

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148236.3 + phase: 0 /pseudo
(116 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
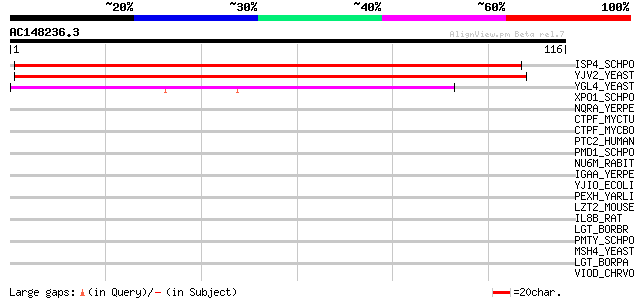
Score E
Sequences producing significant alignments: (bits) Value
ISP4_SCHPO (P40900) Sexual differentiation process protein isp4 112 2e-25
YJV2_YEAST (P40897) Hypothetical 91.6 kDa protein in HXT8-CRT1 i... 105 2e-23
YGL4_YEAST (P53134) Hypothetical 80.0 kDa protein in SNF4-TAF60 ... 41 6e-04
XPO1_SCHPO (P14068) Exportin 1 (Chromosome region maintenance pr... 31 0.44
NQRA_YERPE (Q8ZBZ0) Na(+)-translocating NADH-quinone reductase s... 30 0.75
CTPF_MYCTU (P63687) Probable cation-transporting ATPase F (EC 3.... 30 1.3
CTPF_MYCBO (P63688) Probable cation-transporting ATPase F (EC 3.... 30 1.3
PTC2_HUMAN (Q9Y6C5) Patched protein homolog 2 (PTC2) 29 1.7
PMD1_SCHPO (P36619) Leptomycin B resistance protein pmd1 29 2.2
NU6M_RABIT (O79438) NADH-ubiquinone oxidoreductase chain 6 (EC 1... 29 2.2
IGAA_YERPE (P58722) Putative membrane protein igaA homolog 29 2.2
YJIO_ECOLI (P39386) Hypothetical transport protein yjiO 28 2.9
PEXH_YARLI (P87200) Peroxisomal membrane protein PEX17 (Peroxin-17) 28 3.7
LZT2_MOUSE (Q91YU6) Leucine zipper putative tumor suppressor 2 28 3.7
IL8B_RAT (P35407) High affinity interleukin-8 receptor B (IL-8R ... 28 3.7
LGT_BORBR (Q7WFQ4) Prolipoprotein diacylglyceryl transferase (EC... 28 4.9
PMTY_SCHPO (O42933) Probable dolichyl-phosphate-mannose--protein... 27 6.4
MSH4_YEAST (P40965) MUTS protein homolog 4 27 6.4
LGT_BORPA (Q7W496) Prolipoprotein diacylglyceryl transferase (EC... 27 6.4
VIOD_CHRVO (Q9S3U8) Probable tryptophan hydroxylase vioD (EC 1.-... 27 8.3
>ISP4_SCHPO (P40900) Sexual differentiation process protein isp4
Length = 785
Score = 112 bits (279), Expect = 2e-25
Identities = 48/106 (45%), Positives = 72/106 (67%)
Query: 2 PWWALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGYM 61
PWW +I + ++ +P+ I+ A TN GLNV TE+I+G + PGRP+A + FKT GY+
Sbjct: 489 PWWVIIVGVIFSAVWFIPIGIVQAITNIQLGLNVFTEFIVGYMYPGRPLAMMIFKTVGYI 548
Query: 62 SMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWW 107
+M+Q ++F +D K GHYMK+PPR MF Q++ T+ + V +GV W
Sbjct: 549 TMTQGLAFAADLKFGHYMKLPPRIMFYTQMIATIWSCFVQIGVLDW 594
>YJV2_YEAST (P40897) Hypothetical 91.6 kDa protein in HXT8-CRT1
intergenic region
Length = 799
Score = 105 bits (262), Expect = 2e-23
Identities = 51/107 (47%), Positives = 71/107 (65%)
Query: 2 PWWALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGYM 61
P WA + A ++L+ +P I+ A TNQ GLN+ITE I G +LP RP+AN+ FK YG++
Sbjct: 508 PAWAFVIAILISLVNFIPQGILEAMTNQHVGLNIITELICGYMLPLRPMANLLFKLYGFI 567
Query: 62 SMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
M Q ++ D KL YMK+ PR +F VQI T+I+G V+VGV W+
Sbjct: 568 VMRQGLNLSRDLKLAMYMKVSPRLIFAVQIYATIISGMVNVGVQEWM 614
