

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147959.10 + phase: 1 /pseudo
(378 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
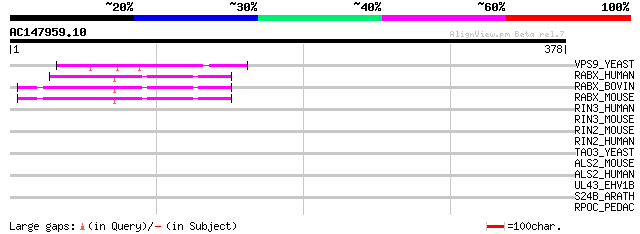
VPS9_YEAST (P54787) Vacuolar protein sorting-associated protein ... 79 1e-14
RABX_HUMAN (Q9UJ41) Rab5 GDP/GTP exchange factor (Rabex-5) 60 7e-09
RABX_BOVIN (O18973) Rab5 GDP/GTP exchange factor (Rabex-5) 59 2e-08
RABX_MOUSE (Q9JM13) Rab5 GDP/GTP exchange factor (Rabex-5) 57 6e-08
RIN3_HUMAN (Q8TB24) Ras and Rab interactor 3 (Ras interaction/in... 42 0.003
RIN3_MOUSE (P59729) Ras and Rab interactor 3 (Ras interaction/in... 40 0.007
RIN2_MOUSE (Q9D684) Ras and Rab interactor 2 (Ras interaction/in... 36 0.18
RIN2_HUMAN (Q8WYP3) Ras and Rab interactor 2 (Ras interaction/in... 35 0.24
TAO3_YEAST (P40468) Cell morphogenesis protein PAG1 (TAO3 protein) 35 0.31
ALS2_MOUSE (Q920R0) Alsin (Amyotrophic lateral sclerosis protein... 34 0.53
ALS2_HUMAN (Q96Q42) Alsin (Amyotrophic lateral sclerosis protein 2) 34 0.70
UL43_EHV1B (P28959) Gene 17 membrane protein 31 4.5
S24B_ARATH (Q9M081) Putative protein transport protein Sec24-lik... 30 7.7
RPOC_PEDAC (P77917) DNA-directed RNA polymerase beta' chain (EC ... 30 7.7
>VPS9_YEAST (P54787) Vacuolar protein sorting-associated protein
VPS9
Length = 451
Score = 79.3 bits (194), Expect = 1e-14
Identities = 55/145 (37%), Positives = 79/145 (53%), Gaps = 18/145 (12%)
Query: 33 AMEGLEKYIMTKLFSRTFAAS-------PEDAKIDHEISEKISLLQT-----FLKPEHLD 80
A EG+EK IM KL+SR F+ S P D + +++ +LL+ F+ P LD
Sbjct: 133 AKEGMEKLIMGKLYSRCFSPSLYEILQKPLDDEHMKDLTNDDTLLEKIRHYRFISPIMLD 192
Query: 81 IPPVLHN---EASWLLAEKELQKINAFKAPQEKLSTIMNCCRVINNLLLNAAMSEYVPAG 137
IP + N LA KEL KIN FK+P++K+ ++N +VI LL + + + G
Sbjct: 193 IPDTMPNARLNKFVHLASKELGKINRFKSPRDKMVCVLNASKVIFGLLKHTKLEQ---NG 249
Query: 138 ADDFIPVLIYVTIKARLALLVQTSN 162
AD FIPVLIY +K ++ LV N
Sbjct: 250 ADSFIPVLIYCILKGQVRYLVSNVN 274
>RABX_HUMAN (Q9UJ41) Rab5 GDP/GTP exchange factor (Rabex-5)
Length = 708
Score = 60.5 bits (145), Expect = 7e-09
Identities = 41/130 (31%), Positives = 70/130 (53%), Gaps = 12/130 (9%)
Query: 28 EDIDCAMEGLEKYIMTKLFSRTFAA-SPEDAKIDHEISEKISLL-----QTFLKPEHLDI 81
E ++ M+ +EKYIMT+L+ F + +D K D I ++I L Q P + DI
Sbjct: 419 ERVEKIMDQIEKYIMTRLYKYVFCPETTDDEKKDLAIQKRIRALRWVTPQMLCVPVNEDI 478
Query: 82 PPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRVINNLLLNAAMSEYVPAGADDF 141
P V A ++ ++++ + P++KL+ I C + I N + +++ PA ADDF
Sbjct: 479 PEVSDMVVK---AITDIIEMDSKRVPRDKLACITKCSKHIFNAI---KITKNEPASADDF 532
