

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147714.1 - phase: 0 /pseudo
(805 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
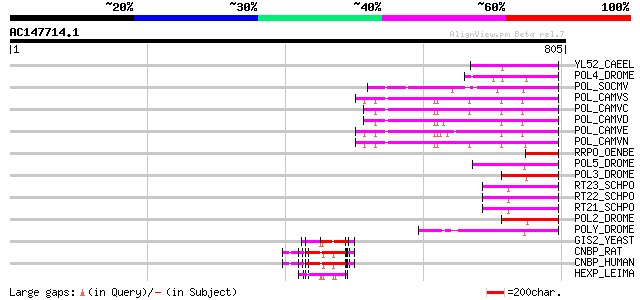
Score E
Sequences producing significant alignments: (bits) Value
YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III 97 1e-19
POL4_DROME (P10394) Retrovirus-related Pol polyprotein from tran... 91 1e-17
POL_SOCMV (P15629) Enzymatic polyprotein [Contains: Aspartic pro... 82 4e-15
POL_CAMVS (P03554) Enzymatic polyprotein [Contains: Aspartic pro... 79 4e-14
POL_CAMVC (P03555) Enzymatic polyprotein [Contains: Aspartic pro... 79 5e-14
POL_CAMVD (P03556) Enzymatic polyprotein [Contains: Aspartic pro... 77 2e-13
POL_CAMVE (Q02964) Enzymatic polyprotein [Contains: Aspartic pro... 76 3e-13
POL_CAMVN (Q00962) Enzymatic polyprotein [Contains: Aspartic pro... 75 5e-13
RRPO_OENBE (P31843) RNA-directed DNA polymerase homolog (Reverse... 75 7e-13
POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from tran... 72 6e-12
POL3_DROME (P04323) Retrovirus-related Pol polyprotein from tran... 71 1e-11
RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protei... 71 1e-11
RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protei... 71 1e-11
RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protei... 71 1e-11
POL2_DROME (P20825) Retrovirus-related Pol polyprotein from tran... 70 3e-11
POLY_DROME (P10401) Retrovirus-related Pol polyprotein from tran... 67 1e-10
GIS2_YEAST (P53849) Zinc-finger protein GIS2 65 5e-10
CNBP_RAT (P62634) Cellular nucleic acid binding protein (CNBP) (... 62 8e-09
CNBP_HUMAN (P62633) Cellular nucleic acid binding protein (CNBP)... 62 8e-09
HEXP_LEIMA (Q04832) DNA-binding protein HEXBP (Hexamer-binding p... 61 1e-08
>YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III
Length = 2186
Score = 97.4 bits (241), Expect = 1e-19
Identities = 48/131 (36%), Positives = 79/131 (59%), Gaps = 3/131 (2%)
Query: 669 VVCDFPEVFPDEIPDVPPEREVEFSIDLVPGTKPVSMAPYRISAS---ELKKQLEDLLEK 725
V+ F +VF ++ E I+L G +P+ P I + E++K ++ +L +
Sbjct: 909 VIEQFQDVFAISDDELGRNSGTECVIELKEGAEPIRQKPRPIPLALKPEIRKMIQKMLNQ 968
Query: 726 KFVRPSVSPWGASVLLVKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGARVF 785
K +R S SPW + V+LVKKKDGS+R+CIDYR++NKV N +PLP I+ + L G +++
