

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147714.2 - phase: 0 /pseudo
(971 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
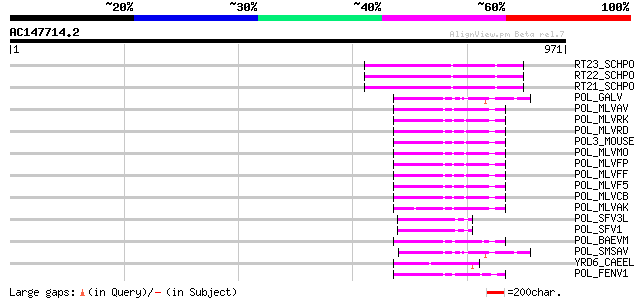
Score E
Sequences producing significant alignments: (bits) Value
RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protei... 137 1e-31
RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protei... 137 1e-31
RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protei... 137 1e-31
POL_GALV (P21414) Pol polyprotein [Contains: Protease (EC 3.4.23... 69 6e-11
POL_MLVAV (P03356) Pol polyprotein [Contains: Protease (EC 3.4.2... 66 4e-10
POL_MLVRK (P31795) Pol polyprotein [Contains: Protease (EC 3.4.2... 63 3e-09
POL_MLVRD (P11227) Pol polyprotein [Contains: Protease (EC 3.4.2... 63 3e-09
POL3_MOUSE (P11367) Retrovirus-related Pol polyprotein (Endonucl... 63 3e-09
POL_MLVMO (P03355) Pol polyprotein [Contains: Protease (EC 3.4.2... 62 6e-09
POL_MLVFP (P26808) Pol polyprotein [Contains: Protease (EC 3.4.2... 62 6e-09
POL_MLVFF (P26809) Pol polyprotein [Contains: Protease (EC 3.4.2... 62 6e-09
POL_MLVF5 (P26810) Pol polyprotein [Contains: Protease (EC 3.4.2... 62 6e-09
POL_MLVCB (P08361) Pol polyprotein [Contains: Reverse transcript... 62 6e-09
POL_MLVAK (P03357) Pol polyprotein [Contains: Reverse transcript... 62 6e-09
POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.2... 62 1e-08
POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23... 61 1e-08
POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC 3.4.2... 61 1e-08
POL_SMSAV (P03359) Pol polyprotein [Contains: Reverse transcript... 61 2e-08
YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II 59 5e-08
POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse transcript... 59 8e-08
>RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protein
type 3
Length = 1333
Score = 137 bits (346), Expect = 1e-31
Identities = 88/279 (31%), Positives = 137/279 (48%), Gaps = 7/279 (2%)
Query: 622 WDSISMIL*RVAKYX*RDXRIGGILXR*RKSAHFLXXNISFPVSQXXEIYIKEIVKCMGV 681
W+S+SM + ++ R K A + S Q ++ + ++ G
Sbjct: 984 WESLSMDFITALPESSGYNALFVVVDRFSKMAILVPCTKSITAEQTARMFDQRVIAYFGN 1043
Query: 682 PSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQ 741
P I++D D FTS+ WK ++ S Y PQTDGQ+ERT Q++E LLR
Sbjct: 1044 PKEIIADNDHIFTSQTWKDFAHKYNFVMKFSLPYRPQTDGQTERTNQTVEKLLRCVCSTH 1103
Query: 742 GGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQQTT 801
TW H+ L++ +YNN+ HS+ M PFE ++ R L E E Q+T
Sbjct: 1104 PNTWVDHISLVQQSYNNAIHSATQMTPFEIVH--RYSPALSPLELPSFSDKTDENSQETI 1161
Query: 802 EKVQMIQEKMKASQSRQKSYHDKRRKDL-EFQEGDHVFLRVTPMTGVXRALKSKKLTPKF 860
+ Q ++E + + + K Y D + +++ EFQ GD V ++ T TG KS KL P F
Sbjct: 1162 QVFQTVKEHLNTNNIKMKKYFDMKIQEIEEFQPGDLVMVKRT-KTGFLH--KSNKLAPSF 1218
Query: 861 IGPYQILERVGTVAYRVGLPPHLSNL-HNVFHVSQLRKY 898
GP+ +L++ G Y + LP + ++ + FHVS L KY
Sbjct: 1219 AGPFYVLQKSGPNNYELDLPDSIKHMFSSTFHVSHLEKY 1257
>RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protein
