

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147714.16 - phase: 0
(322 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
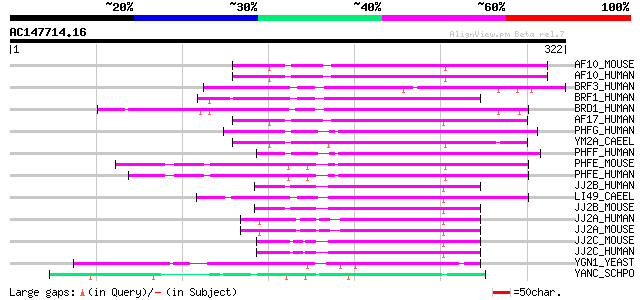
Score E
Sequences producing significant alignments: (bits) Value
AF10_MOUSE (O54826) AF-10 protein 137 4e-32
AF10_HUMAN (P55197) AF-10 protein 137 4e-32
BRF3_HUMAN (Q9ULD4) Bromodomain and PHD finger-containing protei... 137 5e-32
BRF1_HUMAN (P55201) Peregrin (Bromodomain and PHD finger-contain... 133 5e-31
BRD1_HUMAN (O95696) Bromodomain-containing protein 1 (BR140-like... 132 9e-31
AF17_HUMAN (P55198) AF-17 protein 132 2e-30
PHFG_HUMAN (Q92613) PHD finger protein 16 124 2e-28
YM2A_CAEEL (P34447) Hypothetical protein F54F2.2 in chromosome I... 119 1e-26
PHFF_HUMAN (Q9NQC1) PHD finger protein 15 108 2e-23
PHFE_MOUSE (Q9D4H9) PHD finger protein 14 106 7e-23
PHFE_HUMAN (O94880) PHD finger protein 14 105 2e-22
JJ2B_HUMAN (O94953) Jumonji domain containing protein 2B 102 1e-21
LI49_CAEEL (Q20318) Protein lin-49 (Abnormal cell lineage protei... 102 2e-21
JJ2B_MOUSE (Q91VY5) Jumonji domain containing protein 2B 101 2e-21
JJ2A_HUMAN (O75164) Jumonji domain containing protein 2A 99 2e-20
JJ2A_MOUSE (Q8BW72) Jumonji domain containing protein 2A (Fragment) 97 7e-20
JJ2C_MOUSE (Q8VCD7) Jumonji domain containing protein 2C 94 5e-19
JJ2C_HUMAN (Q9H3R0) Jumonji domain containing protein 2C (Gene a... 94 5e-19
YGN1_YEAST (P53127) Hypothetical 163.2 kDa protein in RPL1B-CEG1... 76 1e-13
YANC_SCHPO (Q10077) Hypothetical protein C3H1.12c in chromosome I 69 2e-11
>AF10_MOUSE (O54826) AF-10 protein
Length = 1068
Score = 137 bits (345), Expect = 4e-32
Identities = 68/193 (35%), Positives = 101/193 (52%), Gaps = 17/193 (8%)
Query: 130 CCVCHSTDANAEDPIVFCDG--CNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDT 187
CCVC AE+P+V+CDG C++ VH +CYG + Q+P G WFC +C +
Sbjct: 25 CCVCSDERGWAENPLVYCDGHGCSVAVHQACYG---IVQVPTGPWFCRKCESQER----A 77
Query: 188 GPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKC 247
+RC LCP K+GA+K+T +G W H+VCAL +PEV F + E I +P R+ + C
Sbjct: 78 ARVRCELCPHKDGALKRTDNGGWAHVVCALYIPEVQFANVSTMEPIVLQSVPHDRYNKTC 137
Query: 248 YVCG-------CFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATVVAGFCKTH 300
Y+C G + C++ C FH+TC L E + G+CK H
Sbjct: 138 YICDEQGRESKAATGACMTCNKHGCRQAFHVTCAQFAGLLCEEEGNGADNVQYCGYCKYH 197
Query: 301 -SQIWEKNKGSGK 312
S++ + +GS +
Sbjct: 198 FSKLKKSKRGSNR 210
>AF10_HUMAN (P55197) AF-10 protein
Length = 1027
Score = 137 bits (345), Expect = 4e-32
