

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147960.5 - phase: 0
(617 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
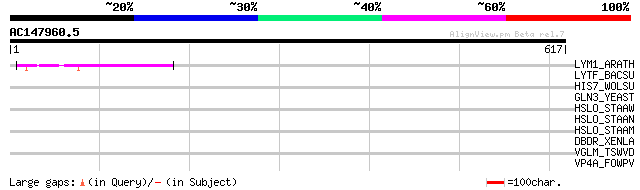
Score E
Sequences producing significant alignments: (bits) Value
LYM1_ARATH (Q93ZH0) LysM-domain GPI-anchored protein 1 precursor 54 9e-07
LYTF_BACSU (O07532) Endopeptidase lytF precursor (EC 3.4.-.-) (G... 35 0.44
HIS7_WOLSU (Q7M7V2) Imidazoleglycerol-phosphate dehydratase (EC ... 33 2.9
GLN3_YEAST (P18494) Nitrogen regulatory protein GLN3 33 2.9
HSLO_STAAW (Q8NXZ3) 33 kDa chaperonin (Heat shock protein 33 hom... 32 3.8
HSLO_STAAN (P99082) 33 kDa chaperonin (Heat shock protein 33 hom... 32 3.8
HSLO_STAAM (P64400) 33 kDa chaperonin (Heat shock protein 33 hom... 32 3.8
DBDR_XENLA (P42290) D(1B) dopamine receptor (D(5) dopamine recep... 32 4.9
VGLM_TSWVD (Q9IKB7) M polyprotein precursor [Contains: Glycoprot... 32 6.4
VP4A_FOWPV (Q9J559) Major core protein P4a 31 8.4
>LYM1_ARATH (Q93ZH0) LysM-domain GPI-anchored protein 1 precursor
Length = 416
Score = 54.3 bits (129), Expect = 9e-07
Identities = 48/188 (25%), Positives = 84/188 (44%), Gaps = 18/188 (9%)
Query: 8 LSFLFMLLAS--------KSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQSNLVTK 59
L F+ ++LAS KS I + TCN AL Y L D ++ V+++ Q +
Sbjct: 10 LIFVSLILASSLTFTATAKSTIEPCSSNDTCN-ALLGYTLYTDLKVSEVASLFQVD---- 64
Query: 60 PEDIVSYNTDTITNKD----FVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATKDTYLSVA 115
P I+ N I+ D + S + +P C C+ Y+ D S+A
Sbjct: 65 PISILLANAIDISYPDVENHILPSKLFLKIPITCSCVDGIRKSVSTHYKTRPSDNLGSIA 124
Query: 116 SNNYSNLTTSEWLQNFNSYPSNDIPDTGT-LNVTVNCSCGNSDVSKDYGLFITYPLRPED 174
+ Y L ++E +Q NS + D GT L + + C+C N + ++++Y ++ D
Sbjct: 125 DSVYGGLVSAEQIQEANSVNDPSLLDVGTSLVIPLPCACFNGTDNSLPAVYLSYVVKEID 184
Query: 175 SLELISNK 182
+L I+ +
Sbjct: 185 TLVGIARR 192
>LYTF_BACSU (O07532) Endopeptidase lytF precursor (EC 3.4.-.-)
(Gamma-D-glutamate-meso-diaminopimelate muropeptidase
lytF) (Cell wall-associated polypeptide CWBP49)
Length = 488
Score = 35.4 bits (80), Expect = 0.44
Identities = 29/98 (29%), Positives = 50/98 (50%), Gaps = 8/98 (8%)
Query: 102 QYQVATKDTYLSVASNNYSNLTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKD 161
+Y V + D+ +A NNY NLT + ++N N+ S+ + L +T S G+S S
Sbjct: 241 KYTVKSGDSLWKIA-NNY-NLTVQQ-IRNINNLKSDVLYVGQVLKLTGKASSGSSSSSSS 297
Query: 162 Y-----GLFITYPLRPEDSLELISNKTEIDAELLQKYN 194
G TY ++ DSL +I+ K + A+ +++ N
Sbjct: 298 SSNASSGTTTTYTVKSGDSLWVIAQKFNVTAQQIREKN 335
>HIS7_WOLSU (Q7M7V2) Imidazoleglycerol-phosphate dehydratase (EC
4.2.1.19) (IGPD)
Length = 190
Score = 32.7 bits (73), Expect = 2.9
Identities = 20/85 (23%), Positives = 41/85 (47%), Gaps = 6/85 (7%)
Query: 134 YPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELISNKTEIDAELLQKY 193
YP+ I G +V ++ SC D+ F+ + + E ++ E DAEL +++
Sbjct: 81 YPAQGIERFGNASVVMDESCVECDLDVSNRPFLVFDMTIEGAVG------EFDAELAEEF 134
Query: 194 NPGVNFSQGSGLVYIPGKDQNRNYV 218
+ F+ G L + + +NR+++
Sbjct: 135 FRALAFNAGLSLHIVQKRGKNRHHL 159
>GLN3_YEAST (P18494) Nitrogen regulatory protein GLN3
Length = 730
Score = 32.7 bits (73), Expect = 2.9
Identities = 21/85 (24%), Positives = 43/85 (49%), Gaps = 8/85 (9%)
Query: 100 IFQYQVATKDTYLSVASNNYSNLTTSEWLQNFNSYP-SNDIPDTGTLNVTVNCSCGNSDV 158
+F Y +T ++ ++ S N ++ T ++ +FN Y N+ + +N+T N + NS++
Sbjct: 182 LFPYNSSTSNSNINQPSINNNSNTNAQSHHSFNIYKLQNNNSSSSAMNITNNNNSNNSNI 241