type 2
Length = 1333
Score = 137 bits (346), Expect = 1e-31
Identities = 88/279 (31%), Positives = 137/279 (48%), Gaps = 7/279 (2%)
Query: 622 WDSISMIL*RVAKYX*RDXRIGGILXR*RKSAHFLXXNISFPVSQXXEIYIKEIVKCMGV 681
W+S+SM + ++ R K A + S Q ++ + ++ G
Sbjct: 984 WESLSMDFITALPESSGYNALFVVVDRFSKMAILVPCTKSITAEQTARMFDQRVIAYFGN 1043
Query: 682 PSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQ 741
P I++D D FTS+ WK ++ S Y PQTDGQ+ERT Q++E LLR
Sbjct: 1044 PKEIIADNDHIFTSQTWKDFAHKYNFVMKFSLPYRPQTDGQTERTNQTVEKLLRCVCSTH 1103
Query: 742 GGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQQTT 801
TW H+ L++ +YNN+ HS+ M PFE ++ R L E E Q+T
Sbjct: 1104 PNTWVDHISLVQQSYNNAIHSATQMTPFEIVH--RYSPALSPLELPSFSDKTDENSQETI 1161
Query: 802 EKVQMIQEKMKASQSRQKSYHDKRRKDL-EFQEGDHVFLRVTPMTGVXRALKSKKLTPKF 860
+ Q ++E + + + K Y D + +++ EFQ GD V ++ T TG KS KL P F
Sbjct: 1162 QVFQTVKEHLNTNNIKMKKYFDMKIQEIEEFQPGDLVMVKRT-KTGFLH--KSNKLAPSF 1218
Query: 861 IGPYQILERVGTVAYRVGLPPHLSNL-HNVFHVSQLRKY 898
GP+ +L++ G Y + LP + ++ + FHVS L KY
Sbjct: 1219 AGPFYVLQKSGPNNYELDLPDSIKHMFSSTFHVSHLEKY 1257
>RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protein
type 1
Length = 1333
Score = 137 bits (346), Expect = 1e-31
Identities = 88/279 (31%), Positives = 137/279 (48%), Gaps = 7/279 (2%)
Query: 622 WDSISMIL*RVAKYX*RDXRIGGILXR*RKSAHFLXXNISFPVSQXXEIYIKEIVKCMGV 681
W+S+SM + ++ R K A + S Q ++ + ++ G
Sbjct: 984 WESLSMDFITALPESSGYNALFVVVDRFSKMAILVPCTKSITAEQTARMFDQRVIAYFGN 1043
Query: 682 PSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQ 741
P I++D D FTS+ WK ++ S Y PQTDGQ+ERT Q++E LLR
Sbjct: 1044 PKEIIADNDHIFTSQTWKDFAHKYNFVMKFSLPYRPQTDGQTERTNQTVEKLLRCVCSTH 1103
Query: 742 GGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQQTT 801
TW H+ L++ +YNN+ HS+ M PFE ++ R L E E Q+T
Sbjct: 1104 PNTWVDHISLVQQSYNNAIHSATQMTPFEIVH--RYSPALSPLELPSFSDKTDENSQETI 1161
Query: 802 EKVQMIQEKMKASQSRQKSYHDKRRKDL-EFQEGDHVFLRVTPMTGVXRALKSKKLTPKF 860
+ Q ++E + + + K Y D + +++ EFQ GD V ++ T TG KS KL P F
Sbjct: 1162 QVFQTVKEHLNTNNIKMKKYFDMKIQEIEEFQPGDLVMVKRT-KTGFLH--KSNKLAPSF 1218
Query: 861 IGPYQILERVGTVAYRVGLPPHLSNL-HNVFHVSQLRKY 898
GP+ +L++ G Y + LP + ++ + FHVS L KY
Sbjct: 1219 AGPFYVLQKSGPNNYELDLPDSIKHMFSSTFHVSHLEKY 1257
>POL_GALV (P21414) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1165
Score = 68.9 bits (167), Expect = 6e-11
Identities = 66/249 (26%), Positives = 107/249 (42%), Gaps = 35/249 (14%)
Query: 672 IKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLE 731
++EI+ G+P + SD P F ++ + L LG +L AY PQ+ GQ ER ++++
Sbjct: 926 LEEILPRFGIPKVLGSDNGPAFVAQVSQGLATQLGINWKLHCAYRPQSSGQVERMNRTIK 985
Query: 732 DLLRICVLEQGG-TWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERV 790
+ L LE GG W + LPL N+ G+ P+E LYG P ESGE
Sbjct: 986 ETLTKLALETGGKDWVTLLPLALLRARNT-PGRFGLTPYEILYG----GPPPILESGE-- 1038
Query: 791 VLGPEIVQQTTEKVQMIQEKMKASQSRQKSYHDKRRKDLE---------FQEGDHVFLRV 841
LGP+ + ++ +KA + + D+ ++ + FQ GD V +