>YGL4_YEAST (P53134) Hypothetical 80.0 kDa protein in SNF4-TAF60
intergenic region
Length = 725
Score = 40.8 bits (94), Expect = 6e-04
Identities = 29/99 (29%), Positives = 45/99 (45%), Gaps = 6/99 (6%)
Query: 1 MPWWALIFASGLALIFTLPVAIITATTNQTP--GLNVITEYIMGVILP----GRPIANVC 54
+P +A+I A LAL ++ T+ P G+ I++ I I+P G + NV
Sbjct: 472 IPLYAIITALILALFLSILGIRALGETDLNPVSGIGKISQLIFAFIIPRDRPGSVLMNVV 531
Query: 55 FKTYGYMSMSQAISFLSDFKLGHYMKIPPRSMFIVQILG 93
S QA + D K GH + PR+ F Q++G
Sbjct: 532 SGGIAEASAQQAGDLMQDLKTGHLLGASPRAQFCAQLIG 570
>XPO1_SCHPO (P14068) Exportin 1 (Chromosome region maintenance
protein 1) (Caffeine resistance protein 2)
Length = 1078
Score = 31.2 bits (69), Expect = 0.44
Identities = 22/67 (32%), Positives = 37/67 (54%), Gaps = 6/67 (8%)
Query: 29 QTPGLNVITEYIMGVILPGRPIANVCFKTYGYMSMSQAISFL---SDFKLGHYMK--IPP 83
Q GLN++ E I + G ++N F+TY Y+S+ Q I ++ SD K G ++ I
Sbjct: 891 QETGLNILLELINNMASMGPDVSNAFFQTY-YISLLQDILYVLTDSDHKSGFKLQSLILA 949
Query: 84 RSMFIVQ 90
R ++V+
Sbjct: 950 RLFYLVE 956
>NQRA_YERPE (Q8ZBZ0) Na(+)-translocating NADH-quinone reductase
subunit A (EC 1.6.5.-) (Na(+)-translocating NQR subunit
A) (Na(+)-NQR subunit A) (NQR complex subunit A) (NQR-1
subunit A)
Length = 447
Score = 30.4 bits (67), Expect = 0.75
Identities = 16/70 (22%), Positives = 32/70 (44%), Gaps = 8/70 (11%)
Query: 40 IMGVILPGRPIANVCFKTYGYMSMSQAISFLSDFK--------LGHYMKIPPRSMFIVQI 91
+ G ++PGR ++ T G+ + +F +D +G+Y ++ P + +
Sbjct: 335 LFGWVMPGRDKYSITRTTLGHFFKRKLFAFSTDMHGGERAMVPIGNYERVMPLDILATHL 394
Query: 92 LGTLIAGTVD 101
L L+AG D
Sbjct: 395 LRDLLAGDTD 404
>CTPF_MYCTU (P63687) Probable cation-transporting ATPase F (EC
3.6.3.-)
Length = 905
Score = 29.6 bits (65), Expect = 1.3
Identities = 25/90 (27%), Positives = 38/90 (41%), Gaps = 7/90 (7%)
Query: 24 TATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGYMSMSQAISFLSDFKLGHYMKIPP 83
T TN GL ++ +GV LP P + +++ ++F + K M PP
Sbjct: 705 TLPTNLGEGLVILAAIAVGVALPILPTQILWINMTTAIALGLMLAF--EPKEAGIMTRPP 762
Query: 84 RSMFIVQILG-----TLIAGTVDVGVAWWL 108
R + G TL+ T+ V AWWL
Sbjct: 763 RDPDQPLLTGWLVRRTLLVSTLLVASAWWL 792
>CTPF_MYCBO (P63688) Probable cation-transporting ATPase F (EC
3.6.3.-)
Length = 905
Score = 29.6 bits (65), Expect = 1.3
Identities = 25/90 (27%), Positives = 38/90 (41%), Gaps = 7/90 (7%)
Query: 24 TATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGYMSMSQAISFLSDFKLGHYMKIPP 83
T TN GL ++ +GV LP P + +++ ++F + K M PP
Sbjct: 705 TLPTNLGEGLVILAAIAVGVALPILPTQILWINMTTAIALGLMLAF--EPKEAGIMTRPP 762
Query: 84 RSMFIVQILG-----TLIAGTVDVGVAWWL 108
R + G TL+ T+ V AWWL
Sbjct: 763 RDPDQPLLTGWLVRRTLLVSTLLVASAWWL 792
>PTC2_HUMAN (Q9Y6C5) Patched protein homolog 2 (PTC2)
Length = 1203
Score = 29.3 bits (64), Expect = 1.7
Identities = 30/116 (25%), Positives = 47/116 (39%), Gaps = 13/116 (11%)