Identities = 68/193 (35%), Positives = 101/193 (52%), Gaps = 17/193 (8%)
Query: 130 CCVCHSTDANAEDPIVFCDG--CNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDT 187
CCVC AE+P+V+CDG C++ VH +CYG + Q+P G WFC +C +
Sbjct: 25 CCVCSDERGWAENPLVYCDGHGCSVAVHQACYG---IVQVPTGPWFCRKCESQER----A 77
Query: 188 GPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKC 247
+RC LCP K+GA+K+T +G W H+VCAL +PEV F + E I +P R+ + C
Sbjct: 78 ARVRCELCPHKDGALKRTDNGGWAHVVCALYIPEVQFANVSTMEPIVLQSVPHDRYNKTC 137
Query: 248 YVCG-------CFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATVVAGFCKTH 300
Y+C G + C++ C FH+TC L E + G+CK H
Sbjct: 138 YICDEQGRESKAATGACMTCNKHGCRQAFHVTCAQFAGLLCEEEGNGADNVQYCGYCKYH 197
Query: 301 -SQIWEKNKGSGK 312
S++ + +GS +
Sbjct: 198 FSKLKKSKRGSNR 210
>BRF3_HUMAN (Q9ULD4) Bromodomain and PHD finger-containing protein 3
(Fragment)
Length = 1214
Score = 137 bits (344), Expect = 5e-32
Identities = 78/224 (34%), Positives = 111/224 (48%), Gaps = 24/224 (10%)
Query: 113 QSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDW 172
+S+ + + D++ CCVC + + + I+FCD CNL VH CYG P IP+G W
Sbjct: 207 ESRSSGAQQSLIDEDAFCCVCLDDECHNSNVILFCDICNLAVHQECYGVPY---IPEGQW 263
Query: 173 FCDQCRFKNDIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDP---EG 229
C C + + + P+ C LCP K GA KQT+DG W H+VCA+ +PEV F + E
Sbjct: 264 LCRCC-----LQSPSRPVDCILCPNKGGAFKQTSDGHWAHVVCAIWIPEVCFANTVFLEP 318
Query: 230 REGIDCSKIPKKRWLEKCYVCGCFD-GCALVCSEQKCGLGFHITCGIKEDLCIE---YKE 285
EGID IP RW CY+C G A+ C + C FH+TC + L ++ +E
Sbjct: 319 IEGID--NIPPARWKLTCYICKQKGLGAAIQCHKVNCYTAFHVTCAQRAGLFMKIEPMRE 376
Query: 286 GKKGATVV----AGFCKTHS---QIWEKNKGSGKYKIVAVEDKE 322
T+ +C+ HS + KG I D+E
Sbjct: 377 TSLNGTIFTVRKTAYCEAHSPPGAATARRKGDSPRSISETGDEE 420
>BRF1_HUMAN (P55201) Peregrin (Bromodomain and PHD finger-containing
protein 1) (BR140 protein)
Length = 1214
Score = 133 bits (335), Expect = 5e-31
Identities = 69/171 (40%), Positives = 93/171 (54%), Gaps = 16/171 (9%)
Query: 110 IEKQS-----QKQDIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLV 164
+EK+S K D VD+D + CC+C+ + + I+FCD CNL VH CYG P
Sbjct: 252 LEKESYFESHNKGDPNALVDEDAV-CCICNDGECQNSNVILFCDMCNLEVHQECYGVPY- 309
Query: 165 KQIPDGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFF 224
IP+G W C +C + + + + C+LCP K GA KQT DG+W H+VCAL +PEV F
Sbjct: 310 --IPEGQWLCRRC-----LQSPSRAVDCALCPNKGGAFKQTDDGRWAHVVCALWIPEVCF 362
Query: 225 VDPEGREGID-CSKIPKKRWLEKCYVC-GCFDGCALVCSEQKCGLGFHITC 273
+ E ID IP RW CY+C G + C + C FH+TC
Sbjct: 363 ANTVFLEPIDSIEHIPPARWKLTCYICKQRGSGACIQCHKANCYTAFHVTC 413