Sbjct: 1039 TLGPD-----DRFLPVLFTHLKALEIVRTQIWDQIKEVYKPGTVTIPHPFQVGDQVLV-- 1091
Query: 842 TPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPD 901
R + L P++ GPY +L T G+ + + H+ PD
Sbjct: 1092 -------RRHRPSSLEPRWKGPYLVLLTTPTAVKVDGIAAWV----HASHLKPAPPSAPD 1140
Query: 902 PSHVIQSDD 910
S ++ D
Sbjct: 1141 ESWELEKTD 1149
>POL_MLVAV (P03356) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1196
Score = 66.2 bits (160), Expect = 4e-10
Identities = 57/199 (28%), Positives = 93/199 (46%), Gaps = 15/199 (7%)
Query: 672 IKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLE 731
++EI G+P + SD P FTS+ +S+ + LG +L AY PQ+ GQ ER ++++
Sbjct: 959 LEEIFPRFGMPQVLGSDNGPAFTSQVSQSVADLLGIDWKLHCAYRPQSSGQVERMNRTIK 1018
Query: 732 DLLRICVLEQG-GTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERV 790
+ L L G W LPL + N+ G+ P+E LYG PL F +
Sbjct: 1019 ETLTKLTLAAGTRDWVLLLPLALYRARNT-PGPHGLTPYEILYG--APPPLVNFHDPDMS 1075
Query: 791 VL-GPEIVQQTTEKVQMIQEKM-KASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGVX 848
L +Q + +Q +Q ++ K + D+ F+ GD V++
Sbjct: 1076 ELTNSPSLQAHLQALQTVQREIWKPLAEAYRDQLDQPVIPHPFRIGDSVWV--------- 1126
Query: 849 RALKSKKLTPKFIGPYQIL 867
R ++K L P++ GPY +L
Sbjct: 1127 RRHQTKNLEPRWKGPYTVL 1145
>POL_MLVRK (P31795) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)] (Fragment)
Length = 581
Score = 63.2 bits (152), Expect = 3e-09
Identities = 57/200 (28%), Positives = 93/200 (46%), Gaps = 17/200 (8%)
Query: 672 IKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLE 731
++EI G+P + +D P F S+ +S+ + LG +L AY PQ+ GQ ER ++++
Sbjct: 344 LEEIFPRFGMPQVLGTDNGPAFVSQVSQSVAKLLGIDWKLHCAYRPQSSGQVERMNRTIK 403
Query: 732 DLLRICVLEQG-GTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGE-- 788
+ L L G W LPL + N+ G+ P+E LYG PL F E
Sbjct: 404 ETLTKLTLATGTRDWVLLLPLALYRARNT-PGPHGLTPYEILYG--APPPLVNFHDPEMS 460
Query: 789 RVVLGPEIVQQTTEKVQMIQ-EKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGV 847
+ P + Q + +Q +Q E K + + D+ F+ GD V++
Sbjct: 461 KFTNSPSL-QAHLQALQAVQREVWKPLAAAYQDQLDQPVIPHPFRVGDTVWV-------- 511
Query: 848 XRALKSKKLTPKFIGPYQIL 867
R ++K L P++ GPY +L
Sbjct: 512 -RRHQTKNLEPRWKGPYTVL 530
>POL_MLVRD (P11227) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1196
Score = 63.2 bits (152), Expect = 3e-09
Identities = 57/200 (28%), Positives = 93/200 (46%), Gaps = 17/200 (8%)
Query: 672 IKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLE 731
++EI G+P + +D P F S+ +S+ + LG +L AY PQ+ GQ ER ++++
Sbjct: 959 LEEIFPRFGMPQVLGTDNGPAFVSQVSQSVAKLLGIDWKLHCAYRPQSSGQVERMNRTIK 1018
Query: 732 DLLRICVLEQG-GTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGE-- 788
+ L L G W LPL + N+ G+ P+E LYG PL F E
Sbjct: 1019 ETLTKLTLATGTRDWVLLLPLALYRARNT-PGPHGLTPYEILYG--APPPLVNFHDPEMS 1075
Query: 789 RVVLGPEIVQQTTEKVQMIQ-EKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGV 847
+ P + Q + +Q +Q E K + + D+ F+ GD V++
Sbjct: 1076 KFTNSPSL-QAHLQALQAVQREVWKPLAAAYQDQLDQPVIPHPFRVGDTVWV-------- 1126