Query: 142 IPVLIYVTIK 151
+P LIY+ +K
Sbjct: 533 LPTLIYIVLK 542
>RABX_BOVIN (O18973) Rab5 GDP/GTP exchange factor (Rabex-5)
Length = 492
Score = 59.3 bits (142), Expect = 2e-08
Identities = 43/152 (28%), Positives = 78/152 (51%), Gaps = 15/152 (9%)
Query: 6 QDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAA-SPEDAKIDHEIS 64
QDF+ ++ ++ E ++ M+ +EKYIMT+L+ F + +D K D I
Sbjct: 184 QDFYQNVAERMQTR---GKVPPERVEKIMDQIEKYIMTRLYKYVFCPETTDDEKKDLAIQ 240
Query: 65 EKISLL-----QTFLKPEHLDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCR 119
++I L Q P + +IP V A ++ ++++ + P++KL+ I C +
Sbjct: 241 KRIRALHWVTPQMLCVPVNEEIPEVSDMVVK---AITDIIEMDSKRVPRDKLACITKCSK 297
Query: 120 VINNLLLNAAMSEYVPAGADDFIPVLIYVTIK 151
I N + +++ PA ADDF+P LIY+ +K
Sbjct: 298 HIFNAI---KITKNEPASADDFLPTLIYIVLK 326
>RABX_MOUSE (Q9JM13) Rab5 GDP/GTP exchange factor (Rabex-5)
Length = 491
Score = 57.4 bits (137), Expect = 6e-08
Identities = 42/152 (27%), Positives = 78/152 (50%), Gaps = 15/152 (9%)
Query: 6 QDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAA-SPEDAKIDHEIS 64
QDF+ ++ ++ E ++ M+ +EK+IMT+L+ F + +D K D I
Sbjct: 183 QDFYQNVAERMQTR---GKVPPEKVEKIMDQIEKHIMTRLYKFVFCPETTDDEKKDLAIQ 239
Query: 65 EKISLL-----QTFLKPEHLDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCR 119
++I L Q P + +IP V A ++ ++++ + P++KL+ I C +
Sbjct: 240 KRIRALHWVTPQMLCVPVNEEIPEVSDMVVK---AITDIIEMDSKRVPRDKLACITRCSK 296
Query: 120 VINNLLLNAAMSEYVPAGADDFIPVLIYVTIK 151
I N + +++ PA ADDF+P LIY+ +K
Sbjct: 297 HIFNAI---KITKNEPASADDFLPTLIYIVLK 325
>RIN3_HUMAN (Q8TB24) Ras and Rab interactor 3 (Ras
interaction/interference protein 3)
Length = 984
Score = 41.6 bits (96), Expect = 0.003
Identities = 23/65 (35%), Positives = 40/65 (61%), Gaps = 5/65 (7%)
Query: 93 LAEKELQKINAFK---APQEKLSTIMNCCRVINNLLLNAAMSEYVPAGADDFIPVLIYVT 149
+ EK LQK + +P++K+S ++ C++I + + A + P GADDF+PVL+YV
Sbjct: 738 MMEKILQKFTSMHKAYSPEKKISILLKTCKLIYDSM--ALGNPGKPYGADDFLPVLMYVL 795
Query: 150 IKARL 154
++ L
Sbjct: 796 ARSNL 800
>RIN3_MOUSE (P59729) Ras and Rab interactor 3 (Ras
interaction/interference protein 3)
Length = 980
Score = 40.4 bits (93), Expect = 0.007
Identities = 23/63 (36%), Positives = 39/63 (61%), Gaps = 5/63 (7%)
Query: 95 EKELQKINAFK---APQEKLSTIMNCCRVINNLLLNAAMSEYVPAGADDFIPVLIYVTIK 151
EK LQK+ + +P +K+S ++ C++I + + A + P GADDF+PVL+YV +
Sbjct: 737 EKILQKLTSMHKAYSPGKKISILLKTCKLIYDSM--ALGNPGKPYGADDFLPVLMYVLAR 794
Query: 152 ARL 154
+ L
Sbjct: 795 SNL 797
>RIN2_MOUSE (Q9D684) Ras and Rab interactor 2 (Ras
interaction/interference protein 2)