>BRD1_HUMAN (O95696) Bromodomain-containing protein 1 (BR140-like
protein)
Length = 1058
Score = 132 bits (333), Expect = 9e-31
Identities = 87/272 (31%), Positives = 125/272 (44%), Gaps = 31/272 (11%)
Query: 52 HTPPSSPTPPPSTYSLPTKKRITALQPHLHHHNNIPNDAVPLIDLNVEYSPSLPSATP-- 109
++PPS+P PP Y +K L + + + + A I +P+ +
Sbjct: 127 YSPPSAPRRPPVYYKF-IEKSAEELDNEVEYDMDEEDYAWLEIVNEKRKGDCVPAVSQSM 185
Query: 110 -------IEKQS----QKQDIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASC 158
EK+S QKQ + + D++ +CC+C + + I+FCD CNL VH C
Sbjct: 186 FEFLMDRFEKESHCENQKQGEQQSLIDEDAVCCICMDGECQNSNVILFCDMCNLAVHQEC 245
Query: 159 YGNPLVKQIPDGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALL 218
YG P IP+G W C C + + P C LCP K GA K+T D +W H+VCAL
Sbjct: 246 YGVP---YIPEGQWLCRHC-----LQSRARPADCVLCPNKGGAFKKTDDDRWGHVVCALW 297
Query: 219 VPEVFFVDPEGREGID-CSKIPKKRWLEKCYVCGCFD-GCALVCSEQKCGLGFHITCGIK 276
+PEV F + E ID IP RW CY+C G + C + C FH+TC K
Sbjct: 298 IPEVGFANTVFIEPIDGVRNIPPARWKLTCYLCKQKGVGACIQCHKANCYTAFHVTCAQK 357
Query: 277 EDLCIE---YKEGKKGATVVA----GFCKTHS 301
L ++ KE G T + +C H+
Sbjct: 358 AGLYMKMEPVKELTGGGTTFSVRKTAYCDVHT 389
>AF17_HUMAN (P55198) AF-17 protein
Length = 1093
Score = 132 bits (331), Expect = 2e-30
Identities = 65/180 (36%), Positives = 93/180 (51%), Gaps = 16/180 (8%)
Query: 130 CCVCHSTDANAEDPIVFCDG--CNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDT 187
CCVC AE+P+V+CDG C++ VH +CYG + Q+P G WFC +C +
Sbjct: 8 CCVCSDERGWAENPLVYCDGHACSVAVHQACYG---IVQVPTGPWFCRKCESQER----A 60
Query: 188 GPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKC 247
+RC LCP K+GA+K+T +G W H+VCAL +PEV F + E I +P R+ + C
Sbjct: 61 ARVRCELCPHKDGALKRTDNGGWAHVVCALYIPEVQFANVLTMEPIVLQYVPHDRFNKTC 120
Query: 248 YVC-------GCFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATVVAGFCKTH 300
Y+C G + C+ C FH+TC L E + + G+CK H
Sbjct: 121 YICEETGRESKAASGACMTCNRHGCRQAFHVTCAQMAGLLCEEEVLEVDNVKYCGYCKYH 180
>PHFG_HUMAN (Q92613) PHD finger protein 16
Length = 823
Score = 124 bits (312), Expect = 2e-28
Identities = 67/184 (36%), Positives = 94/184 (50%), Gaps = 12/184 (6%)
Query: 125 DDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDID 184
D++++C VC S D+ + +VFCD CN+ VH +CYG + ++P+G W C C
Sbjct: 198 DEDVICDVCRSPDSEEGNDMVFCDKCNVCVHQACYG---ILKVPEGSWLCRSCVL----- 249
Query: 185 TDTGPIRCSLCPTKEGAMKQTTDG-KWVHLVCALLVPEVFFVDPEGREGI-DCSKIPKKR 242
P +C LCP K GA+K T G KW H+ CAL +PEV PE E I S IP R
Sbjct: 250 -GIYP-QCVLCPKKGGALKTTKTGTKWAHVSCALWIPEVSIACPERMEPITKISHIPPSR 307