Query: 5 ALIFASGLALIFTLPVAIITATTNQTP----GLNVITEYIMG----VILPGRPI---ANV 53
AL ASGL L L + ATT P G+ V +++ LPG P+
Sbjct: 434 ALAVASGLGLCALLGITFNAATTQVLPFLALGIGVDDVFLLAHAFTEALPGTPLQERMGE 493
Query: 54 CFKTYGYMSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWLP 109
C + G + +I+ ++ F + + IP F +Q ++ G V V P
Sbjct: 494 CLQRTGTSVVLTSINNMAAFLMAALVPIPALRAFSLQ--AAIVVGCTFVAVMLVFP 547
>PMD1_SCHPO (P36619) Leptomycin B resistance protein pmd1
Length = 1362
Score = 28.9 bits (63), Expect = 2.2
Identities = 21/75 (28%), Positives = 32/75 (42%), Gaps = 10/75 (13%)
Query: 34 NVITEYIMGVILPGRPIANVCFKTYGYMSMSQAISFLSD----------FKLGHYMKIPP 83
NV EY +I PGR A K+ + S +Q ++FL + + G Y +
Sbjct: 998 NVFAEYCDSLIKPGRESAIASLKSGLFFSAAQGVTFLINALTFWYGSTLMRKGEYNIVQF 1057
Query: 84 RSMFIVQILGTLIAG 98
+ FI + G AG
Sbjct: 1058 YTCFIAIVFGIQQAG 1072
>NU6M_RABIT (O79438) NADH-ubiquinone oxidoreductase chain 6 (EC
1.6.5.3)
Length = 174
Score = 28.9 bits (63), Expect = 2.2
Identities = 19/71 (26%), Positives = 34/71 (47%), Gaps = 10/71 (14%)
Query: 6 LIFASGLALIFTLPVAIITATTNQTPGLNV--ITEYIMGVILPGRPIANVCFKTYGYMSM 63
LI+ G+ ++F A+ T +T G NV + +++GV++ + YM M
Sbjct: 57 LIYLGGMLVVFGYTTAMATEEYPETWGSNVMILGMFVLGVLMEVGLVV--------YMVM 108
Query: 64 SQAISFLSDFK 74
S + + DFK
Sbjct: 109 SDGVEIVVDFK 119
>IGAA_YERPE (P58722) Putative membrane protein igaA homolog
Length = 715
Score = 28.9 bits (63), Expect = 2.2
Identities = 13/28 (46%), Positives = 17/28 (60%), Gaps = 1/28 (3%)
Query: 88 IVQILGTLIAGTVDVGVAWWLPRFNKEH 115
IV IL L+ + VG+ WWL RF + H
Sbjct: 4 IVLILALLLTSLIAVGLLWWL-RFRRPH 30
>YJIO_ECOLI (P39386) Hypothetical transport protein yjiO
Length = 410
Score = 28.5 bits (62), Expect = 2.9
Identities = 28/84 (33%), Positives = 37/84 (43%), Gaps = 14/84 (16%)
Query: 13 ALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCF-KTYGYMSMSQAISFLS 71
ALIFTL A TT+ T L I I G ++CF T GY+++ +A
Sbjct: 85 ALIFTLACAATMFTTSMTQFL--IARAIQG--------TSICFIATVGYVTVQEAFGQTK 134
Query: 72 DFKLGHYMKIPPRSMFIVQILGTL 95
KL M I + I I+G L
Sbjct: 135 GIKL---MAIITSIVLIAPIIGPL 155
>PEXH_YARLI (P87200) Peroxisomal membrane protein PEX17 (Peroxin-17)
Length = 671
Score = 28.1 bits (61), Expect = 3.7
Identities = 18/53 (33%), Positives = 26/53 (48%), Gaps = 6/53 (11%)
Query: 55 FKTYGYMSMSQAISFLSDFKLGHYMK----IPPRSMFIVQILGTLIAGTVDVG 103
F Y Y+ S LS++ Y+K IPP F+ + L +AGT +VG
Sbjct: 372 FAAYDYVFFSAIDVLLSEY--APYIKNRGTIPPNKEFVAERLAANLAGTSNVG 422
>LZT2_MOUSE (Q91YU6) Leucine zipper putative tumor suppressor 2
Length = 671
Score = 28.1 bits (61), Expect = 3.7
Identities = 15/42 (35%), Positives = 23/42 (54%), Gaps = 4/42 (9%)
Query: 12 LALIFTLPVAII----TATTNQTPGLNVITEYIMGVILPGRP 49
+A++ TLPV + TAT +TP + ++ I G PG P
Sbjct: 1 MAIVHTLPVPLEPARETATAPKTPAMGSVSSLISGRPCPGGP 42
>IL8B_RAT (P35407) High affinity interleukin-8 receptor B (IL-8R B)
(CXCR-2) (GRO/MGSA receptor)
Length = 359