Length = 903
Score = 35.8 bits (81), Expect = 0.18
Identities = 33/137 (24%), Positives = 63/137 (45%), Gaps = 16/137 (11%)
Query: 20 PLWATATEEDIDCAME-GLEKYIMTKLFSRTFAASPEDAKID---HEISEKISLLQTFLK 75
P+ + E+ ID +E + K I+ L A + D ++ E + L++
Sbjct: 586 PIESLIPEDQIDVVLEKAMHKCILKPLKGHVEAMLKDFHTADGSWKQLKENLQLVRQ-RN 644
Query: 76 PEHLDI----PPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRVINNLLLNAAMS 131
P+ L + P ++ E L + + +P++K+ ++ C++I ++ N +
Sbjct: 645 PQELGVFAPTPDLMELEKIKL----KFMTMQKMYSPEKKVMLLLRVCKLIYTVMENNSGR 700
Query: 132 EYVPAGADDFIPVLIYV 148
Y GADDF+PVL YV
Sbjct: 701 MY---GADDFLPVLTYV 714
>RIN2_HUMAN (Q8WYP3) Ras and Rab interactor 2 (Ras
interaction/interference protein 2) (Ras inhibitor
JC265) (Ras association domain family 4)
Length = 895
Score = 35.4 bits (80), Expect = 0.24
Identities = 30/133 (22%), Positives = 60/133 (44%), Gaps = 8/133 (6%)
Query: 20 PLWATATEEDIDCAME-GLEKYIMTKLFSRTFAASPEDAKID---HEISEKISLLQTFLK 75
P+ + E+ ID +E + K I+ L A + D ++ E + L++
Sbjct: 577 PIESLIPEDQIDVVLEKAMHKCILKPLKGHVEAMLKDFHMADGSWKQLKENLQLVRQ-RN 635
Query: 76 PEHLDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRVINNLLLNAAMSEYVP 135
P+ L + + + + + +P++K+ ++ C++I ++ N + Y
Sbjct: 636 PQELGVFAPTPDFVDVEKIKVKFMTMQKMYSPEKKVMLLLRVCKLIYTVMENNSGRMY-- 693
Query: 136 AGADDFIPVLIYV 148
GADDF+PVL YV
Sbjct: 694 -GADDFLPVLTYV 705
>TAO3_YEAST (P40468) Cell morphogenesis protein PAG1 (TAO3 protein)
Length = 2376
Score = 35.0 bits (79), Expect = 0.31
Identities = 26/116 (22%), Positives = 53/116 (45%), Gaps = 9/116 (7%)
Query: 82 PPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRVINNLLLNAAMSEYVPAGADDF 141
P + +++ L E+ K AF A ++ L +I CRV+N ++ A+ +E D
Sbjct: 323 PSIANSQLKSLENTIEVAKEEAFLADRKSLISIYILCRVLNEIVKQASSNE--EEDLSDK 380
Query: 142 IPVLIYVTIKARLALLVQTS-------NSLSCIEGKQNSSQKLSIISQILCQLRHL 190
+ +++ +K L + TS NS + + G + + LS+ + + L +
Sbjct: 381 LEEIVFTQLKTTDPLSISTSLIKSSNWNSFAELLGSMSEKKFLSVSDRFIADLEKI 436
>ALS2_MOUSE (Q920R0) Alsin (Amyotrophic lateral sclerosis protein 2
homolog)
Length = 1651
Score = 34.3 bits (77), Expect = 0.53
Identities = 21/67 (31%), Positives = 35/67 (51%), Gaps = 1/67 (1%)
Query: 88 EASWLLAEKELQKINAFKAPQEKLSTIMNCCRVINNLLLNAAMSEYVPAGADDFIPVLIY 147
+A + A + LQ+I+ P +KL I I+ +L A++ E DD PV +Y
Sbjct: 1543 DACFASAVECLQQISTTFTPSDKLKVIQQTFEEISQSVL-ASLQEDFLWSMDDLFPVFLY 1601
Query: 148 VTIKARL 154
V ++AR+
Sbjct: 1602 VVLRARI 1608
>ALS2_HUMAN (Q96Q42) Alsin (Amyotrophic lateral sclerosis protein 2)
Length = 1657
Score = 33.9 bits (76), Expect = 0.70