Query: 243 WLEKCYVCGCFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATVVAGFCKTHSQ 302
W C +C G + CS + C FH+TC + L ++ + +C HSQ
Sbjct: 308 WALVCNLCKLKTGACIQCSIKSCITAFHVTCAFEHGLEMKTILDEGDEVKFKSYCLKHSQ 367
Query: 303 IWEK 306
+K
Sbjct: 368 NRQK 371
>YM2A_CAEEL (P34447) Hypothetical protein F54F2.2 in chromosome III,
isoform a
Length = 867
Score = 119 bits (298), Expect = 1e-26
Identities = 57/183 (31%), Positives = 95/183 (51%), Gaps = 17/183 (9%)
Query: 130 CCVCHSTDANAEDPIVFCDG--CNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDI---D 184
CCVC + ++P+++CDG C + VH CYG ++++P+G+WFC +C + +
Sbjct: 8 CCVCADENGWTDNPLIYCDGENCEVAVHQGCYG---IQEVPEGEWFCAKCTKASAMMPGS 64
Query: 185 TDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWL 244
+ C LCP GA+K+T W H++CAL +PEV F + E + + +P ++
Sbjct: 65 INEATFCCQLCPFDYGALKKTDRNGWAHVICALYIPEVRFGNVHSMEPVILNDVPTDKFN 124
Query: 245 EKCYVCG------CFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATV-VAGFC 297
+ CY+C G + C++ C FH+TC ++ L E EG V G+C
Sbjct: 125 KLCYICNEERPNDAKKGACMSCNKSTCKRSFHVTCAQRKGLLCE--EGAISRNVKYCGYC 182
Query: 298 KTH 300
+ H
Sbjct: 183 ENH 185
Score = 31.2 bits (69), Expect = 3.6
Identities = 23/67 (34%), Positives = 32/67 (47%), Gaps = 8/67 (11%)
Query: 47 NSSLFHTPPSSPTPPPSTYSLPTKKRITALQ---PHLHHHNNIPNDAVPLIDLNVEYSPS 103
NS +F PP++ +PP L T R + Q P L + N + + PLI S +
Sbjct: 271 NSGIFVPPPTAFSPP-----LTTSSRSSVAQDPSPPLTINKNSLSSSGPLIPSTAHLSAT 325
Query: 104 LPSATPI 110
SATPI
Sbjct: 326 TASATPI 332
>PHFF_HUMAN (Q9NQC1) PHD finger protein 15
Length = 576
Score = 108 bits (270), Expect = 2e-23
Identities = 61/167 (36%), Positives = 79/167 (46%), Gaps = 12/167 (7%)
Query: 144 IVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAMK 203
+VFCD CN+ VH +CYG + ++P G W C C P +C LCP + GA+K
Sbjct: 1 MVFCDKCNVCVHQACYG---ILKVPTGSWLCRTCAL------GVQP-KCLLCPKRGGALK 50
Query: 204 QTTDG-KWVHLVCALLVPEVFFVDPEGREGID-CSKIPKKRWLEKCYVCGCFDGCALVCS 261
T G KWVH+ CAL +PEV PE E I S IP RW C +C G + CS
Sbjct: 51 PTRSGTKWVHVSCALWIPEVSIGCPEKMEPITKISHIPASRWALSCSLCKECTGTCIQCS 110
Query: 262 EQKCGLGFHITCGIKEDLCIEYKEGKKGATVVAGFCKTHSQIWEKNK 308
C FH+TC L + FC+ HS +N+
Sbjct: 111 MPSCVTAFHVTCAFDHGLEMRTILADNDEVKFKSFCQEHSDGGPRNE 157
>PHFE_MOUSE (Q9D4H9) PHD finger protein 14
Length = 881
Score = 106 bits (265), Expect = 7e-23
Identities = 71/253 (28%), Positives = 112/253 (44%), Gaps = 28/253 (11%)
Query: 62 PSTYSLPTKKRITALQPHLHHHNNIPNDAVPLIDLNVEYSPSLPSATPIEKQSQKQDIEF 121
P++ + KKR L + + ND+ L + S S + +EK Q+
Sbjct: 255 PASEAGGKKKRSKVLSRNSADDEELTNDS-----LTLSQSKSNEDSLILEKS---QNWSS 306
