAAYYTLADLILLIQCMWYDKEKKSILQEVKKNVDPVHLPPANPINETVLQDVFN 130
Query: 151 EKRPLNPK----PSQVYSGIAI 168
E PL P+ SQ YS + +
Sbjct: 131 EYEPLLPRIEEEDSQSYSSLEL 152
Score = 32.0 bits (71), Expect = 2.8
Identities = 16/49 (32%), Positives = 26/49 (52%)
Query: 36 LGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLVGCL 84
LG +S + + + IPQI+ F+ KS G+S F L G+ ++ L
Sbjct: 211 LGYLSAILYLGSRIPQIVLNFKRKSCEGVSFLFFLFACLGNTSFIISVL 259
>YMP0_YEAST (Q03687) Hypothetical 46.9 kDa protein in PLB1-HXT2
intergenic region
Length = 405
Score = 38.9 bits (89), Expect = 0.023
Identities = 24/91 (26%), Positives = 45/91 (49%), Gaps = 5/91 (5%)
Query: 287 GQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFES 346
G +G + + + +PQI + K SV+G + V L +T + ++ +
Sbjct: 253 GSIIGSLGLLVESLLPLPQIAILYKLKSVQGFKLILLVSWLCGDTLKITYLIFGAKNISA 312
Query: 347 IKANLPWLLDATVCVALDFFIISQYIYYRYF 377
+ +++ A ++LDF+I QYIYYRY+
Sbjct: 313 L-----FVIFALFQMSLDFYIGGQYIYYRYY 338
Score = 35.8 bits (81), Expect = 0.19
Identities = 25/102 (24%), Positives = 46/102 (44%), Gaps = 8/102 (7%)
Query: 14 VKWVETYFKDCLCN---LRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLL 70
V+ + T+F + N L + +G + L+ + +PQI +++ KS G L L+
Sbjct: 231 VQILVTFFISNILNWDSLAQGLGSIIGSLGLLVESLLPLPQIAILYKLKSVQGFKLILLV 290
Query: 71 TWVAGDICNLVGCLLEPATLPTQFYTALTVINCSFMQALQSY 112
+W+ GD + + + +AL VI F +L Y
Sbjct: 291 SWLCGDTLKITYLIFGAKNI-----SALFVIFALFQMSLDFY 327
>YD03_SCHPO (Q10227) Hypothetical protein C2E12.03c in chromosome
I
Length = 283
Score = 38.5 bits (88), Expect = 0.030
Identities = 17/40 (42%), Positives = 24/40 (59%)
Query: 38 LMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDI 77
++ V W V IPQII +R KS+ G+ F+L+WV I
Sbjct: 24 ILGTVCWCVQLIPQIIKNYRAKSTEGLDTLFILSWVVASI 63
>CTNS_DROME (Q9VCR7) Cystinosin homolog
Length = 397
Score = 37.4 bits (85), Expect = 0.066
Identities = 17/43 (39%), Positives = 23/43 (52%)
Query: 291 GWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSY 333
GW+ ++ S PQIW N +R SVEGLN ++ T Y
Sbjct: 136 GWVYFVAWSVSFYPQIWSNYRRKSVEGLNFDFLALNIVGFTLY 178
Score = 33.1 bits (74), Expect = 1.2
Identities = 31/108 (28%), Positives = 50/108 (45%), Gaps = 25/108 (23%)
Query: 10 NKECVKWVETYFKDCLCNLRDDISFSL--GLMSLVSWGVAEIPQIITIFRNKSSHGISLA 67
NKE VK + + + + R I S+ G + V+W V+ PQI + +R KS G++
Sbjct: 109 NKEIVK--DVFVRVTVAKSRALIYTSIIFGWVYFVAWSVSFYPQIWSNYRRKSVEGLNFD 166
Query: 68 FLLTWVAGDICNLVGCLLEPATLPTQFYTALTVINCS--FMQALQSYY 113
FL N+VG +T ++ NC F++ LQ+ Y
Sbjct: 167 FL-------ALNIVG------------FTLYSMFNCGLYFIEDLQNEY 195
Score = 31.6 bits (70), Expect = 3.6
Identities = 32/137 (23%), Positives = 57/137 (41%), Gaps = 17/137 (12%)
Query: 264 RYGYIGFTGIKLLKVYVVVHSTY--GQYLGWIMAAIYTCSRI----------PQIWLNIK 311
R +I + + + V VVV + G + W+ +Y CS + PQ +N +
Sbjct: 237 RVSFIAYGILAIFAVVVVVSAGLAGGSVIHWL-DFLYYCSYVKLTITIIKYVPQALMNYR 295
Query: 312 RGSVEGLNPFMFVFALIANTSYVGSILVRTTEFE---SIKANLPWLLDATVCVALD-FFI 367
R S G + + T + +++ ++ SI + V D FF+
Sbjct: 296 RKSTSGWSIGNILLDFTGGTLSMLQMILNAHNYDDWVSIFGDPTKFGLGLFSVLFDVFFM 355
Query: 368 ISQYIYYRYFRSSESSD 384
+ Y++YR+ R S SSD
Sbjct: 356 LQHYVFYRHSRESSSSD 372
>ERS1_YEAST (P17261) Transmembrane protein ERS1 (ERD suppressor)
Length = 260
Score = 37.0 bits (84), Expect = 0.086
Identities = 19/46 (41%), Positives = 30/46 (64%), Gaps = 3/46 (6%)
Query: 30 DDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAG 75
DDI LG++ + SW ++ P IIT +R+KS+ IS+ F++ AG
Sbjct: 5 DDI---LGIVYVTSWSISMYPPIITNWRHKSASAISMDFVMLNTAG 47
>CTNS_ARATH (P57758) Cystinosin homolog
Length = 270
Score = 35.0 bits (79), Expect = 0.33
Identities = 18/47 (38%), Positives = 28/47 (59%)
Query: 288 QYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYV 334