Sbjct: 969 KVIRESKSPWSSPVVLVKKKDGSIRMCIDYRKVNKVVKNNAHPLPNIEATLQSLAGKKLY 1028
Query: 786 SKIDLRSGYHQ 796
+ D+ +G+ Q
Sbjct: 1029 TVFDMIAGFWQ 1039
Score = 39.3 bits (90), Expect = 0.042
Identities = 29/114 (25%), Positives = 45/114 (39%), Gaps = 12/114 (10%)
Query: 393 NERKGKGQQSRPKPYSAPADKGKQKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACPEEIK 452
N R S+ ++ +Q + P+K C +C ++G C ++ K
Sbjct: 504 NSRNSWNNNSQNNSAASQNISREQSWKTISVPQKHQNPSDRCSDCQQRGWHMFWCSKKSK 563
Query: 453 -----KCVRCGKKGHVVADCNRT-DIVCFNCNGEGHISSQC------TQPKRAP 494
KC C + G +A C + + CF CN GHI+ C T K AP
Sbjct: 564 DNASQKCDECQQSGWHMASCFKLKNRACFRCNEMGHIAWNCPKKNENTSEKEAP 617
>POL4_DROME (P10394) Retrovirus-related Pol polyprotein from
transposon 412 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1237
Score = 90.9 bits (224), Expect = 1e-17
Identities = 58/160 (36%), Positives = 90/160 (56%), Gaps = 25/160 (15%)
Query: 660 NQAVIDRLPVVCDFPEVFPDEIPDVPPEREVEFSIDLVPGT--------------KPVSM 705
N+ V+ +L +FPE+F ++ ++ E F+++ P T +PV
Sbjct: 260 NKTVLSQLKK--NFPELFKSQLENICSEYIDIFALESEPITVNNLYKQQLRLKDDEPVYT 317
Query: 706 APYRISAS---ELKKQLEDLLEKKFVRPSVSPWGASVLLVKKKDG------SMRLCIDYR 756
YR S E++ Q++ L++ K V PSVS + + +LLV KK RL IDYR
Sbjct: 318 KNYRSPHSQVEEIQAQVQKLIKDKIVEPSVSQYNSPLLLVPKKSSPNSDKKKWRLVIDYR 377
Query: 757 QLNKVTIKNRYPLPRIDDLMDQLVGARVFSKIDLRSGYHQ 796
Q+NK + +++PLPRIDD++DQL A+ FS +DL SG+HQ
Sbjct: 378 QINKKLLADKFPLPRIDDILDQLGRAKYFSCLDLMSGFHQ 417
>POL_SOCMV (P15629) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 692
Score = 82.4 bits (202), Expect = 4e-15
Identities = 74/293 (25%), Positives = 140/293 (47%), Gaps = 30/293 (10%)
Query: 520 INNTPLVAIIDTGATHCFIAFDCVSALGLDLSDMNGEMVVETPAKGSVTTSLVCLKCPLS 579
I +A IDTGAT CF + ++ E+++ +K + ++ + L
Sbjct: 26 IGKRNFLAYIDTGATLCFGKRKISN--NWEILKQPKEIIIADKSKHYIREAISNVF--LK 81
Query: 580 MFGRDFEMDLVCLPLSGMDVILGMNWLE-YNHVLINCFSKSVHFSSVEEESGAEFLSTKQ 638
+ ++F + ++ L SG+D+I+G N+L+ Y + + + + ++ ++ +STK
Sbjct: 82 IENKEFLIPIIYLHDSGLDLIIGNNFLKLYQPFIQRLETIELRWKNLNNPKESQMISTKI 141
Query: 639 LKQ-------LERDGILMFSLMATLSIENQAVIDRLPVVCDFPEVFPDEIPDVPPEREVE 691
L + E+ I + + +IE Q L VC + + + +
Sbjct: 142 LTKNEVLKLSFEKIHICLEKYLFFKTIEEQ-----LEEVCS-----EHPLDETKNKNGLL 191
Query: 692 FSIDLVPGTKPVSMA---PYRI-SASELKKQLEDLLEKKFVRPSVSPWGASVLLVKK--- 744
I L + +++ PY I E K++ EDLL+K +R S SP A V+