Query: 122 EVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYG-----NPLVKQIPDGD---WF 173
+ D ++CCVC ++ D I+ CD C + VH CYG + ++ + WF
Sbjct: 307 QKMDHILICCVCLGDNSEDADEIIQCDNCGITVHEGCYGVDGESDSIMSSASENSTEPWF 366
Query: 174 CDQCRFKNDIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGI 233
CD C+ P C LCP ++G K+T G+WVH+VCAL VP V F D + +
Sbjct: 367 CDACK------CGVSP-SCELCPNQDGIFKETDAGRWVHIVCALYVPGVAFGDIDKLRPV 419
Query: 234 DCSKIPKKRW-LEKCYVCG----CFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKK 288
+++ ++ ++C C G + C C FH+TC KE L E +
Sbjct: 420 TLTEMNYSKYGAKECSFCEDPRFARTGVCISCDAGMCRAYFHVTCAQKEGLLSEAAAEED 479
Query: 289 GATVVAGFCKTHS 301
A +CK H+
Sbjct: 480 IADPFFAYCKQHA 492
>PHFE_HUMAN (O94880) PHD finger protein 14
Length = 888
Score = 105 bits (261), Expect = 2e-22
Identities = 69/245 (28%), Positives = 108/245 (43%), Gaps = 28/245 (11%)
Query: 70 KKRITALQPHLHHHNNIPNDAVPLIDLNVEYSPSLPSATPIEKQSQKQDIEFEVDDDEIL 129
KK+ L + + ND+ L + S S + +EK Q+ + D ++
Sbjct: 270 KKKSKVLSRNSADDEELTNDS-----LTLSQSKSNEDSLILEKS---QNWSSQKMDHILI 321
Query: 130 CCVCHSTDANAEDPIVFCDGCNLMVHASCYG-----NPLVKQIPDGD---WFCDQCRFKN 181
CCVC ++ D I+ CD C + VH CYG + ++ + WFCD C+
Sbjct: 322 CCVCLGDNSEDADEIIQCDNCGITVHEGCYGVDGESDSIMSSASENSTEPWFCDACK--- 378
Query: 182 DIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKK 241
P C LCP ++G K+T G+WVH+VCAL VP V F D + + +++
Sbjct: 379 ---CGVSP-SCELCPNQDGIFKETDAGRWVHIVCALYVPGVAFGDIDKLRPVTLTEMNYS 434
Query: 242 RW-LEKCYVCG----CFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATVVAGF 296
++ ++C C G + C C FH+TC KE L E + A +
Sbjct: 435 KYGAKECSFCEDPRFARTGVCISCDAGMCRAYFHVTCAQKEGLLSEAAAEEDIADPFFAY 494
Query: 297 CKTHS 301
CK H+
Sbjct: 495 CKQHA 499
>JJ2B_HUMAN (O94953) Jumonji domain containing protein 2B
Length = 1096
Score = 102 bits (254), Expect = 1e-21
Identities = 48/135 (35%), Positives = 71/135 (52%), Gaps = 12/135 (8%)
Query: 143 PIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAM 202
P++ C C L VHASCYG + ++ + W C +C C LC + GA+
Sbjct: 756 PLIACGKCCLQVHASCYG--IRPELVNEGWTCSRCA------AHAWTAECCLCNLRGGAL 807
Query: 203 KQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKCYVC----GCFDGCAL 258
+ TTD +W+H++CA+ VPE F++ R +D S IP++RW KC C G +
Sbjct: 808 QMTTDRRWIHVICAIAVPEARFLNVIERHPVDISAIPEQRWKLKCVYCRKRMKKVSGACI 867
Query: 259 VCSEQKCGLGFHITC 273
CS + C FH+TC
Sbjct: 868 QCSYEHCSTSFHVTC 882
>LI49_CAEEL (Q20318) Protein lin-49 (Abnormal cell lineage protein
49)
Length = 1042
Score = 102 bits (253), Expect = 2e-21
