

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
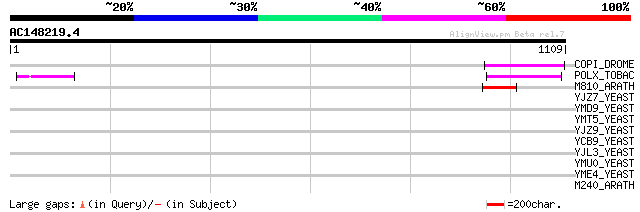
Query= AC148219.4 + phase: 0 /pseudo
(1109 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 107 2e-22
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 91 2e-17
M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810... 73 4e-12
YJZ7_YEAST (P47098) Transposon Ty1 protein B 39 0.060
YMD9_YEAST (Q03434) Transposon Ty1 protein B 38 0.13
YMT5_YEAST (Q04214) Transposon Ty1 protein B 37 0.23
YJZ9_YEAST (P47100) Transposon Ty1 protein B 37 0.23
YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B) 37 0.23
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 36 0.50
YMU0_YEAST (Q04670) Transposon Ty1 protein B 36 0.66
YME4_YEAST (Q04711) Transposon Ty1 protein B 36 0.66
M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240... 32 9.5
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 107 bits (266), Expect = 2e-22
Identities = 60/160 (37%), Positives = 88/160 (54%), Gaps = 1/160 (0%)
Query: 950 KVFFFPDVDWAGCLDSRRSIFGQCFFLGN-SLISWRTMKQLTISRSSSEAEYRALSAATY 1008
K+ + D DWAG R+S G F + + +LI W T +Q +++ SS+EAEY AL A
Sbjct: 1247 KIIGYVDSDWAGSEIDRKSTTGYLFKMFDFNLICWNTKRQNSVAASSTEAEYMALFEAVR 1306
Query: 1009 ELQWLLYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGI 1068
E WL +LL +++ +Y DNQ + I NP H+ KHI+I H R+++Q +
Sbjct: 1307 EALWLKFLLTSINIKLENPIKIYEDNQGCISIANNPSCHKRAKHIDIKYHFAREQVQNNV 1366
Query: 1069 IKFLLVSSKDQLADIFTNPLLPQPFSTLLSNLGMLNSYHS 1108
I + +++QLADIFT PL F L LG+L S
Sbjct: 1367 ICLEYIPTENQLADIFTKPLPAARFVELRDKLGLLQDDQS 1406
Score = 42.0 bits (97), Expect = 0.009
Identities = 31/120 (25%), Positives = 56/120 (45%), Gaps = 8/120 (6%)
Query: 15 SFAL*HFRLGHVSYERLAHMSQLYPF*---LFFDSNATCDICH---FARQKQLPFHLSSS 68
+F L H R GH+S +L + + F L + +C+IC +Q +LPF
Sbjct: 414 NFRLWHERFGHISDGKLLEIKRKNMFSDQSLLNNLELSCEICEPCLNGKQARLPFKQLKD 473
Query: 69 VASNKFEL--LHFDI*GPLIVPFIHNHKYFLTILDDSSRFVWIVLLKSKAEVSQHVKNFI 126
K L +H D+ GP+ + + YF+ +D + + L+K K++V ++F+
Sbjct: 474 KTHIKRPLFVVHSDVCGPITPVTLDDKNYFVIFVDQFTHYCVTYLIKYKSDVFSMFQDFV 533
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 90.5 bits (223), Expect = 2e-17
Identities = 50/149 (33%), Positives = 81/149 (53%), Gaps = 1/149 (0%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
+ D D AG +D+R+S G F ISW++ Q ++ S++EAEY A + E+ WL
Sbjct: 1178 YTDADMAGDIDNRKSSTGYLFTFSGGAISWQSKLQKCVALSTTEAEYIAATETGKEMIWL 1237
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
L +L + K V+YCD+QSA+ + N M+H TKHI++ H +R+ + +K L
Sbjct: 1238 KRFLQELGLHQ-KEYVVYCDSQSAIDLSKNSMYHARTKHIDVRYHWIREMVDDESLKVLK 1296
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGM 1102
+S+ + AD+ T + F +GM
Sbjct: 1297 ISTNENPADMLTKVVPRNKFELCKELVGM 1325
Score = 66.6 bits (161), Expect = 3e-10
Identities = 38/118 (32%), Positives = 67/118 (56%), Gaps = 5/118 (4%)
Query: 14 LSFAL*HFRLGHVSYERLAHMSQLYPF*LFFDSNAT---CDICHFARQKQLPFHLSSSVA 70
+S L H R+GH+S + L +++ + + T CD C F +Q ++ F SS
Sbjct: 420 ISVDLWHKRMGHMSEKGLQILAKKSL--ISYAKGTTVKPCDYCLFGKQHRVSFQTSSERK 477
Query: 71 SNKFELLHFDI*GPLIVPFIHNHKYFLTILDDSSRFVWIVLLKSKAEVSQHVKNFITL 128
N +L++ D+ GP+ + + +KYF+T +DD+SR +W+ +LK+K +V Q + F L
Sbjct: 478 LNILDLVYSDVCGPMEIESMGGNKYFVTFIDDASRKLWVYILKTKDQVFQVFQKFHAL 535
>M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810