Sbjct: 192 IEIRLKDPLQEINVTNRIPYTIRDVQEFKEECEDLLKKGLIRESQSPHSAPAFYVENHNE 251
Query: 745 -KDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGARVFSKIDLRSGYHQ 796
K G R+ I+Y+++N+ TI + Y LPR D +++++ G+ FS +D +SGY+Q
Sbjct: 252 IKRGKRRMVINYKKMNEATIGDSYKLPRKDFILEKIKGSLWFSSLDAKSGYYQ 304
>POL_CAMVS (P03554) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 679
Score = 79.3 bits (194), Expect = 4e-14
Identities = 87/347 (25%), Positives = 155/347 (44%), Gaps = 56/347 (16%)
Query: 502 LTGTQTENEDRL---------IRGTCYINNTPLVAI---IDTGATHCFIAFDCVSALGLD 549
L TQT+ E + I+G Y + + +DTGA+ C + + +
Sbjct: 5 LLKTQTQTEQVMNVTNPNSIYIKGRLYFKGYKKIELHCFVDTGASLCIASKFVIP----E 60
Query: 550 LSDMNGE--MVVETPAKGSVTTSLVCLKCPLSMFGRDFEMDLVCLPLSGMDVILGMNWLE 607
+N E ++V+ S+T S VC L + G F + V SG+D I+G N+ +
Sbjct: 61 EHWVNAERPIMVKIADGSSITISKVCKDIDLIIAGEIFRIPTVYQQESGIDFIIGNNFCQ 120
Query: 608 YNHVLIN-----CFSKS----VHFSSVEE--ESGAE-FLST--KQLKQLERDGILMFSLM 653
I F+K+ VH + + G E FL + K+ K + + + + +
Sbjct: 121 LYEPFIQFTDRVIFTKNKSYPVHIAKLTRAVRVGTEGFLESMKKRSKTQQPEPVNISTNK 180
Query: 654 ATLSIENQAVID---------------RLPVVCDFPEVFPDEIPDVPPERE--VEFSIDL 696
+E A++ R+ + + E E P P + + ++ SI L
Sbjct: 181 IENPLEEIAILSEGRRLSEEKLFITQQRMQKIEELLEKVCSENPLDPNKTKQWMKASIKL 240
Query: 697 VPGTKPVSMAPYRISA---SELKKQLEDLLEKKFVRPSVSPWGASVLLV----KKKDGSM 749
+K + + P + S E KQ+++LL+ K ++PS SP A LV +K+ G
Sbjct: 241 SDPSKAIKVKPMKYSPMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKK 300
Query: 750 RLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGARVFSKIDLRSGYHQ 796
R+ ++Y+ +NK T+ + Y LP D+L+ + G ++FS D +SG+ Q
Sbjct: 301 RMVVNYKAMNKATVGDAYNLPNKDELLTLIRGKKIFSSFDCKSGFWQ 347
>POL_CAMVC (P03555) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 679
Score = 79.0 bits (193), Expect = 5e-14
Identities = 83/326 (25%), Positives = 149/326 (45%), Gaps = 47/326 (14%)
Query: 514 IRGTCYINNTPLVAI---IDTGATHCFIAFDCVSALGLDLSDMNGE--MVVETPAKGSVT 568
I+G Y + + +DTGA+ C + + + +N E ++V+ S+T
Sbjct: 26 IKGRLYFKGYKKIELHCFVDTGASLCIASKFVIP----EEHWVNAERPIMVKIADGSSIT 81
Query: 569 TSLVCLKCPLSMFGRDFEMDLVCLPLSGMDVILGMNWLEYNHVLIN-----CFSKS---- 619
S VC L + G F++ V SG+D I+G N+ + I F+K+
Sbjct: 82 ISKVCKDIDLIIAGEIFKIPTVYQQESGIDFIIGNNFCQLYEPFIQFTDRVIFTKNKSYP 141
Query: 620 VHFSSVEE--ESGAE-FLST--KQLKQLERDGILMFSLMATLSIENQAVID--------- 665