Identities = 63/198 (31%), Positives = 89/198 (44%), Gaps = 16/198 (8%)
Query: 109 PIEKQSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIP 168
P E K + E+DD +C +C D + + IV+CD CNL VH CYG P IP
Sbjct: 180 PKEFHKLKDENGEELDD---VCNICLDGDTSNCNQIVYCDRCNLSVHQDCYGIPF---IP 233
Query: 169 DGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPE 228
+G C +C + G + C LCP+ GA KQ +WVH++C + V E F +
Sbjct: 234 EGCLECRRCGI-----SPAGRVNCVLCPSTTGAFKQVDQKRWVHVLCVIWVDETHFGNTI 288
Query: 229 GREGI-DCSKIPKKRWLEKCYVC----GCFDGCALVCSEQKCGLGFHITCGIKEDLCIEY 283
E + + K R C +C G + CSE KC FH+TC L +
Sbjct: 289 FMENVQNVEKALHDRRALSCLLCKNRQNARMGACIQCSETKCTASFHVTCARDSGLVMRI 348
Query: 284 KEGKKGATVVAGFCKTHS 301
E + G +C H+
Sbjct: 349 NETEDGQVNRFVWCPKHA 366
>JJ2B_MOUSE (Q91VY5) Jumonji domain containing protein 2B
Length = 1086
Score = 101 bits (252), Expect = 2e-21
Identities = 48/135 (35%), Positives = 72/135 (52%), Gaps = 12/135 (8%)
Query: 143 PIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAM 202
P++ C C L VHASCYG + ++ W C +C C LC + GA+
Sbjct: 744 PLISCAHCCLQVHASCYG--VRPELAKEGWTCSRCA------AHAWTAECCLCNLRGGAL 795
Query: 203 KQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKCYVC----GCFDGCAL 258
++TT+ +W+H++CA+ VPEV F++ R +D S IP++RW KC C G +
Sbjct: 796 QRTTEHRWIHVICAIAVPEVRFLNVIERNPVDVSAIPEQRWKLKCIYCRKRMKRVSGACI 855
Query: 259 VCSEQKCGLGFHITC 273
CS + C FH+TC
Sbjct: 856 QCSYEHCSTSFHVTC 870
>JJ2A_HUMAN (O75164) Jumonji domain containing protein 2A
Length = 1064
Score = 98.6 bits (244), Expect = 2e-20
Identities = 55/152 (36%), Positives = 76/152 (49%), Gaps = 21/152 (13%)
Query: 135 STDANAEDP---------IVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDT 185
STD N P +V C C++ VHASCYG P K DW C +C N ++
Sbjct: 717 STDINLSTPYLEEDGTSILVSCKKCSVRVHASCYGVPPAKA--SEDWMCSRCS-ANALEE 773
Query: 186 DTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLE 245
D C LC + GA+++ D +WVH+ CA+ + E FV+ R +D SKIP R+
Sbjct: 774 D-----CCLCSLRGGALQRANDDRWVHVSCAVAILEARFVNIAERSPVDVSKIPLPRFKL 828
Query: 246 KCYVC----GCFDGCALVCSEQKCGLGFHITC 273
KC C GC + CS +C FH++C
Sbjct: 829 KCIFCKKRRKRTAGCCVQCSHGRCPTAFHVSC 860
>JJ2A_MOUSE (Q8BW72) Jumonji domain containing protein 2A (Fragment)
Length = 1036
Score = 96.7 bits (239), Expect = 7e-20
Identities = 54/152 (35%), Positives = 76/152 (49%), Gaps = 21/152 (13%)
Query: 135 STDANAEDP---------IVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDT 185
STD N P +V C C++ VHASCYG P K +W C +C N ++
Sbjct: 717 STDINLSTPYLEEDGTSMLVSCKKCSVRVHASCYGVPPAKA--SEEWMCSRCS-ANALEE 773