Identities = 21/67 (31%), Positives = 35/67 (51%), Gaps = 1/67 (1%)
Query: 88 EASWLLAEKELQKINAFKAPQEKLSTIMNCCRVINNLLLNAAMSEYVPAGADDFIPVLIY 147
+A + A + LQ+I+ P +KL I I+ +L A++ E DD PV +Y
Sbjct: 1549 DACFASAVECLQQISTTFTPSDKLKVIQQTFEEISQSVL-ASLHEDFLWSMDDLFPVFLY 1607
Query: 148 VTIKARL 154
V ++AR+
Sbjct: 1608 VVLRARI 1614
>UL43_EHV1B (P28959) Gene 17 membrane protein
Length = 401
Score = 31.2 bits (69), Expect = 4.5
Identities = 23/87 (26%), Positives = 37/87 (42%), Gaps = 2/87 (2%)
Query: 78 HLDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRVINNLLLNAAMSEYVPAG 137
H+D+ + N +L + ++ + P K+ TI+ CR I L A S +V
Sbjct: 59 HIDL--LTRNSTCLILMIISMYVLSLIRVPISKMETIVTVCRSIQALATLVAASVWVAGS 116
Query: 138 ADDFIPVLIYVTIKARLALLVQTSNSL 164
A +LI VT+ + T SL
Sbjct: 117 AVKKEHLLIVVTVCILFVFIAGTQISL 143
>S24B_ARATH (Q9M081) Putative protein transport protein Sec24-like
At4g32640
Length = 1069
Score = 30.4 bits (67), Expect = 7.7
Identities = 27/114 (23%), Positives = 47/114 (40%), Gaps = 5/114 (4%)
Query: 79 LDIPPVLHNEASWLLAEKE----LQKINAFKAPQEKLSTIMNCCRVINNLLLNAAMSEYV 134
L PP L+N+ W + + + ++ + Q + + C R+ ++ L A +
Sbjct: 728 LSDPPKLYNDLKWNITRPQGFEAVMRVRCSQGIQVQEYSGNFCKRIPTDIDLPAHDDKLQ 787
Query: 135 PAGADDFIPVLIYVTIKARLALLVQTSNSLSCIEGKQNSSQKLSIISQILCQLR 188
F L+Y TI + V T+ SLSC N + + SQ C L+
Sbjct: 788 DGAECAFQCALLYTTIYGERRIRV-TTLSLSCTNMLSNLFRAADLDSQFACMLK 840
>RPOC_PEDAC (P77917) DNA-directed RNA polymerase beta' chain (EC
2.7.7.6) (RNAP beta' subunit) (Transcriptase beta'
chain) (RNA polymerase beta' subunit) (Fragment)
Length = 1055
Score = 30.4 bits (67), Expect = 7.7
Identities = 21/61 (34%), Positives = 34/61 (55%), Gaps = 4/61 (6%)
Query: 30 IDCAMEGLEKYIMTKLFSRTFAASPEDA--KIDHEISEKISLLQTFLK--PEHLDIPPVL 85
+ AME + +IM +L SR A++ ++A KID + E +L+ +K P L+ P L
Sbjct: 359 VPMAMELFKPFIMKELVSRNLASNIKNAKRKIDRKDEEVYDVLEDVIKEHPVLLNRAPTL 418
Query: 86 H 86
H
Sbjct: 419 H 419
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.328 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,378,216
Number of Sequences: 164201
Number of extensions: 1472132
Number of successful extensions: 4458
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 4445
Number of HSP's gapped (non-prelim): 14
length of query: 378
length of database: 59,974,054
effective HSP length: 112
effective length of query: 266
effective length of database: 41,583,542
effective search space: 11061222172
effective search space used: 11061222172
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC147959.10