VH + + G E FL + K+ K + + + + + +E A++
Sbjct: 142 VHITKLTRAVRVGIEGFLESMKKRSKTQQPEPVNISTNKIENPLEEIAILSEGRRLSEEK 201
Query: 666 ------RLPVVCDFPEVFPDEIPDVPPERE--VEFSIDLVPGTKPVSMAPYRISA---SE 714
R+ + + E E P P + + ++ SI L +K + + P + S E
Sbjct: 202 LFITQQRMQKIEELLEKVCSENPLDPNKTKQWMKASIKLSDPSKAIKVKPMKYSPMDREE 261
Query: 715 LKKQLEDLLEKKFVRPSVSPWGASVLLV----KKKDGSMRLCIDYRQLNKVTIKNRYPLP 770
KQ+++LL+ K ++PS SP A LV +K+ G R+ ++Y+ +NK TI + Y LP
Sbjct: 262 FDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATIGDAYNLP 321
Query: 771 RIDDLMDQLVGARVFSKIDLRSGYHQ 796
D+L+ + G ++FS D +SG+ Q
Sbjct: 322 NKDELLTLIRGKKIFSSFDCKSGFWQ 347
>POL_CAMVD (P03556) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 674
Score = 77.0 bits (188), Expect = 2e-13
Identities = 81/319 (25%), Positives = 144/319 (44%), Gaps = 40/319 (12%)
Query: 514 IRGTCYINNTPLVAI---IDTGATHCFIAFDCVSALGLDLSDMNGE--MVVETPAKGSVT 568
I+G Y + + +DTGA+ C + + + +N E ++V+ S+T
Sbjct: 28 IKGRLYFKGYKKIELHCFVDTGASLCIASKFVIP----EEHWINAERPIMVKIADGSSIT 83
Query: 569 TSLVCLKCPLSMFGRDFEMDLVCLPLSGMDVILGMNWLEYNHVLIN-----CFSKS---- 619
+ VC L + G F + V SG+D I+G N+ + I F+K
Sbjct: 84 INKVCRDIDLIIAGEIFHIPTVYQQESGIDFIIGNNFCQLYEPFIQFTDRVIFTKDRTYP 143
Query: 620 VHFSSVEE----------ESGAEFLSTKQLK--QLERDGILMFSLMATLSIENQAVID-R 666
VH + + ES + T+Q + + + I + S LS E + R
Sbjct: 144 VHIAKLTRAVRVGTEGFLESMKKRSKTQQPEPVNISTNKIAILSEGRRLSEEKLFITQQR 203
Query: 667 LPVVCDFPEVFPDEIPDVPPERE--VEFSIDLVPGTKPVSMAPYRISA---SELKKQLED 721
+ + + E E P P + + ++ SI L +K + + P + S E KQ+++
Sbjct: 204 MQKIEELLEKVCSENPLDPNKTKQWMKASIKLSDPSKAIKVKPMKYSPMDREEFDKQIKE 263
Query: 722 LLEKKFVRPSVSPWGASVLLV----KKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMD 777
LL+ K ++PS SP A LV +K+ G R+ ++Y+ +NK T+ + Y P D+L+
Sbjct: 264 LLDLKVIKPSKSPHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATVGDAYNPPNKDELLT 323
Query: 778 QLVGARVFSKIDLRSGYHQ 796
+ G ++FS D +SG+ Q
Sbjct: 324 LIRGKKIFSSFDCKSGFWQ 342
>POL_CAMVE (Q02964) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 679
Score = 76.3 bits (186), Expect = 3e-13
Identities = 88/348 (25%), Positives = 157/348 (44%), Gaps = 58/348 (16%)
Query: 502 LTGTQTENEDRL---------IRGTCYINNTPLVAI---IDTGATHCFIAFDCVSALGLD 549
L TQT+ E + I+G Y + + +DTGA+ C + + +
Sbjct: 5 LLKTQTQTEQVMNVTNPNSIYIKGRLYFKGYKKIELHCFVDTGASLCIASKFVIP----E 60
