

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148098.5 + phase: 0
(189 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
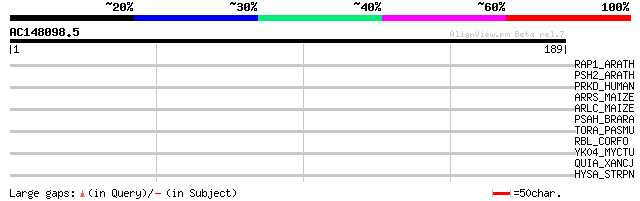
Score E
Sequences producing significant alignments: (bits) Value
RAP1_ARATH (Q39204) Transcription factor AtMYC2 (R-homologous Ar... 32 0.67
PSH2_ARATH (Q9SUI6) Photosystem I reaction center subunit VI-2, ... 31 1.9
PRKD_HUMAN (P78527) DNA-dependent protein kinase catalytic subun... 30 2.5
ARRS_MAIZE (P13027) Anthocyanin regulatory R-S protein 30 2.5
ARLC_MAIZE (P13526) Anthocyanin regulatory Lc protein 30 2.5
PSAH_BRARA (O04006) Photosystem I reaction center subunit VI, ch... 30 3.3
TORA_PASMU (Q9CK41) Trimethylamine-N-oxide reductase precursor (... 29 5.7
RBL_CORFO (Q31983) Ribulose bisphosphate carboxylase large chain... 29 5.7
YK04_MYCTU (Q10852) Hypothetical protein Rv2004c/MT2060/Mb2027c 28 9.7
QUIA_XANCJ (Q9XD78) Probable quinate dehydrogenase [Pyrroloquino... 28 9.7
HYSA_STRPN (Q54873) Hyaluronate lyase precursor (EC 4.2.2.1) (Hy... 28 9.7
>RAP1_ARATH (Q39204) Transcription factor AtMYC2 (R-homologous
Arabidopsis protein-1) (RAP-1) (Basic helix-loop-helix
protein 6) (bHLH6) (AtbHLH006) (rd22BP1)
Length = 623
Score = 32.3 bits (72), Expect = 0.67
Identities = 48/217 (22%), Positives = 71/217 (32%), Gaps = 76/217 (35%)
Query: 5 LPLLHSLLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKR 64
+P Q TL+ T W YA+FW+ P ++F G S
Sbjct: 57 IPAQAGFNQETLQQRLQALIEGTHEGWTYAIFWQ--------PSYDFSGAS--------- 99
Query: 65 NWIIVWEDGFCDFNECE---QRKSGC------------------LNERFGADV------- 96
++ W DG+ E + +R+S LN V
Sbjct: 100 --VLGWGDGYYKGEEDKANPRRRSSSPPFSTPADQEYRKKVLRELNSLISGGVAPSDDAV 157
Query: 97 ---------FFKMS-HEVYSYGEGLVGKVAADNGHKWVYSDTQ---NGCEANYVGPWNAS 143
FF +S + ++ G GL GK A WV Q +GCE G
Sbjct: 158 DEEVTDTEWFFLVSMTQSFACGAGLAGKAFATGNAVWVSGSDQLSGSGCERAKQGG---- 213
Query: 144 IDHQPRAWEFQLNSGIQTIAVIAVKEGLVQLGSFDKV 180
G+ TIA I G+V++GS + +
Sbjct: 214 ------------VFGMHTIACIPSANGVVEVGSTEPI 238
>PSH2_ARATH (Q9SUI6) Photosystem I reaction center subunit VI-2,
chloroplast precursor (PSI-H1)
Length = 145
Score = 30.8 bits (68), Expect = 1.9
Identities = 22/80 (27%), Positives = 32/80 (39%), Gaps = 2/80 (2%)
Query: 93 GADVFFKMSHEVYSYGEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWE 152
GA +F K S + + G V A G K VY D ++ N G W+ P +
Sbjct: 26 GAKLFIKPSRQSFKTKSTRAGAVVAKYGDKSVYFDLED--LGNTTGQWDVYGSDAPSPYN 83
Query: 153 FQLNSGIQTIAVIAVKEGLV 172
+ +T A K GL+
Sbjct: 84 PLQSKFFETFAAPFTKRGLL 103
>PRKD_HUMAN (P78527) DNA-dependent protein kinase catalytic subunit
(EC 2.7.1.37) (DNA-PKcs) (DNPK1) (p460)
Length = 4128
Score = 30.4 bits (67), Expect = 2.5
Identities = 8/32 (25%), Positives = 15/32 (46%)
Query: 56 LDRSKGNKRNWIIVWEDGFCDFNECEQRKSGC 87
L + + +W++ W +C + C R GC
Sbjct: 3262 LHKESKTRDDWLVSWVQSYCRLSHCRSRSQGC 3293
>ARRS_MAIZE (P13027) Anthocyanin regulatory R-S protein
Length = 612
Score = 30.4 bits (67), Expect = 2.5
Identities = 29/101 (28%), Positives = 48/101 (46%), Gaps = 16/101 (15%)
Query: 83 RKSGCLN-ERFGADVFFKMSHEVYSY--GEGLVGKVAADNGHKWVYSDTQNGCEANYVGP 139
R +G L+ E G ++ + Y++ G+GL G+ A + H W+ C A+ G
Sbjct: 108 RPAGSLSPEDLGDTEWYYVVSMTYAFRPGQGLPGRSFASDEHVWL-------CNAHLAGS 160
Query: 140 WNASIDHQPRAWEFQLNSGIQTIAVIAVKEGLVQLGSFDKV 180
PRA ++ IQ+I I V G+++LG+ D V
Sbjct: 161 -----KAFPRAL-LAKSASIQSILCIPVMGGVLELGTTDTV 195
>ARLC_MAIZE (P13526) Anthocyanin regulatory Lc protein
Length = 610
Score = 30.4 bits (67), Expect = 2.5