(ORF240b)
Length = 240
Score = 73.2 bits (178), Expect = 4e-12
Identities = 35/68 (51%), Positives = 44/68 (64%)
Query: 945 RNVLVKVFFFPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALS 1004
+N + V F D DWAGC +RRS G C FLG ++ISW +Q T+SRSS+E EYRAL+
Sbjct: 158 KNSKLNVQAFCDSDWAGCTSTRRSTTGFCTFLGCNIISWSAKRQPTVSRSSTETEYRALA 217
Query: 1005 AATYELQW 1012
EL W
Sbjct: 218 LTAAELTW 225
>YJZ7_YEAST (P47098) Transposon Ty1 protein B
Length = 1755
Score = 39.3 bits (90), Expect = 0.060
Identities = 33/132 (25%), Positives = 59/132 (44%), Gaps = 4/132 (3%)
Query: 967 RSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWLLYLLNDLHVTTVK 1026
+S G F L +I ++ K S++EAE A+S + L L YL+ +L+ +
Sbjct: 1620 KSQIGNIFLLNGKVIGGKSTKASLTCTSTTEAEIHAISESVPLLNNLSYLIQELNKKPI- 1678
Query: 1027 LPVLYCDNQSALHI--GTNPMFHEITKHIEIICHLVRDKLQAGIIKFLLVSSKDQLADIF 1084
+ L D++S + I TN + +RD++ + + +K +AD+
Sbjct: 1679 IKGLLTDSRSTISIIKSTNEEKFR-NRFFGTKAMRLRDEVSGNNLYVYYIETKKNIADVM 1737
Query: 1085 TNPLLPQPFSTL 1096
T PL + F L
Sbjct: 1738 TKPLPIKTFKLL 1749
>YMD9_YEAST (Q03434) Transposon Ty1 protein B
Length = 1328
Score = 38.1 bits (87), Expect = 0.13
Identities = 32/132 (24%), Positives = 59/132 (44%), Gaps = 4/132 (3%)
Query: 967 RSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWLLYLLNDLHVTTVK 1026
+S G + L +I ++ K S++EAE A+S + L L YL+ +L+ +
Sbjct: 1193 KSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAEIHAISESVPLLNNLSYLIQELNKKPI- 1251
Query: 1027 LPVLYCDNQSALHI--GTNPMFHEITKHIEIICHLVRDKLQAGIIKFLLVSSKDQLADIF 1084
+ L D++S + I TN + +RD++ + + +K +AD+
Sbjct: 1252 IKGLLTDSRSTISIIKSTNEEKFR-NRFFGTKAMRLRDEVSGNNLYVYYIETKKNIADVM 1310
Query: 1085 TNPLLPQPFSTL 1096
T PL + F L
Sbjct: 1311 TKPLPIKTFKLL 1322
>YMT5_YEAST (Q04214) Transposon Ty1 protein B
Length = 1328
Score = 37.4 bits (85), Expect = 0.23
Identities = 29/130 (22%), Positives = 54/130 (41%)
Query: 967 RSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWLLYLLNDLHVTTVK 1026
+S G + L +I ++ K S++EAE A+S + L L YL+ +L +
Sbjct: 1193 KSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAEIHAISESVPLLNNLSYLIQELDKKPIT 1252
Query: 1027 LPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLLVSSKDQLADIFTN 1086
+L + I +N + +RD++ + + +K +AD+ T
Sbjct: 1253 KGLLTDSKSTISIIISNNEEKFRNRFFGTKAMRLRDEVSGNHLHVCYIETKKNIADVMTK 1312
Query: 1087 PLLPQPFSTL 1096
PL + F L
Sbjct: 1313 PLPIKTFKLL 1322
>YJZ9_YEAST (P47100) Transposon Ty1 protein B
Length = 1755
Score = 37.4 bits (85), Expect = 0.23
Identities = 29/130 (22%), Positives = 54/130 (41%)
Query: 967 RSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWLLYLLNDLHVTTVK 1026
+S G + L +I ++ K S++EAE A+S + L L YL+ +L +
Sbjct: 1620 KSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAEIHAISESVPLLNNLSYLIQELDKKPIT 1679
Query: 1027 LPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLLVSSKDQLADIFTN 1086
+L + I +N + +RD++ + + +K +AD+ T
Sbjct: 1680 KGLLTDSKSTISIIISNNEEKFRNRFFGTKAMRLRDEVSGNHLHVCYIETKKNIADVMTK 1739
Query: 1087 PLLPQPFSTL 1096
PL + F L
Sbjct: 1740 PLPIKTFKLL 1749
>YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B)
Length = 1770
Score = 37.4 bits (85), Expect = 0.23
Identities = 31/131 (23%), Positives = 59/131 (44%), Gaps = 2/131 (1%)
Query: 967 RSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWLLYLLNDLHVTTVK 1026
+S G F L +I ++ K S++EAE A+S A L L +L+ +L+ +