Query: 848 XRALKSKKLTPKFIGPYQIL 867
R ++K L P++ GPY +L
Sbjct: 1127 -RRHQTKNLEPRWKGPYTVL 1145
>POL3_MOUSE (P11367) Retrovirus-related Pol polyprotein
(Endonuclease) (Fragment)
Length = 390
Score = 63.2 bits (152), Expect = 3e-09
Identities = 57/200 (28%), Positives = 93/200 (46%), Gaps = 17/200 (8%)
Query: 672 IKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLE 731
++EI G+P + +D P F S+ +S+ + LG +L AY PQ+ GQ ER ++++
Sbjct: 168 LEEIFPRFGMPQVLGTDNGPAFVSQVSQSVAKLLGIDWKLHCAYRPQSSGQVERMNRTIK 227
Query: 732 DLLRICVLEQG-GTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGE-- 788
+ L L G W LPL + N+ G+ P+E LYG PL F E
Sbjct: 228 ETLTKLTLATGTRDWVLLLPLALYRARNT-PGPHGLTPYEILYG--APPPLVNFHDPEMS 284
Query: 789 RVVLGPEIVQQTTEKVQMIQ-EKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGV 847
+ P + Q + +Q +Q E K + + D+ F+ GD V++
Sbjct: 285 KFTNSPSL-QAHLQALQAVQREVWKPLAAAYQDQLDQPVIPHPFRVGDTVWV-------- 335
Query: 848 XRALKSKKLTPKFIGPYQIL 867
R ++K L P++ GPY +L
Sbjct: 336 -RRHQTKNLEPRWKGPYTVL 354
>POL_MLVMO (P03355) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1199
Score = 62.4 bits (150), Expect = 6e-09
Identities = 54/200 (27%), Positives = 94/200 (47%), Gaps = 17/200 (8%)
Query: 672 IKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLE 731
++EI G+P + +D P F S+ +++ + LG +L AY PQ+ GQ ER ++++
Sbjct: 959 LEEIFPRFGMPQVLGTDNGPAFVSKVSQTVADLLGIDWKLHCAYRPQSSGQVERMNRTIK 1018
Query: 732 DLLRICVLEQGG-TWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGE-- 788
+ L L G W LPL + N+ G+ P+E LYG PL F +
Sbjct: 1019 ETLTKLTLATGSRDWVLLLPLALYRARNT-PGPHGLTPYEILYG--APPPLVNFPDPDMT 1075
Query: 789 RVVLGPEIVQQTTEKVQMIQ-EKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGV 847
RV P + Q + + ++Q E + + + D+ ++ GD V++
Sbjct: 1076 RVTNSPSL-QAHLQALYLVQHEVWRPLAAAYQEQLDRPVVPHPYRVGDTVWV-------- 1126
Query: 848 XRALKSKKLTPKFIGPYQIL 867
R ++K L P++ GPY +L
Sbjct: 1127 -RRHQTKNLEPRWKGPYTVL 1145
>POL_MLVFP (P26808) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1204
Score = 62.4 bits (150), Expect = 6e-09
Identities = 54/200 (27%), Positives = 94/200 (47%), Gaps = 17/200 (8%)
Query: 672 IKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLE 731
++EI G+P + +D P F S+ +++ + LG +L AY PQ+ GQ ER ++++
Sbjct: 964 LEEIFPRFGMPQVLGTDNGPAFVSKVSQTVADLLGVDWKLHCAYRPQSSGQVERMNRTIK 1023
Query: 732 DLLRICVLEQGG-TWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGE-- 788
+ L L G W LPL + N+ G+ P+E LYG PL F +
Sbjct: 1024 ETLTKLTLATGSRDWVLLLPLALYRARNT-PGPHGLTPYEILYG--APPPLVNFPDPDMA 1080
Query: 789 RVVLGPEIVQQTTEKVQMIQ-EKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGV 847
+V P + Q + + ++Q E + + + D+ F+ GD V++
Sbjct: 1081 KVTHNPSL-QAHLQALYLVQHEVWRPLAAAYQEQLDRPVVPHPFRVGDTVWV-------- 1131
Query: 848 XRALKSKKLTPKFIGPYQIL 867
R ++K L P++ GPY +L
Sbjct: 1132 -RRHQTKNLEPRWKGPYTVL 1150
>POL_MLVFF (P26809) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1204
Score = 62.4 bits (150), Expect = 6e-09