Query: 550 LSDMNGE--MVVETPAKGSVTTSLVCLKCPLSMFGRDFEMDLVCLPLSGMDVILGMNWLE 607
+N E ++V+ S+T S VC L + F++ V SG+D I+G N+ +
Sbjct: 61 EHWVNAERPIMVKIADGSSITISKVCKDIDLIIAREIFKIPTVYQQESGIDFIIGNNFCQ 120
Query: 608 YNHVLIN-----CFSKS----VHFS---------------SVEEESGAEF-----LSTKQ 638
I F+K+ VH + S+++ S + +ST +
Sbjct: 121 LYEPFIQFTDRVIFTKNKSYPVHIAKLTRAVRVGTEGFLESMKKRSKTQQPEPVNISTNK 180
Query: 639 LKQLERDGILMFSLMATLSIENQAVID-RLPVVCDFPEVFPDEIPDVPPERE--VEFSID 695
++ ++ I + S LS E + R+ + + E E P P + + ++ SI
Sbjct: 181 IENPLKE-IAILSEGRRLSEEKLFITQQRMQKIEELLEKVCSENPLDPNKTKQWMKASIK 239
Query: 696 LVPGTKPVSMAPYRISA---SELKKQLEDLLEKKFVRPSVSPWGASVLLV----KKKDGS 748
L +K + + P + S E KQ+++LL+ K ++PS SP A LV +K+ G
Sbjct: 240 LSDPSKAIKVKPMKYSPMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAEKRRGK 299
Query: 749 MRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGARVFSKIDLRSGYHQ 796
R+ ++Y+ +NK TI + Y LP D+L+ + G ++FS D +SG+ Q
Sbjct: 300 KRMVVNYKAMNKATIGDAYNLPNKDELLTLIRGKKIFSSFDCKSGFWQ 347
>POL_CAMVN (Q00962) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 680
Score = 75.5 bits (184), Expect = 5e-13
Identities = 86/347 (24%), Positives = 154/347 (43%), Gaps = 56/347 (16%)
Query: 502 LTGTQTENEDRL---------IRGTCYINNTPLVAI---IDTGATHCFIAFDCVSALGLD 549
L TQT+ E + I+G Y + + +DTGA+ C + + +
Sbjct: 6 LLKTQTQTEQVMNVTNPNSIYIKGRLYFKGYKKIELHCFVDTGASLCIASKFVIP----E 61
Query: 550 LSDMNGE--MVVETPAKGSVTTSLVCLKCPLSMFGRDFEMDLVCLPLSGMDVILGMNWLE 607
+N E ++V+ S+T S VC L + G F++ V SG+D I+G N+ +
Sbjct: 62 EHWVNAERPIMVKIADGSSITISKVCKDIDLIIVGVIFKIPTVYQQESGIDFIIGNNFCQ 121
Query: 608 YNHVLIN-----CFSKS----VHFSSVEE--ESGAE-FLST--KQLKQLERDGILMFSLM 653
I F+K+ VH + + G E FL + K+ K + + + + +
Sbjct: 122 LYEPFIQFTDRVIFTKNKSYPVHIAKLTRAVRVGTEGFLESMKKRSKTQQPEPVNISTNK 181
Query: 654 ATLSIENQAVID---------------RLPVVCDFPEVFPDEIPDVPPERE--VEFSIDL 696
+E A++ R+ + E E P P + + ++ SI L
Sbjct: 182 IENPLEEIAILSEGRRLSEEKLFITQQRMQKTEELLEKVCSENPLDPNKTKQWMKASIKL 241
Query: 697 VPGTKPVSMAPYRISA---SELKKQLEDLLEKKFVRPSVSPWGASVLLVKKKD----GSM 749
+K + + P + S E KQ+++LL+ K ++PS SP A LV + G+
Sbjct: 242 SDPSKAIKVKPMKYSPMDREEFDKQIKELLDLKVIKPSKSPHMAPAFLVNNEAENGRGNK 301
Query: 750 RLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGARVFSKIDLRSGYHQ 796
R+ ++Y+ +NK T+ + Y LP D+L+ + G ++FS D +SG+ Q