Query: 159 SKDYGLFITYPLRPEDSLELISNKT 183
+ L+ DS+ L S+ T
Sbjct: 242 QHPF-------LKKSDSIGLSSSNT 259
>HSLO_STAAW (Q8NXZ3) 33 kDa chaperonin (Heat shock protein 33
homolog) (HSP33)
Length = 293
Score = 32.3 bits (72), Expect = 3.8
Identities = 18/58 (31%), Positives = 32/58 (55%), Gaps = 6/58 (10%)
Query: 43 TNLTYVSNIMQSNLVTKPEDIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHI 100
+ +T VS +++ L PE +++ I +D VQ ++ V F C+C H++FL I
Sbjct: 198 SEMTPVSKLIEQGLT--PEGLLN----EILGEDHVQILEKMPVQFECNCSHEKFLNAI 249
>HSLO_STAAN (P99082) 33 kDa chaperonin (Heat shock protein 33
homolog) (HSP33)
Length = 293
Score = 32.3 bits (72), Expect = 3.8
Identities = 18/58 (31%), Positives = 32/58 (55%), Gaps = 6/58 (10%)
Query: 43 TNLTYVSNIMQSNLVTKPEDIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHI 100
+ +T VS +++ L PE +++ I +D VQ ++ V F C+C H++FL I
Sbjct: 198 SEMTPVSKLIEQGLT--PEGLLN----EILGEDHVQILEKMPVQFECNCSHEKFLNAI 249
>HSLO_STAAM (P64400) 33 kDa chaperonin (Heat shock protein 33
homolog) (HSP33)
Length = 293
Score = 32.3 bits (72), Expect = 3.8
Identities = 18/58 (31%), Positives = 32/58 (55%), Gaps = 6/58 (10%)
Query: 43 TNLTYVSNIMQSNLVTKPEDIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHDEFLGHI 100
+ +T VS +++ L PE +++ I +D VQ ++ V F C+C H++FL I
Sbjct: 198 SEMTPVSKLIEQGLT--PEGLLN----EILGEDHVQILEKMPVQFECNCSHEKFLNAI 249
>DBDR_XENLA (P42290) D(1B) dopamine receptor (D(5) dopamine
receptor)
Length = 457
Score = 32.0 bits (71), Expect = 4.9
Identities = 17/57 (29%), Positives = 31/57 (53%), Gaps = 3/57 (5%)
Query: 62 DIVSYNTDTITNKDFVQSFTRVNVPFPCDCIHD--EFLGHIFQYQVATKDTYLSVAS 116
+++SYN DT+ +KD V ++ + +P DCI D + H+ Q + + L+ S
Sbjct: 378 ELISYNQDTLFHKDIVTAYVNM-IPNVVDCIDDNEDAFDHMSQISQTSANNELATDS 433
>VGLM_TSWVD (Q9IKB7) M polyprotein precursor [Contains: Glycoprotein
G1; Glycoprotein G2]
Length = 1135
Score = 31.6 bits (70), Expect = 6.4
Identities = 17/61 (27%), Positives = 31/61 (49%), Gaps = 2/61 (3%)
Query: 111 YLSVASNNY--SNLTTSEWLQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSKDYGLFITY 168
+ SV +N + +LT + +NSYP+N T+ ++ +C S+ + Y + IT
Sbjct: 182 HFSVGTNFFIPESLTQDNYPITYNSYPTNGTVSLQTVKLSGDCKITKSNFANPYTVSITS 241
Query: 169 P 169
P
Sbjct: 242 P 242
>VP4A_FOWPV (Q9J559) Major core protein P4a
Length = 891
Score = 31.2 bits (69), Expect = 8.4
Identities = 20/88 (22%), Positives = 40/88 (44%), Gaps = 10/88 (11%)
Query: 9 SFLFMLLASKSFIAESKCSKTCNIALASYYLQDDTNLTYVSNIMQ---------SNLVTK 59
+F+FML + F + + + SYY+ + Y++N++Q L++
Sbjct: 522 TFIFMLFKAMGFNVRVNRTTISSHSYTSYYVSPRVSKRYLTNMLQKASCSQSEAEKLLSS 581
Query: 60 PEDIVSYNTDTITNKDFVQSFTRVNVPF 87
D+VS+ ++ N S+ R+N F
Sbjct: 582 AHDLVSFML-SVNNSSNRDSYRRINRSF 608
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.351 0.153 0.577
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 64,441,848
Number of Sequences: 164201
Number of extensions: 2459079
Number of successful extensions: 8898
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 8895
Number of HSP's gapped (non-prelim): 12
length of query: 617
length of database: 59,974,054
effective HSP length: 116
effective length of query: 501
effective length of database: 40,926,738
effective search space: 20504295738
effective search space used: 20504295738
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 69 (31.2 bits)
Medicago: description of AC147960.5