Score = 28.1 bits (61), Expect = 3.7
Identities = 17/64 (26%), Positives = 28/64 (43%)
Query: 12 LALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGYMSMSQAISFLS 71
L+L+ +LP+ I+ T P V E I R + + +TYG++ + F
Sbjct: 171 LSLVLSLPIFILRTTVKANPSTVVCYENIGNNTSKWRVVLRILPQTYGFLLPLLIMLFCY 230
Query: 72 DFKL 75
F L
Sbjct: 231 GFTL 234
>LGT_BORBR (Q7WFQ4) Prolipoprotein diacylglyceryl transferase (EC
2.4.99.-)
Length = 262
Score = 27.7 bits (60), Expect = 4.9
Identities = 21/72 (29%), Positives = 38/72 (52%), Gaps = 4/72 (5%)
Query: 3 WWALIFASGLALIFTLPVAIITA--TTNQTPG--LNVITEYIMGVILPGRPIANVCFKTY 58
W+ L++ G AL++ L IT+ TT+ T ++I ++GV+L GR + +K
Sbjct: 21 WYGLMYLIGFALVYALGRRRITSGHTTSMTVRDLEDLIFYSVLGVVLGGRLGYVLFYKPA 80
Query: 59 GYMSMSQAISFL 70
Y++ I +L
Sbjct: 81 HYLANPLEIFYL 92
>PMTY_SCHPO (O42933) Probable dolichyl-phosphate-mannose--protein
mannosyltransferase C16C6.09 (EC 2.4.1.109)
Length = 778
Score = 27.3 bits (59), Expect = 6.4
Identities = 13/37 (35%), Positives = 19/37 (51%)
Query: 2 PWWALIFASGLALIFTLPVAIITATTNQTPGLNVITE 38
PWWA +F +G L T+ + T + GL+V E
Sbjct: 227 PWWAWLFFTGFFLSCTISTKYVGFFTFLSIGLSVCLE 263
>MSH4_YEAST (P40965) MUTS protein homolog 4
Length = 878
Score = 27.3 bits (59), Expect = 6.4
Identities = 14/45 (31%), Positives = 22/45 (48%)
Query: 28 NQTPGLNVITEYIMGVILPGRPIANVCFKTYGYMSMSQAISFLSD 72
N GL IT+Y+M I + KT+ + S AIS++ +
Sbjct: 195 NSQDGLAAITKYLMDDTKKDLKIEEIIDKTFALCAASAAISYMEE 239
>LGT_BORPA (Q7W496) Prolipoprotein diacylglyceryl transferase (EC
2.4.99.-)
Length = 262
Score = 27.3 bits (59), Expect = 6.4
Identities = 21/72 (29%), Positives = 38/72 (52%), Gaps = 4/72 (5%)
Query: 3 WWALIFASGLALIFTLPVAIITA--TTNQTPG--LNVITEYIMGVILPGRPIANVCFKTY 58
W+ L++ G AL++ L IT+ TT+ T ++I ++GV+L GR + +K
Sbjct: 21 WYGLMYLIGFALVYALGRRRITSGHTTSMTVRDLEDLIFYSVLGVVLGGRLGYVLFYKPA 80
Query: 59 GYMSMSQAISFL 70
Y++ I +L
Sbjct: 81 YYLANPLEIFYL 92
>VIOD_CHRVO (Q9S3U8) Probable tryptophan hydroxylase vioD (EC
1.-.-.-)
Length = 373
Score = 26.9 bits (58), Expect = 8.3
Identities = 15/39 (38%), Positives = 22/39 (55%), Gaps = 3/39 (7%)
Query: 42 GVILPGRPIANVCFKTYGYMSMSQAIS--FLSDFKLGHY 78
GV+LPGRP + Y+ + ++ FL DFKL H+
Sbjct: 43 GVVLPGRPGQHPA-NPLSYLDAPERLNPQFLEDFKLVHH 80
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.328 0.142 0.459
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,385,880
Number of Sequences: 164201
Number of extensions: 475825
Number of successful extensions: 1424
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 1416
Number of HSP's gapped (non-prelim): 20
length of query: 116
length of database: 59,974,054
effective HSP length: 92
effective length of query: 24
effective length of database: 44,867,562
effective search space: 1076821488
effective search space used: 1076821488
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC148236.3