Sbjct: 1635 KSQIGNIFLLNGKVIGGKSTKASLTCTSTTEAEIHAVSEAIPLLNNLSHLVQELNKKPI- 1693
Query: 1027 LPVLYCDNQSALHIGTNPMFHEI-TKHIEIICHLVRDKLQAGIIKFLLVSSKDQLADIFT 1085
+ L D++S + I + + + +RD++ + + +K +AD+ T
Sbjct: 1694 IKGLLTDSRSTISIIKSTNEEKFRNRFFGTKAMRLRDEVSGNNLYVYYIETKKNIADVMT 1753
Query: 1086 NPLLPQPFSTL 1096
PL + F L
Sbjct: 1754 KPLPIKTFKLL 1764
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 36.2 bits (82), Expect = 0.50
Identities = 31/135 (22%), Positives = 60/135 (43%), Gaps = 2/135 (1%)
Query: 967 RSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWLLYLLNDLHVTTVK 1026
+S G + G ++ + + K SS+EAE A+ + + L L +L
Sbjct: 1655 QSRIGVILWYGMNIFNVYSNKSTNRCVSSTEAELHAIYEGYADSETLKVTLKELGEGDNN 1714
Query: 1027 LPVLYCDNQSALHIGTNPMFHEIT-KHIEIICHLVRDKLQAGIIKFLLVSSKDQLADIFT 1085
V+ D++ A+ G N + + K I ++++K++ IK L ++ K +AD+ T
Sbjct: 1715 DIVMITDSKPAIQ-GLNRSYQQPKEKFTWIKTEIIKEKIKEKSIKLLKITGKGNIADLLT 1773
Query: 1086 NPLLPQPFSTLLSNL 1100
P+ F + L
Sbjct: 1774 KPVSASDFKRFIQVL 1788
>YMU0_YEAST (Q04670) Transposon Ty1 protein B
Length = 1328
Score = 35.8 bits (81), Expect = 0.66
Identities = 28/130 (21%), Positives = 55/130 (41%)
Query: 967 RSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWLLYLLNDLHVTTVK 1026
+S G + L +I ++ K S++EAE A+S + L L +L+ +L+ +
Sbjct: 1193 KSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAEIHAISESVPLLNNLSHLVQELNKKPIT 1252
Query: 1027 LPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLLVSSKDQLADIFTN 1086
+L + I +N + +RD++ + + +K +AD+ T
Sbjct: 1253 KGLLTDSKSTISIIISNNEEKFRNRFFGTKAMRLRDEVSGNHLHVCYIETKKNIADVMTK 1312
Query: 1087 PLLPQPFSTL 1096
PL + F L
Sbjct: 1313 PLPIKTFKLL 1322
>YME4_YEAST (Q04711) Transposon Ty1 protein B
Length = 1328
Score = 35.8 bits (81), Expect = 0.66
Identities = 28/130 (21%), Positives = 55/130 (41%)
Query: 967 RSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWLLYLLNDLHVTTVK 1026
+S G + L +I ++ K S++EAE A+S + L L +L+ +L+ +
Sbjct: 1193 KSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAEIHAISESVPLLNNLSHLVQELNKKPIT 1252
Query: 1027 LPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLLVSSKDQLADIFTN 1086
+L + I +N + +RD++ + + +K +AD+ T
Sbjct: 1253 KGLLTDSKSTISIIISNNEEKFRNRFFGTKAMRLRDEVSGNHLHVCYIETKKNIADVMTK 1312
Query: 1087 PLLPQPFSTL 1096
PL + F L
Sbjct: 1313 PLPIKTFKLL 1322
>M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240
(ORF111a)
Length = 111
Score = 32.0 bits (71), Expect = 9.5
Identities = 15/29 (51%), Positives = 18/29 (61%), Gaps = 5/29 (17%)
Query: 954 FPDVDWAGCLDSRRSIFGQC-----FFLG 977
F D DWA C D+RRS+ G C +FLG
Sbjct: 60 FADSDWASCPDTRRSVTGFCSLVPLWFLG 88
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.362 0.160 0.603
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 106,230,886
Number of Sequences: 164201
Number of extensions: 3874581
Number of successful extensions: 15521
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 15479
Number of HSP's gapped (non-prelim): 37
length of query: 1109
length of database: 59,974,054
effective HSP length: 121
effective length of query: 988
effective length of database: 40,105,733
effective search space: 39624464204
effective search space used: 39624464204
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (22.0 bits)
S2: 71 (32.0 bits)
Medicago: description of AC148219.4