Identities = 54/200 (27%), Positives = 94/200 (47%), Gaps = 17/200 (8%)
Query: 672 IKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLE 731
++EI G+P + +D P F S+ +++ + LG +L AY PQ+ GQ ER ++++
Sbjct: 964 LEEIFPRFGMPQVLGTDNGPAFVSKVSQTVADLLGVDWKLHCAYRPQSSGQVERMNRTIK 1023
Query: 732 DLLRICVLEQGG-TWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGE-- 788
+ L L G W LPL + N+ G+ P+E LYG PL F +
Sbjct: 1024 ETLTKLTLATGSRDWVLLLPLALYRARNT-PGPHGLTPYEILYG--APPPLVNFPDPDMA 1080
Query: 789 RVVLGPEIVQQTTEKVQMIQ-EKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGV 847
+V P + Q + + ++Q E + + + D+ F+ GD V++
Sbjct: 1081 KVTHNPSL-QAHLQALYLVQHEVWRPLAAAYQEQLDRPVVPHPFRVGDTVWV-------- 1131
Query: 848 XRALKSKKLTPKFIGPYQIL 867
R ++K L P++ GPY +L
Sbjct: 1132 -RRHQTKNLEPRWKGPYTVL 1150
>POL_MLVF5 (P26810) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1204
Score = 62.4 bits (150), Expect = 6e-09
Identities = 54/200 (27%), Positives = 94/200 (47%), Gaps = 17/200 (8%)
Query: 672 IKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLE 731
++EI G+P + +D P F S+ +++ + LG +L AY PQ+ GQ ER ++++
Sbjct: 964 LEEIFPRFGMPQVLGTDNGPAFVSKVSQTVADLLGVDWKLHCAYRPQSSGQVERMNRTIK 1023
Query: 732 DLLRICVLEQGG-TWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGE-- 788
+ L L G W LPL + N+ G+ P+E LYG PL F +
Sbjct: 1024 ETLTKLTLATGSRDWVLLLPLALYRARNT-PGPHGLTPYEILYG--APPPLVNFPDPDMA 1080
Query: 789 RVVLGPEIVQQTTEKVQMIQ-EKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGV 847
+V P + Q + + ++Q E + + + D+ F+ GD V++
Sbjct: 1081 KVTHNPSL-QAHLQALYLVQHEVWRPLAAAYQEQLDRPVVPHPFRVGDTVWV-------- 1131
Query: 848 XRALKSKKLTPKFIGPYQIL 867
R ++K L P++ GPY +L
Sbjct: 1132 -RRHQTKNLEPRWKGPYTVL 1150
>POL_MLVCB (P08361) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 282
Score = 62.4 bits (150), Expect = 6e-09
Identities = 54/200 (27%), Positives = 94/200 (47%), Gaps = 17/200 (8%)
Query: 672 IKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLE 731
++EI G+P + +D P F S+ +++ + LG +L AY PQ+ GQ ER ++++
Sbjct: 45 LEEIFPRFGMPQVLGTDNGPAFVSKVSQTVADLLGIDWKLHCAYRPQSSGQVERMNRTIK 104
Query: 732 DLLRICVLEQGG-TWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGE-- 788
+ L L G W LPL + N+ G+ P+E LYG PL F +
Sbjct: 105 ETLTKLTLATGSRDWVLLLPLALYRARNT-PGPHGLTPYEILYG--APPPLVNFPDPDMT 161
Query: 789 RVVLGPEIVQQTTEKVQMIQ-EKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGV 847
RV P + Q + + ++Q E + + + D+ ++ GD V++
Sbjct: 162 RVTNSPSL-QAHLQALYLVQHEVWRPLAAAYQEQLDRPVVPHPYRVGDTVWV-------- 212
Query: 848 XRALKSKKLTPKFIGPYQIL 867
R ++K L P++ GPY +L
Sbjct: 213 -RRHQTKNLEPRWKGPYTVL 231
>POL_MLVAK (P03357) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 843
Score = 62.4 bits (150), Expect = 6e-09
Identities = 57/199 (28%), Positives = 93/199 (46%), Gaps = 16/199 (8%)
Query: 672 IKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLE 731
++EI G+P + SD P FTS+ +S+ + LG +L AY PQ+ GQ ER ++++
Sbjct: 607 LEEIFPRFGMPQVLGSDNGPAFTSQVSQSVADLLGID-KLHCAYRPQSSGQVERMNRTIK 665
