

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148236.2 - phase: 0 /pseudo
(362 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
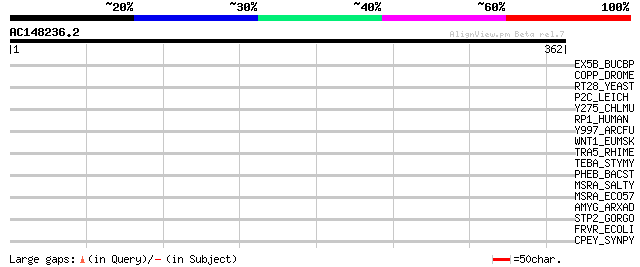
Score E
Sequences producing significant alignments: (bits) Value
EX5B_BUCBP (Q89AB3) Exodeoxyribonuclease V beta chain (EC 3.1.11.5) 36 0.17
COPP_DROME (O62621) Coatomer beta' subunit (Beta'-coat protein) ... 35 0.39
RT28_YEAST (P21771) 40S ribosomal protein S28, mitochondrial pre... 34 0.66
P2C_LEICH (P36982) Protein phosphatase 2C (EC 3.1.3.16) (PP2C) 34 0.66
Y275_CHLMU (Q9PL33) Hypothetical protein TC0275 31 4.3
RP1_HUMAN (P56715) Oxygen-regulated protein 1 (Retinitis pigment... 31 4.3
Y997_ARCFU (O29265) Hypothetical protein AF0997 31 5.6
WNT1_EUMSK (P28113) Wnt-1 protein (Fragment) 31 5.6
TRA5_RHIME (Q52873) Transposase for insertion sequence element I... 31 5.6
TEBA_STYMY (P29550) Telomere-binding protein alpha subunit 30 7.3
PHEB_BACST (P31003) Metapyrocatechase (EC 1.13.11.2) (MPC) (CatO... 30 7.3
MSRA_SALTY (Q8ZK71) Peptide methionine sulfoxide reductase msrA ... 30 7.3
MSRA_ECO57 (Q8XCG3) Peptide methionine sulfoxide reductase msrA ... 30 7.3
AMYG_ARXAD (P42042) Glucoamylase precursor (EC 3.2.1.3) (Glucan ... 30 7.3
STP2_GORGO (Q9N1A5) Nuclear transition protein 2 (TP-2) (TP2) (F... 30 9.5
FRVR_ECOLI (P32152) Putative frv operon regulatory protein 30 9.5
CPEY_SYNPY (Q02174) Bilin biosynthesis protein cpeY 30 9.5
>EX5B_BUCBP (Q89AB3) Exodeoxyribonuclease V beta chain (EC 3.1.11.5)
Length = 1180
Score = 35.8 bits (81), Expect = 0.17
Identities = 24/87 (27%), Positives = 40/87 (45%), Gaps = 6/87 (6%)
Query: 143 FDLLSKEVGGRTNLGFIRLDQKNYLRKKR-----ERSLVQGEAGYLLQYFQRKTVENPTF 197
+ +L K + NL +LD KNY+++ + ++ L + + Q Q ++EN
Sbjct: 1003 YSILYKNLDKNYNLRLSKLDSKNYIKELKFLLPIKKKLTALKLNNIFQTHQCSSLENKLC 1062
Query: 198 YHAYQLDIEDQITNVF-WADARMLVDY 223
+H Q + I VF W LVDY
Sbjct: 1063 FHPIQGILSGSIDLVFLWKTKYFLVDY 1089
>COPP_DROME (O62621) Coatomer beta' subunit (Beta'-coat protein)
(Beta'-COP)
Length = 914
Score = 34.7 bits (78), Expect = 0.39
Identities = 37/113 (32%), Positives = 50/113 (43%), Gaps = 11/113 (9%)
Query: 136 GVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQYFQRKTVENP 195
G Q+ + L S E R N G +RL +N+ +K E+ Y YF KT
Sbjct: 391 GSAQEFVWALESNEYAIRENNGTVRLF-RNFKERKSFTPEYGAESIYGGYYFGVKTSSGL 449
Query: 196 TFYHAYQLD----IEDQITNVFWADARMLV----DYSYFGDVVSLDSTYCTNS 240
FY L IE Q NVFW ++ LV D SYF V+ +D+ N+
Sbjct: 450 AFYDWETLQLVRRIEVQPKNVFWNESGSLVCLATDDSYF--VLGVDTAQVANA 500
>RT28_YEAST (P21771) 40S ribosomal protein S28, mitochondrial
precursor
Length = 286
Score = 33.9 bits (76), Expect = 0.66
Identities = 27/91 (29%), Positives = 46/91 (49%), Gaps = 8/91 (8%)
Query: 159 IRLDQKNYLRKKRERSLVQGEAGYLLQYFQRKTVENPTFYH--AYQLDIEDQ-ITNVFWA 215
