

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148154.6 - phase: 0
(199 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
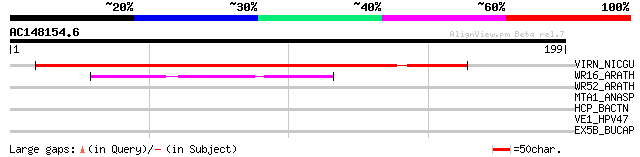
Score E
Sequences producing significant alignments: (bits) Value
VIRN_NICGU (Q40392) TMV resistance protein N 142 7e-34
WR16_ARATH (Q9FL92) Probable WRKY transcription factor 16 (WRKY ... 50 4e-06
WR52_ARATH (Q9FH83) Probable WRKY transcription factor 52 (WRKY ... 38 0.018
MTA1_ANASP (P70802) Modification methylase AvaI (EC 2.1.1.113) (... 30 4.8
HCP_BACTN (Q8A9X8) Hydroxylamine reductase (EC 1.7.-.-) (Hybrid-... 29 6.2
VE1_HPV47 (P22419) Replication protein E1 29 8.2
EX5B_BUCAP (Q8K9A9) Exodeoxyribonuclease V beta chain (EC 3.1.11.5) 29 8.2
>VIRN_NICGU (Q40392) TMV resistance protein N
Length = 1144
Score = 142 bits (357), Expect = 7e-34
Identities = 74/155 (47%), Positives = 102/155 (65%), Gaps = 3/155 (1%)
Query: 10 SSSLISYGFTYQVFLNFRGSDTRDGFTGHLYKALTDKGIHTFIDDCDLKRGDEITPSLIK 69
+SS S ++Y VFL+FRG DTR FT HLY+ L DKGI TF DD L+ G I L K
Sbjct: 2 ASSSSSSRWSYDVFLSFRGEDTRKTFTSHLYEVLNDKGIKTFQDDKRLEYGATIPGELCK 61
Query: 70 AIEESRIFIPVFSINYASSKFCLDELVHIIHCYKTKGRLVLPVFYGVDPTQIRHQSGSYG 129
AIEES+ I VFS NYA+S++CL+ELV I+ C + V+P+FY VDP+ +R+Q S+
Sbjct: 62 AIEESQFAIVVFSENYATSRWCLNELVKIMECKTRFKQTVIPIFYDVDPSHVRNQKESFA 121
Query: 130 EHLTKHEESFQNNKKNKERLHQWKLALTQAANLSG 164
+ +HE + K + E + +W++AL +AANL G
Sbjct: 122 KAFEEHETKY---KDDVEGIQRWRIALNEAANLKG 153
>WR16_ARATH (Q9FL92) Probable WRKY transcription factor 16 (WRKY
DNA-binding protein 16)
Length = 1372
Score = 49.7 bits (117), Expect = 4e-06
Identities = 29/87 (33%), Positives = 45/87 (51%), Gaps = 7/87 (8%)
Query: 30 DTRDGFTGHLYKALTDKGIHTFIDDCDLKRGDEITPSLIKAIEESRIFIPVFSINYASSK 89
+ R F HL KAL KG++ D D D ++ +E +R+ + + N S
Sbjct: 15 EVRYSFVSHLSKALQRKGVNDVFIDSD----DSLSNESQSMVERARVSVMILPGNRTVS- 69
Query: 90 FCLDELVHIIHCYKTKGRLVLPVFYGV 116
LD+LV ++ C K K ++V+PV YGV
Sbjct: 70 --LDKLVKVLDCQKNKDQVVVPVLYGV 94
>WR52_ARATH (Q9FH83) Probable WRKY transcription factor 52 (WRKY
DNA-binding protein 52) (Disease resistance protein
RRS1) (Resistance to Ralstonia solanacearum 1 protein)
Length = 1378
Score = 37.7 bits (86), Expect = 0.018
Identities = 25/87 (28%), Positives = 42/87 (47%), Gaps = 3/87 (3%)
Query: 30 DTRDGFTGHLYKALTDKGIHTFIDDCDLKRGDEITPSLIKAIEESRIFIPVFSINYASSK 89
+ R F HL +AL KGI+ + D D+ D + IE++ + + V N S+
Sbjct: 18 EVRYSFVSHLSEALRRKGINNVVVDVDI--DDLLFKESQAKIEKAGVSVMVLPGNCDPSE 75