Identities = 29/101 (28%), Positives = 48/101 (46%), Gaps = 16/101 (15%)
Query: 83 RKSGCLN-ERFGADVFFKMSHEVYSY--GEGLVGKVAADNGHKWVYSDTQNGCEANYVGP 139
R +G L+ E G ++ + Y++ G+GL G+ A + H W+ C A+ G
Sbjct: 108 RPAGSLSPEDLGDTEWYYVVSMTYAFRPGQGLPGRSFASDEHVWL-------CNAHLAGS 160
Query: 140 WNASIDHQPRAWEFQLNSGIQTIAVIAVKEGLVQLGSFDKV 180
PRA ++ IQ+I I V G+++LG+ D V
Sbjct: 161 -----KAFPRAL-LAKSASIQSILCIPVMGGVLELGTTDTV 195
>PSAH_BRARA (O04006) Photosystem I reaction center subunit VI,
chloroplast precursor (PSI-H) (Light-harvesting complex
I 11 kDa protein)
Length = 145
Score = 30.0 bits (66), Expect = 3.3
Identities = 22/80 (27%), Positives = 32/80 (39%), Gaps = 2/80 (2%)
Query: 93 GADVFFKMSHEVYSYGEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWE 152
GA +F K S + + G V A G K VY D ++ N G W+ P +
Sbjct: 26 GAKLFIKPSRQSFKPKSTRAGAVVAKYGDKSVYFDLED--LGNTTGQWDLYGSDAPSPYN 83
Query: 153 FQLNSGIQTIAVIAVKEGLV 172
+ +T A K GL+
Sbjct: 84 PLQSKFFETFAAPFTKRGLL 103
>TORA_PASMU (Q9CK41) Trimethylamine-N-oxide reductase precursor (EC
1.7.2.3) (TMAO reductase) (Trimethylamine oxidase)
Length = 823
Score = 29.3 bits (64), Expect = 5.7
Identities = 19/45 (42%), Positives = 24/45 (53%), Gaps = 6/45 (13%)
Query: 101 SHEVYSYGEGLVGKVAADNGH----KWVYSDTQN--GCEANYVGP 139
+HE Y Y E L KVAA + + V S TQN GC+ Y+ P
Sbjct: 234 THEAYKYLEQLKAKVAAKDVNVICVDPVKSKTQNYLGCDFQYINP 278
>RBL_CORFO (Q31983) Ribulose bisphosphate carboxylase large chain
(EC 4.1.1.39) (RuBisCO large subunit) (Fragment)
Length = 465
Score = 29.3 bits (64), Expect = 5.7
Identities = 19/61 (31%), Positives = 25/61 (40%), Gaps = 4/61 (6%)
Query: 23 PTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVWEDGFCDFNECEQ 82
PT T + A F RV P+ PP + + S G WI VW DG + +
Sbjct: 17 PTYETKDTDILAAF-RVTPQPGVPPEEAGAAVAAESSTGT---WITVWTDGLTSLDRYKG 72
Query: 83 R 83
R
Sbjct: 73 R 73
>YK04_MYCTU (Q10852) Hypothetical protein Rv2004c/MT2060/Mb2027c
Length = 498
Score = 28.5 bits (62), Expect = 9.7
Identities = 13/34 (38%), Positives = 17/34 (49%), Gaps = 5/34 (14%)
Query: 74 FCDFNECEQRKSGC-----LNERFGADVFFKMSH 102
FCDF EQR+ C LN R A + ++H
Sbjct: 49 FCDFRTAEQRERACIREFELNSRLAAQSYLGIAH 82
>QUIA_XANCJ (Q9XD78) Probable quinate dehydrogenase
[Pyrroloquinoline-quinone] (EC 1.1.99.25)
Length = 790
Score = 28.5 bits (62), Expect = 9.7
Identities = 16/61 (26%), Positives = 23/61 (37%), Gaps = 7/61 (11%)
Query: 105 YSYGEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAV 164
+ YG ++ A KWVY N W+ + QP +F G T AV
Sbjct: 430 HRYGASVLALDATTGAEKWVYQTVHNDL-------WDFDLPMQPSLIDFPNQDGSHTPAV 482
Query: 165 I 165
+
Sbjct: 483 V 483
>HYSA_STRPN (Q54873) Hyaluronate lyase precursor (EC 4.2.2.1)
(Hyaluronidase) (HYase)
Length = 1066
Score = 28.5 bits (62), Expect = 9.7
Identities = 14/42 (33%), Positives = 20/42 (47%)
Query: 139 PWNASIDHQPRAWEFQLNSGIQTIAVIAVKEGLVQLGSFDKV 180
PW S AW Q NS + VI K+G + + S +K+
Sbjct: 70 PWTGSKAQGWSAWVDQKNSADASTRVIEAKDGAITISSHEKL 111
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.138 0.457
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,442,790
Number of Sequences: 164201
Number of extensions: 1089598
Number of successful extensions: 2151
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 2147
Number of HSP's gapped (non-prelim): 14
length of query: 189
length of database: 59,974,054
effective HSP length: 104
effective length of query: 85
effective length of database: 42,897,150
effective search space: 3646257750
effective search space used: 3646257750
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC148098.5