Query: 186 DTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLE 245
D C LC + GA+++ D +WVH+ CA+ + E FV+ R +D SKIP R+
Sbjct: 774 D-----CCLCSLRGGALQRANDDRWVHVSCAVAILEARFVNIAERSPVDVSKIPLPRFKL 828
Query: 246 KCYVC----GCFDGCALVCSEQKCGLGFHITC 273
KC C GC + CS +C FH++C
Sbjct: 829 KCVFCKKRRKRNAGCCVQCSHGRCPTAFHVSC 860
>JJ2C_MOUSE (Q8VCD7) Jumonji domain containing protein 2C
Length = 1054
Score = 94.0 bits (232), Expect = 5e-19
Identities = 49/134 (36%), Positives = 70/134 (51%), Gaps = 12/134 (8%)
Query: 144 IVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAMK 203
++ C C + VHASCYG P ++ DG W C +C+ + C LC + GA+K
Sbjct: 713 LISCAKCFVRVHASCYGVPS-HEVCDG-WLCARCK------RNAWTAECCLCNLRGGALK 764
Query: 204 QTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKCYVC----GCFDGCALV 259
QT + +W H++CA+ VPEV F + R ID +IP +R KC C G +
Sbjct: 765 QTKNNQWAHVICAVAVPEVRFTNVPERTQIDVDRIPLQRLKLKCIFCRHRVKKVSGACIQ 824
Query: 260 CSEQKCGLGFHITC 273
CS +C FH+TC
Sbjct: 825 CSYGRCPASFHVTC 838
>JJ2C_HUMAN (Q9H3R0) Jumonji domain containing protein 2C (Gene
amplified in squamous cell carcinoma 1 protein) (GASC-1
protein)
Length = 1056
Score = 94.0 bits (232), Expect = 5e-19
Identities = 50/134 (37%), Positives = 70/134 (51%), Gaps = 12/134 (8%)
Query: 144 IVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAMK 203
++ C C + VHASCYG P +I DG W C +C+ + C LC + GA+K
Sbjct: 715 LISCAKCCVRVHASCYGIPS-HEICDG-WLCARCK------RNAWTAECCLCNLRGGALK 766
Query: 204 QTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKCYVC----GCFDGCALV 259
QT + +W H++CA+ VPEV F + R ID +IP +R KC C G +
Sbjct: 767 QTKNNRWAHVMCAVAVPEVRFTNVPERTQIDVGRIPLQRLKLKCIFCRHRVKRVSGACIQ 826
Query: 260 CSEQKCGLGFHITC 273
CS +C FH+TC
Sbjct: 827 CSYGRCPASFHVTC 840
>YGN1_YEAST (P53127) Hypothetical 163.2 kDa protein in RPL1B-CEG1
intergenic region
Length = 1403
Score = 76.3 bits (186), Expect = 1e-13
Identities = 62/263 (23%), Positives = 107/263 (40%), Gaps = 45/263 (17%)
Query: 38 PTKKRKESRNSSLFHTPPSSPTPPPSTYSLPTKKRITALQPHLHHHNNIPNDAVPLIDLN 97
P+ K+ + N P + P Y++ + + T++ +H N P+ + D+
Sbjct: 959 PSTKKIKMINDIALSNPLNEPNGASYNYTVISHSKETSVALEKYHDGNKPSKMLEK-DMI 1017
Query: 98 VEYSPSLPSATPIEKQSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHAS 157
++++ + P + D + + C VC + ++ V C C L VH
Sbjct: 1018 LKHTKNKP---------KNPDTAWANNSARTFCSVCKEKFNDNDNYEVVCGNCGLTVHYF 1068
Query: 158 CYGNPLVKQIPDGD------WFCDQCRFKNDIDTDTGPI-----RCSLCPTKE------- 199
CY L K + W CD C D PI +CS+CPTK+
Sbjct: 1069 CYAIKLPKDMKKNTNLKTFKWLCDPC------SNDLNPIISTTYQCSMCPTKDYDYDRYR 1122