I+ +K++ + R LVQ +L+Y +R +NP Y+ +L + D IT+ F
Sbjct: 192 IKEHRKDFANTRNLRILVQQRQA-ILRYLKR---DNPEKYYWTIQKLGLNDAAITDEFNM 247
Query: 216 DARMLVDYSYFGDVVSL-DSTYCTNSSHRPL 245
D R + DY +FGD + + DS N + +
Sbjct: 248 DRRYMQDYEFFGDKILIRDSKKVANQKRKEI 278
>P2C_LEICH (P36982) Protein phosphatase 2C (EC 3.1.3.16) (PP2C)
Length = 406
Score = 33.9 bits (76), Expect = 0.66
Identities = 21/63 (33%), Positives = 28/63 (44%), Gaps = 1/63 (1%)
Query: 45 GSVSSCRFVCCKEGMRKPDKRDYKTKNPRLETR-TNCEARLGLKNVDGKLMVHDFVEDHN 103
G+V R V C +G+ P D+K N R NC R+ VDG L V D
Sbjct: 125 GNVGDSRVVACIDGVCVPLTEDHKPNNEGERQRIENCAGRVENNRVDGSLAVSRAFGDRE 184
Query: 104 HEL 106
++L
Sbjct: 185 YKL 187
>Y275_CHLMU (Q9PL33) Hypothetical protein TC0275
Length = 316
Score = 31.2 bits (69), Expect = 4.3
Identities = 35/159 (22%), Positives = 64/159 (40%), Gaps = 6/159 (3%)
Query: 84 LGLKNVDGKLMVHDFVEDHNHELLLPETTHMLSSQRKVSEIHCQQIELADSVGVQQKKSF 143
L L N ++ V+ N E+ +LSS+ SE I + + VG+ K+++
Sbjct: 135 LALTNWQWSVLFSGMVDPENIEMGSGVYQVVLSSKYHASEAFAVVIGVINEVGLHDKQAW 194
Query: 144 DLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQYFQRKTVENPT-----FY 198
LL L + N+ + + S+ A Y L F++K +N +
Sbjct: 195 PLLGFSYKPEDKLTLNCIYPVNFSAEYQCTSVCDLGAAYRLTRFRKKFPKNSLTTSEGIF 254
Query: 199 HAYQLDIEDQITNVFWADARMLVDYSY-FGDVVSLDSTY 236
+IE + +FW + V Y G+ +SL + +
Sbjct: 255 EYSGREIEGSVKLIFWPGQSLKVFGGYSVGNDISLTNPH 293
>RP1_HUMAN (P56715) Oxygen-regulated protein 1 (Retinitis pigmentosa
RP1 protein) (Retinitis pigmentosa 1 protein)
Length = 2156
Score = 31.2 bits (69), Expect = 4.3
Identities = 34/138 (24%), Positives = 59/138 (42%), Gaps = 15/138 (10%)
Query: 46 SVSSCRFVCCKEGMRKPDKRDYKTKNPRLETRTNCEARLGLKNVDGKLMVHDFVEDHNHE 105
++SSC +C E + DK K+ L+ NC R + D E E
Sbjct: 1395 NLSSCG-LCLSEKEAELDK-----KHSSLDDFENCSLRKFQD--ENAYTSFDMEEPRTSE 1446
Query: 106 LLLPETTHMLSSQRKVSEIHC-QQIELADS------VGVQQKKSFDLLSKEVGGRTNLGF 158
T M SS+R +SE+ +++E D+ V ++ + +L+ +EV L
Sbjct: 1447 EPGSITNSMTSSERNISELESFEELENHDTDIFNTVVNGGEQATEELIQEEVEASKTLEL 1506
Query: 159 IRLDQKNYLRKKRERSLV 176
I + KN + +KR ++
Sbjct: 1507 IDISSKNIMEEKRMNGII 1524
>Y997_ARCFU (O29265) Hypothetical protein AF0997
Length = 416
Score = 30.8 bits (68), Expect = 5.6
Identities = 37/164 (22%), Positives = 59/164 (35%), Gaps = 14/164 (8%)
Query: 104 HELLLPETTHMLSSQRKVSEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQ 163
H + L E + ++R VS HC L S G+ + + VG T+ G +
Sbjct: 242 HAVWLSEAEMKILAERGVSVAHCPTSNLKLSSGIAKVSELLEMGVNVGIGTD-GAASNNM 300
Query: 164 KNYLRKKRERSLVQGEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDY 223
+ L R +L+Q G L+ + Y AY L ++
Sbjct: 301 LSVLSDARVGALLQNLRGRTLKPGHWLEMATEGGYRAYNLK-------------GGRIEE 347
Query: 224 SYFGDVVSLDSTYCTNSSHRPLAIISGFNHHRGAVIFGAALLYD 267
Y D+V T H P A++ N AV+ G ++ D
Sbjct: 348 GYLADIVVFSKTCRNAPMHDPAAMLYVENQALHAVVDGVLVMED 391