Query: 732 DLLRICVLEQG-GTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERV 790
+ L L G W LPL + N+ G+ P+E LYG PL F +
Sbjct: 666 ETLTKLTLAAGTRDWVLLLPLALYRARNT-PGPHGLTPYEILYG--APPPLVNFHDPDMS 722
Query: 791 VL-GPEIVQQTTEKVQMIQEKM-KASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGVX 848
L +Q + +Q +Q ++ K + D+ F+ GD V++
Sbjct: 723 ELTNSPSLQAHLQALQTVQREIWKPLAEAYRDQLDQPVIPHPFRIGDSVWV--------- 773
Query: 849 RALKSKKLTPKFIGPYQIL 867
R ++K L P++ GPY +L
Sbjct: 774 RRHQTKNLEPRWKGPYTVL 792
>POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1157
Score = 61.6 bits (148), Expect = 1e-08
Identities = 39/131 (29%), Positives = 64/131 (48%), Gaps = 9/131 (6%)
Query: 679 MGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICV 738
+ VP I SD+ FTS + + G +L S+ YHPQ+ G+ ER ++ LL +
Sbjct: 940 IAVPKVIHSDQGAAFTSATFADWAKNKGIQLEFSTPYHPQSSGKVERKNSDIKRLLTKLL 999
Query: 739 LEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQ 798
+ + W LP+++ NNSY S P + L+G TP F + + + L E
Sbjct: 1000 VGRPAKWYDLLPVVQLALNNSYSPSSKYTPHQLLFGIDSNTP---FANSDTLDLSRE--- 1053
Query: 799 QTTEKVQMIQE 809
E++ ++QE
Sbjct: 1054 ---EELSLLQE 1061
>POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1161
Score = 61.2 bits (147), Expect = 1e-08
Identities = 37/131 (28%), Positives = 64/131 (48%), Gaps = 9/131 (6%)
Query: 679 MGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICV 738
+ +P + SD+ FTS + + G +L S+ YHPQ+ G+ ER ++ LL +
Sbjct: 938 IAIPKVLHSDQGAAFTSSTFADWAKEKGIQLEFSTPYHPQSSGKVERKNSDIKRLLTKLL 997
Query: 739 LEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQ 798
+ + W LP+++ NNSY S P + L+G TP F + + + L E
Sbjct: 998 IGRPAKWYDLLPVVQLALNNSYSPSSKYTPHQLLFGVDSNTP---FANSDTLDLSRE--- 1051
Query: 799 QTTEKVQMIQE 809
E++ ++QE
Sbjct: 1052 ---EELSLLQE 1059
>POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1189
Score = 61.2 bits (147), Expect = 1e-08
Identities = 56/198 (28%), Positives = 89/198 (44%), Gaps = 16/198 (8%)
Query: 672 IKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLE 731
++EI G+P I SD P F S+ + L LG +L AY PQ+ GQ ER ++++
Sbjct: 954 LEEIFPRFGLPKVIGSDNGPAFVSQVSQGLARILGINWKLHCAYRPQSSGQVERMNRTIK 1013
Query: 732 DLLRICVLEQG-GTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERV 790
+ L LE G W L L N+ + G+ P+E LYG PL +
Sbjct: 1014 ETLTKLTLETGLKDWRRLLSLALLRARNT-PNRFGLTPYEILYGG--PPPLSTLLNSFSP 1070
Query: 791 VLGPEIVQQTTEKVQMIQEKMKASQSR-QKSYHDKRRKDLEFQEGDHVFLRVTPMTGVXR 849
+Q + +Q +Q ++ A + + H + FQ GD V++ R
Sbjct: 1071 SNSKTDLQARLKGLQAVQAQIWAPLAELYRPGHSQTSH--PFQVGDSVYV---------R 1119
Query: 850 ALKSKKLTPKFIGPYQIL 867
+S+ L P++ GPY +L
Sbjct: 1120 RHRSQGLEPRWKGPYIVL 1137
>POL_SMSAV (P03359) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 294
Score = 60.8 bits (146), Expect = 2e-08
Identities = 63/240 (26%), Positives = 101/240 (41%), Gaps = 35/240 (14%)
Query: 681 VPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLE 740
+P + SD P F ++ + L LG +L AY PQ+ GQ ER +++++ L LE