Query: 90 FCLDELVHIIHCYK-TKGRLVLPVFYG 115
LD+ ++ C + K + V+ V YG
Sbjct: 76 VWLDKFAKVLECQRNNKDQAVVSVLYG 102
>MTA1_ANASP (P70802) Modification methylase AvaI (EC 2.1.1.113) (N-4
cytosine-specific methyltransferase AvaI) (M.AvaI)
Length = 482
Score = 29.6 bits (65), Expect = 4.8
Identities = 12/48 (25%), Positives = 26/48 (54%)
Query: 104 TKGRLVLPVFYGVDPTQIRHQSGSYGEHLTKHEESFQNNKKNKERLHQ 151
T L P+ DPT + +S S+ + +++ ++ + ++K RLH+
Sbjct: 2 TSFELESPIEIKTDPTDLDQESDSFVQEISRFNKALEQRFRDKMRLHE 49
>HCP_BACTN (Q8A9X8) Hydroxylamine reductase (EC 1.7.-.-)
(Hybrid-cluster protein) (HCP)
Length = 543
Score = 29.3 bits (64), Expect = 6.2
Identities = 20/78 (25%), Positives = 32/78 (40%), Gaps = 22/78 (28%)
Query: 109 VLPVFYGVDPTQIRHQSGSYGEHLTKHEESFQN-----------------NKKNKERLHQ 151
+LP Y + +H +G+YG K +E F++ N K+R++
Sbjct: 261 MLPAHYYPQLKKYKHLAGNYGNAWWKQKEEFESFNGPILFTSNCIVPPRANASYKDRIY- 319
Query: 152 WKLALTQAANLSGYHYSP 169
+T A L G HY P
Sbjct: 320 ----ITGACGLEGAHYIP 333
>VE1_HPV47 (P22419) Replication protein E1
Length = 605
Score = 28.9 bits (63), Expect = 8.2
Identities = 18/53 (33%), Positives = 24/53 (44%), Gaps = 1/53 (1%)
Query: 9 SSSSLISYGFTYQVFLNFRGSDTRDGFTGHLYKALTDKGIHTFIDDCDLKRGD 61
S S GF L SD D G L++ TD I +D+CDL +G+
Sbjct: 4 SKGSTSKEGFGDWCILEADCSDVEDDL-GQLFERDTDSDISDLLDNCDLDQGN 55
>EX5B_BUCAP (Q8K9A9) Exodeoxyribonuclease V beta chain (EC 3.1.11.5)
Length = 1179
Score = 28.9 bits (63), Expect = 8.2
Identities = 30/115 (26%), Positives = 50/115 (43%), Gaps = 13/115 (11%)
Query: 69 KAIEESRIFIPVFSINYASSKFCLDELVHIIHCYKTKGRLVLPVF-------YGVDPTQI 121
K +E++ I + N+A SK ++ II +K+KG L P+ + V +
Sbjct: 723 KILEKNNISENEYIKNFAESK-----IIRIITIHKSKG-LEYPIVWIPFIVDFNVSKSYF 776
Query: 122 RHQSGSYGEHLTKHEESFQNNKKNKERLHQWKLALTQAANLSGYHYSPGYPTLLK 176
H+ + ++ S K ++ERL + L A S YH S G L+K
Sbjct: 777 YHEKKTLKIFFDNNKSSETLKKSDEERLAEDLRFLYVALTRSIYHCSIGISYLVK 831
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.140 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,013,409
Number of Sequences: 164201
Number of extensions: 1068454
Number of successful extensions: 2293
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 2288
Number of HSP's gapped (non-prelim): 7
length of query: 199
length of database: 59,974,054
effective HSP length: 105
effective length of query: 94
effective length of database: 42,732,949
effective search space: 4016897206
effective search space used: 4016897206
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC148154.6