>WNT1_EUMSK (P28113) Wnt-1 protein (Fragment)
Length = 126
Score = 30.8 bits (68), Expect = 5.6
Identities = 15/56 (26%), Positives = 22/56 (38%), Gaps = 8/56 (14%)
Query: 25 TYXGKXGFGVRKDYTHRKKDGSVSSCRFVCCKEGMRKPDKRDYKTKNPRLETRTNC 80
TY GK G + ++ C +CC G Y+T+ R+ R NC
Sbjct: 77 TYSGKTGTAGTAGRSCNSSSPALDGCELLCCGRG--------YRTRTQRVTERCNC 124
>TRA5_RHIME (Q52873) Transposase for insertion sequence element
ISRM5
Length = 398
Score = 30.8 bits (68), Expect = 5.6
Identities = 15/48 (31%), Positives = 24/48 (49%)
Query: 281 LEAHKQKMPQTVFTDQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHL 328
L+ K + V +D + A+ EV+PEA C H ++N + HL
Sbjct: 216 LKGRGLKGVELVVSDDHAGLVAAIGEVIPEAAWQRCYVHFLRNALDHL 263
>TEBA_STYMY (P29550) Telomere-binding protein alpha subunit
Length = 493
Score = 30.4 bits (67), Expect = 7.3
Identities = 34/148 (22%), Positives = 64/148 (42%), Gaps = 35/148 (23%)
Query: 148 KEVGGR---TNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQYFQRKTVENPTFY------ 198
+E+GG+ ++L KNY +K E +L+Q + +QYFQ+ V + +
Sbjct: 155 QEIGGQEPASDLTPFAFSGKNYTFEKSEAALLQNIRKWAVQYFQQYNVISSDMFTPLNKA 214
Query: 199 ---------------------HAYQLDIEDQITNVFWADA-RMLVDYSYFGDVVSLDS-T 235
+ +L ++DQ VF+ A ++ + G+VV + S T
Sbjct: 215 QAQKGDFDVVAKILQVFELDEYTNELKLKDQSGQVFYTLALKLKFPHLRAGEVVRIRSAT 274
Query: 236 YCTNSSHRPLAIISGFNHHRGAVIFGAA 263
Y S+ + + ++S H+ V F +A
Sbjct: 275 YDETSTQKKVLLLS---HYSNIVTFVSA 299
>PHEB_BACST (P31003) Metapyrocatechase (EC 1.13.11.2) (MPC)
(CatO2ase) (Catechol 2,3-dioxygenase)
Length = 327
Score = 30.4 bits (67), Expect = 7.3
Identities = 18/60 (30%), Positives = 26/60 (43%), Gaps = 12/60 (20%)
Query: 313 HGLCTWHLMQNGIKHLGNLMKGESYFLS--------DFEKCMYGYED----VEQFEDVGH 360
H LC W+ + + L +L+K YF+ CMY YE +E F D G+
Sbjct: 220 HHLCYWYGIPQNLYDLADLLKDHEYFIEVPPNKHGISQAFCMYVYEPGGNRIELFGDAGY 279
>MSRA_SALTY (Q8ZK71) Peptide methionine sulfoxide reductase msrA (EC
1.8.4.6) (Protein-methionine-S-oxide reductase) (Peptide
Met(O) reductase)
Length = 212
Score = 30.4 bits (67), Expect = 7.3
Identities = 22/75 (29%), Positives = 30/75 (39%), Gaps = 6/75 (8%)
Query: 236 YCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTD 295
Y N ++R + SG H AV ++YD SYE L +TF E H D
Sbjct: 77 YTPNPTYRE--VCSGQTGHAEAV----RIVYDPAVISYEQLLQTFWENHDPTQGMQQGND 130
Query: 296 QAKSMAKALAEVMPE 310
A+ + PE
Sbjct: 131 HGTQYRSAIYPLTPE 145
>MSRA_ECO57 (Q8XCG3) Peptide methionine sulfoxide reductase msrA (EC
1.8.4.6) (Protein-methionine-S-oxide reductase) (Peptide
Met(O) reductase)
Length = 211
Score = 30.4 bits (67), Expect = 7.3
Identities = 24/99 (24%), Positives = 36/99 (36%), Gaps = 7/99 (7%)
Query: 213 FWADARMLVD-YSYFGDVVSLDSTYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAE 271
FW R+ + + Y N ++R + SG H AV ++YD +