Query: 200 --------GAMKQTTDGKWVHLVCALLVPEVFFVDPEGRE-GIDCSKIPKKRWLEKCYVC 250
A+K T+ G WVHLVC+L ++ + + + + ++ + + K C VC
Sbjct: 1123 SQSFKICPDALKCTSLGTWVHLVCSLFNEDIKYGNGQSMQPALNTTAVLIKHSRFTCGVC 1182
Query: 251 GCFDGCALVCSEQKCGLGFHITC 273
G + C+ KC +HITC
Sbjct: 1183 RINGGGLVKCN--KCQYRYHITC 1203
>YANC_SCHPO (Q10077) Hypothetical protein C3H1.12c in chromosome I
Length = 1131
Score = 68.9 bits (167), Expect = 2e-11
Identities = 73/309 (23%), Positives = 96/309 (30%), Gaps = 90/309 (29%)
Query: 24 QQQEEQDLSHCSLLPTKKRKES------------------RNSSLFHTPPSSPTPPPSTY 65
QQ DL +C P KKRK RN+S P T +
Sbjct: 717 QQATPSDLRNCDFEPVKKRKADWDHLSNHDNEVKKENNRIRNASSLMENPRVSTKTFDNF 776
Query: 66 SLPTKKRITALQPHLHH--HNNIPNDAVPLIDLNVEYSPSLPSATPIEKQSQKQDIEFEV 123
+L I + NNI N K D+ F
Sbjct: 777 TLTHDSTINVKADTVKRARQNNIKN---------------------------KDDVNFS- 808
Query: 124 DDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCY-----------------------G 160
+D + C +C ++ C C VH CY
Sbjct: 809 EDRKKCCALCGIVGTEG---LLVCFKCGTCVHERCYVCDDYAENEQMLVSASHLSGRTTR 865
Query: 161 NPLVKQIPDG-----------DWFCDQCRFKNDIDTDTGPIRCSLC--PTKEGAMKQTTD 207
N I G W C CR ND C LC MK+T +
Sbjct: 866 NSASPGIVSGKKSYAKKDQVLSWACLSCR-SNDNLGQNNDNHCVLCLQSASHSLMKKTVE 924
Query: 208 GKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKCYVCGCFDGCALVCSEQKCGL 267
G WVHL+CA P+V+ E +++P RW +KC VCG + + S GL
Sbjct: 925 GNWVHLICASWTPDVYVPAEESEPVCGIAQLPPNRWEKKCEVCG--NSFGVCVSSPNSGL 982
Query: 268 GFHITCGIK 276
H+TC K
Sbjct: 983 TSHVTCAEK 991
Score = 30.0 bits (66), Expect = 8.1
Identities = 16/41 (39%), Positives = 18/41 (43%), Gaps = 1/41 (2%)
Query: 140 AEDPIVFCDGCNLMVHASCYGNPLVKQIPDG-DWFCDQCRF 179
A D V C C H C PL+K+ P G W C C F
Sbjct: 270 AFDFSVQCADCKKYYHMDCVVPPLLKKPPHGFGWTCATCSF 310
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.137 0.445
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 45,167,978
Number of Sequences: 164201
Number of extensions: 2202762
Number of successful extensions: 8731
Number of sequences better than 10.0: 177
Number of HSP's better than 10.0 without gapping: 51
Number of HSP's successfully gapped in prelim test: 126
Number of HSP's that attempted gapping in prelim test: 8320
Number of HSP's gapped (non-prelim): 390
length of query: 322
length of database: 59,974,054
effective HSP length: 110
effective length of query: 212
effective length of database: 41,911,944
effective search space: 8885332128
effective search space used: 8885332128
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC147714.16


