

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
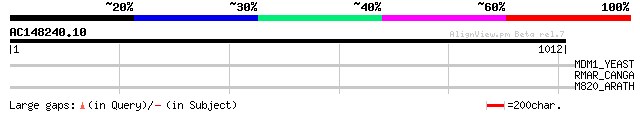
Query= AC148240.10 - phase: 0 /pseudo
(1012 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
MDM1_YEAST (Q01846) Structural protein MDM1 34 1.7
RMAR_CANGA (P21358) Mitochondrial ribosomal protein VAR1 32 6.6
M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820... 32 6.6
>MDM1_YEAST (Q01846) Structural protein MDM1
Length = 1127
Score = 34.3 bits (77), Expect = 1.7
Identities = 32/92 (34%), Positives = 47/92 (50%), Gaps = 9/92 (9%)
Query: 421 IESLEIKLQSKVKVMQV--QLILK--MHQNLINLLILKSIQKLNLVQKLKSLQKLSLIQK 476
I SL I + K +++ LILK + N + L ILK Q+ LK L+ L+++
Sbjct: 712 IASLTISIDQIEKELELLRHLILKADLTNNQMQLKILKKSQRT----LLKELEMKELLKQ 767
Query: 477 QKLVQ*NRMKMLLRMFKITLSRLFSQNSNTNL 508
Q +VQ N L R KI + FS+NS+ L
Sbjct: 768 QYMVQENG-NSLFRKTKIYIRSYFSENSSNGL 798
>RMAR_CANGA (P21358) Mitochondrial ribosomal protein VAR1
Length = 339
Score = 32.3 bits (72), Expect = 6.6
Identities = 31/111 (27%), Positives = 57/111 (50%), Gaps = 13/111 (11%)
Query: 605 KPDLLLKVTVNKKALI-------TLKHLLQLQDWKQSGYFYPMQLTMA*YC-IKWMSKVL 656
K L+ K++ N K L +L+HL + +WK Y Y T+ Y K ++K+L
Sbjct: 30 KNRLMNKLSNNNKYLSELNNKGNSLQHLNNMNNWKLQNYNYNKNNTINNYINSKLINKLL 89
Query: 657 FLMVSLK--KKYMSNNL--LGLRILSILTMFINLRNHYMA*NKLPELGMID 703
+ ++SLK K +S L + + +++I + N+ N+Y N + + MI+
Sbjct: 90 YKLMSLKNNKIIISKPLYKINMNVINIRFYYYNM-NNYNYNNNIYYINMIN 139
>M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820
(ORF170)
Length = 170
Score = 32.3 bits (72), Expect = 6.6
Identities = 18/36 (50%), Positives = 22/36 (61%)
Query: 649 IKWMSKVLFLMVSLKKKYMSNNLLGLRILSILTMFI 684
I WM K+ F M KKK++ NLL LRIL I M +
Sbjct: 134 INWMFKMHFSMGIFKKKFICINLLVLRILFIHPMCV 169
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.359 0.160 0.523
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 85,891,376
Number of Sequences: 164201
Number of extensions: 2836281
Number of successful extensions: 10229
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 10227
Number of HSP's gapped (non-prelim): 3
length of query: 1012
length of database: 59,974,054
effective HSP length: 120
effective length of query: 892
effective length of database: 40,269,934
effective search space: 35920781128
effective search space used: 35920781128
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 71 (32.0 bits)
Medicago: description of AC148240.10