Sbjct: 52 FWGVERLFWQLHGVYSTAAGYTGGYTPNPTYRE--VCSGDTGHAEAV----RIVYDPSVI 105
Query: 272 SYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALAEVMPE 310
SYE L + F E H D A+ + PE
Sbjct: 106 SYEQLLQVFWENHDPAQGMRQGNDHGTQYRSAIYPLTPE 144
>AMYG_ARXAD (P42042) Glucoamylase precursor (EC 3.2.1.3) (Glucan
1,4-alpha-glucosidase) (1,4-alpha-D-glucan
glucohydrolase)
Length = 624
Score = 30.4 bits (67), Expect = 7.3
Identities = 32/148 (21%), Positives = 61/148 (40%), Gaps = 26/148 (17%)
Query: 213 FWADARMLVDYSYFGDVVSLDSTYCTNSSHRPLAIISGFNH-HRGAVIFGAALLYDETAE 271
FW AR L+ Y Y G V L Y S++ ++++ G H + +F + D+
Sbjct: 393 FWDSARQLILYEY-GPV--LRGKY----SYKDISVVLGVMHGYANDNVF--SYTNDQILA 443
Query: 272 SYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNL 331
+ + +FL+ +K + T + K + + + Y G+ T
Sbjct: 444 TAYQVSTSFLDVYK--VANTTSDESGKPLGIPVGRYPEDVYDGVGT-------------- 487
Query: 332 MKGESYFLSDFEKCMYGYEDVEQFEDVG 359
+G ++L+ + Y V++FED G
Sbjct: 488 SQGNPWYLTTMAMAEFLYRSVQEFEDAG 515
>STP2_GORGO (Q9N1A5) Nuclear transition protein 2 (TP-2) (TP2)
(Fragment)
Length = 133
Score = 30.0 bits (66), Expect = 9.5
Identities = 20/59 (33%), Positives = 25/59 (41%), Gaps = 4/59 (6%)
Query: 39 THRKKDGSVSSCRFVCCKEGMRKPDKRDYKTKN----PRLETRTNCEARLGLKNVDGKL 93
+HR G+ SS P KR KT N PR T +C KN++GKL
Sbjct: 55 SHRNPTGAHSSSGHQSQSPNTSPPPKRHKKTMNSHHSPRRPTILHCSCPKNRKNLEGKL 113
>FRVR_ECOLI (P32152) Putative frv operon regulatory protein
Length = 582
Score = 30.0 bits (66), Expect = 9.5
Identities = 20/66 (30%), Positives = 34/66 (51%), Gaps = 3/66 (4%)
Query: 127 QQIELADSVGVQQKKSFDLLSKE-VGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQ 185
+Q+++ D + Q + +L + V GRT L I D N+ + R G AGY L+
Sbjct: 5 RQLKIVDLLEQQPRTPGELAQQTGVSGRTILRDI--DYLNFTLNGKARIFASGSAGYQLE 62
Query: 186 YFQRKT 191
F+R++
Sbjct: 63 IFERRS 68
>CPEY_SYNPY (Q02174) Bilin biosynthesis protein cpeY
Length = 419
Score = 30.0 bits (66), Expect = 9.5
Identities = 20/64 (31%), Positives = 32/64 (49%), Gaps = 8/64 (12%)
Query: 253 HHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKA------LAE 306
H+ +FG+ ++D +AES W+FE L + PQ + A +A A L E
Sbjct: 301 HYCFICLFGS--IFDWSAESKRWIFEVLLSSISNLRPQFQKSRAASILALAKLNPSMLCE 358
Query: 307 VMPE 310
++PE
Sbjct: 359 LIPE 362
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 43,353,732
Number of Sequences: 164201
Number of extensions: 1745564
Number of successful extensions: 3948
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 3944
Number of HSP's gapped (non-prelim): 17
length of query: 362
length of database: 59,974,054
effective HSP length: 111
effective length of query: 251
effective length of database: 41,747,743
effective search space: 10478683493
effective search space used: 10478683493
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 66 (30.0 bits)
Medicago: description of AC148236.2