Sbjct: 64 IPKVLGSDNGPAFVAQVSQGLATQLGINWKLHCAYRPQSSGQVERMNRTIKETLTKLALE 123
Query: 741 QGG-TWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQQ 799
GG W + LPL N+ S G+ P+E LYG P ESG LGP+
Sbjct: 124 TGGKDWVALLPLALLRAKNT-PSRFGLTPYEILYG----GPPPILESGG--TLGPD---- 172
Query: 800 TTEKVQMIQEKMKASQSRQKSYHDKRRKDLE---------FQEGDHVFLRVTPMTGVXRA 850
+ ++ +KA + + D+ ++ + FQ GD V + R
Sbjct: 173 -DNFLPVLFTHLKALEVVRTQIWDQIKEVYKPGTVAIPHPFQVGDQVLV---------RR 222
Query: 851 LKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDD 910
+ L P++ GPY +L T G+ + + H+ PD S ++ D
Sbjct: 223 HRPGSLEPRWKGPYLVLLTTPTAVKVDGIAAWV----HASHLKPAPPSAPDESWELEKAD 278
>YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II
Length = 1268
Score = 59.3 bits (142), Expect = 5e-08
Identities = 40/155 (25%), Positives = 76/155 (48%), Gaps = 6/155 (3%)
Query: 672 IKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLE 731
++EI G P +I+SD + TS + + ++ G + + S+ Y+P+++G +ER + +L+
Sbjct: 893 LEEIFSIHGYPETIISDNGTQLTSHLFAQMCQSHGIEHKTSAVYYPRSNGAAERFVDTLK 952
Query: 732 DLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSI-GMAPFEALYGRRCRTPLCWFESGERV 790
+ + +G L +Y N+ HS++ G P E +GR+ RT + +RV
Sbjct: 953 RGI-AKIKGEGSVNQQILNKFLISYRNTPHSALNGSTPAECHFGRKIRTTMSLLMPTDRV 1011
Query: 791 VLGPEIVQQTTEKVQMIQ----EKMKASQSRQKSY 821
+ P++ Q + + KA Q QK Y
Sbjct: 1012 LKVPKLTQYQQNMKHHYELRNGARAKAFQVNQKVY 1046
>POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 1046
Score = 58.5 bits (140), Expect = 8e-08
Identities = 55/198 (27%), Positives = 88/198 (43%), Gaps = 16/198 (8%)
Query: 672 IKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLE 731
++EI G+P I SD P F S+ + L LG +L AY PQ+ GQ ER ++++
Sbjct: 811 LEEIFPRFGLPKVIGSDNGPAFVSQVSQGLARTLGINWKLHCAYRPQSSGQVERMNRTIK 870
Query: 732 DLLRICVLEQG-GTWDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERV 790
+ L LE G W L L N+ + G+ P+E LYG PL +
Sbjct: 871 ETLTKLTLETGLKDWRRLLSLALLRARNT-PNRFGLTPYEILYGG--PPPLSTLLNSFSP 927
Query: 791 VLGPEIVQQTTEKVQMIQEKMKASQSR-QKSYHDKRRKDLEFQEGDHVFLRVTPMTGVXR 849
+Q + +Q +Q ++ + + H + FQ GD V++R
Sbjct: 928 SDPKTDLQARLKGLQAVQAQIWTPLAELYRPGHP--QTSYPFQVGDSVYVRWH------- 978
Query: 850 ALKSKKLTPKFIGPYQIL 867
+S+ L P++ GPY +L
Sbjct: 979 --RSQGLEPRWKGPYIVL 994
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.350 0.155 0.524
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 88,881,453
Number of Sequences: 164201
Number of extensions: 3146247
Number of successful extensions: 12359
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 30
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 12288
Number of HSP's gapped (non-prelim): 53
length of query: 971
length of database: 59,974,054
effective HSP length: 120
effective length of query: 851
effective length of database: 40,269,934
effective search space: 34269713834
effective search space used: 34269713834
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 71 (32.0 bits)
Medicago: description of AC147714.2