+ +GWI A ++ S PQ+ LN +R SV GLN + L ++SY+
Sbjct: 14 EIVGWIAFASWSISFYPQLILNFRRRSVVGLNFDFVMLNLTKHSSYM 60
>MPU1_ARATH (Q9LTI3) Mannose-P-dolichol utilization defect 1
protein homolog
Length = 239
Score = 32.3 bits (72), Expect = 2.1
Identities = 23/63 (36%), Positives = 31/63 (48%), Gaps = 3/63 (4%)
Query: 22 KDCLCNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLV 81
KDCL L IS LG + + ++PQI+ I NKS G+S+ V G +L
Sbjct: 23 KDCLLPL---ISKLLGYFLVAASMTVKLPQIMKIVDNKSVKGLSVVAFELEVIGYTISLA 79
Query: 82 GCL 84
CL
Sbjct: 80 YCL 82
Score = 32.0 bits (71), Expect = 2.8
Identities = 25/79 (31%), Positives = 36/79 (44%), Gaps = 7/79 (8%)
Query: 297 IYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESIKANLPWLLD 356
I+ +RIPQIW N + S L+ F+ L+ G L R KA L LL
Sbjct: 153 IFLSARIPQIWKNFRNKSTGQLS---FLTCLM----NFGGALARVFTSIQEKAPLSMLLG 205
Query: 357 ATVCVALDFFIISQYIYYR 375
+ + + I+SQ + YR
Sbjct: 206 IVLSIFTNGIIMSQILLYR 224
>CTNS_CAEEL (Q09500) Cystinosin homolog
Length = 404
Score = 32.3 bits (72), Expect = 2.1
Identities = 15/32 (46%), Positives = 20/32 (61%)
Query: 288 QYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLN 319
Q +GW ++ S PQ++LN KR SV GLN
Sbjct: 130 QIVGWTYFFAWSISFYPQMYLNFKRKSVVGLN 161
>CTNS_MOUSE (P57757) Cystinosin
Length = 367
Score = 31.6 bits (70), Expect = 3.6
Identities = 20/62 (32%), Positives = 32/62 (51%), Gaps = 3/62 (4%)
Query: 273 IKLLKVYVVVHSTYGQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTS 332
I+ L ++ + S Q +GWI ++ S PQ+ N +R SV GL+ F F + T
Sbjct: 113 IRFLVIHSRIVSIINQVIGWIYFMAWSVSFYPQVIQNWRRKSVIGLS---FDFLALNLTG 169
Query: 333 YV 334
+V
Sbjct: 170 FV 171
>CTNS_HUMAN (O60931) Cystinosin
Length = 367
Score = 31.6 bits (70), Expect = 3.6
Identities = 20/62 (32%), Positives = 31/62 (49%), Gaps = 3/62 (4%)
Query: 273 IKLLKVYVVVHSTYGQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTS 332
I+ L + S Q +GWI ++ S PQ+ +N +R SV GL+ F F + T
Sbjct: 113 IRFLVIRSSAISIINQVIGWIYFVAWSISFYPQVIMNWRRKSVIGLS---FDFVALNLTG 169
Query: 333 YV 334
+V
Sbjct: 170 FV 171
>HUTH_STAAN (P64416) Histidine ammonia-lyase (EC 4.3.1.3)
(Histidase)
Length = 504
Score = 30.8 bits (68), Expect = 6.2
Identities = 17/46 (36%), Positives = 26/46 (55%), Gaps = 2/46 (4%)
Query: 146 QDNEEEKRPLNPKPS--QVYSGIAIPNGTQKEAARGEYYYMSARSL 189
+D+++ R LN +P Q G+A+ NGTQ A+G Y+ A L
Sbjct: 169 KDSDDVLRELNRQPLNLQAKEGLALINGTQAMTAQGVISYIEAEDL 214
>HUTH_STAAM (P64415) Histidine ammonia-lyase (EC 4.3.1.3)
(Histidase)
Length = 504
Score = 30.8 bits (68), Expect = 6.2
Identities = 17/46 (36%), Positives = 26/46 (55%), Gaps = 2/46 (4%)
Query: 146 QDNEEEKRPLNPKPS--QVYSGIAIPNGTQKEAARGEYYYMSARSL 189
+D+++ R LN +P Q G+A+ NGTQ A+G Y+ A L
Sbjct: 169 KDSDDVLRELNRQPLNLQAKEGLALINGTQAMTAQGVISYIEAEDL 214
>Y041_UREPA (Q9PRA4) Hypothetical protein UU041
Length = 120
Score = 30.4 bits (67), Expect = 8.1
Identities = 12/36 (33%), Positives = 22/36 (60%)
Query: 36 LGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLT 71
+G+ + + IPQ+I + + K + GISL FL++
Sbjct: 11 IGIFGSIFLVIGYIPQVIKVIKTKRTDGISLTFLIS 46
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.137 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 46,083,569
Number of Sequences: 164201
Number of extensions: 1885370
Number of successful extensions: 3967
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 3935
Number of HSP's gapped (non-prelim): 33
length of query: 390
length of database: 59,974,054
effective HSP length: 112
effective length of query: 278
effective length of database: 41,583,542
effective search space: 11560224676
effective search space used: 11560224676
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC148241.11