Sbjct: 302 RMVVNYKAMNKATVGDAYNLPNKDELLTLIRGKKIFSSFDCKSGFWQ 348
>RRPO_OENBE (P31843) RNA-directed DNA polymerase homolog (Reverse
transcriptase homolog)
Length = 142
Score = 75.1 bits (183), Expect = 7e-13
Identities = 33/49 (67%), Positives = 41/49 (83%)
Query: 748 SMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGARVFSKIDLRSGYHQ 796
S+R+CIDYR L KVTIKN+YP+PR+DDL D+L A F+K+DLRSGY Q
Sbjct: 5 SLRMCIDYRALTKVTIKNKYPIPRVDDLFDRLAQATWFTKLDLRSGYWQ 53
>POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from
transposon opus [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1003
Score = 72.0 bits (175), Expect = 6e-12
Identities = 40/132 (30%), Positives = 75/132 (56%), Gaps = 7/132 (5%)
Query: 672 DFPEVFPDEIPDVPPEREVEFSIDLVPGTKPVSMA-PYRISA-SELKKQLEDLLEKKFVR 729
+FP +F + + E V+ I + + PY ++ E+++Q+++LL+ +R
Sbjct: 94 EFPRIFEPPLSGMSVETAVKAEIRTNTQDPIYAKSYPYPVNMRGEVERQIDELLQDGIIR 153
Query: 730 PSVSPWGASVLLVKKK-----DGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGARV 784
PS SP+ + + +V KK + R+ +D+++LN VTI + YP+P I+ + L A+
Sbjct: 154 PSNSPYNSPIWIVPKKPKPNGEKQYRMVVDFKRLNTVTIPDTYPIPDINATLASLGNAKY 213
Query: 785 FSKIDLRSGYHQ 796
F+ +DL SG+HQ
Sbjct: 214 FTTLDLTSGFHQ 225
>POL3_DROME (P04323) Retrovirus-related Pol polyprotein from
transposon 17.6 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1058
Score = 71.2 bits (173), Expect = 1e-11
Identities = 33/88 (37%), Positives = 60/88 (67%), Gaps = 5/88 (5%)
Query: 714 ELKKQLEDLLEKKFVRPSVSPWGASVLLV-KKKDGS----MRLCIDYRQLNKVTIKNRYP 768
E++ Q++D+L + +R S SP+ + + +V KK+D S R+ IDYR+LN++T+ +R+P
Sbjct: 222 EVESQIQDMLNQGIIRTSNSPYNSPIWVVPKKQDASGKQKFRIVIDYRKLNEITVGDRHP 281
Query: 769 LPRIDDLMDQLVGARVFSKIDLRSGYHQ 796
+P +D+++ +L F+ IDL G+HQ
Sbjct: 282 IPNMDEILGKLGRCNYFTTIDLAKGFHQ 309
>RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protein
type 3
Length = 1333
Score = 70.9 bits (172), Expect = 1e-11
Identities = 34/113 (30%), Positives = 66/113 (58%), Gaps = 3/113 (2%)
Query: 686 PEREVEFSIDLVPGTKPVSMAPYRISASELKKQLEDL---LEKKFVRPSVSPWGASVLLV 742
P + +EF ++L + + Y + +++ +++ L+ +R S + V+ V
Sbjct: 396 PIKGLEFEVELTQENYRLPIRNYPLPPGKMQAMNDEINQGLKSGIIRESKAINACPVMFV 455
Query: 743 KKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGARVFSKIDLRSGYH 795
KK+G++R+ +DY+ LNK N YPLP I+ L+ ++ G+ +F+K+DL+S YH
Sbjct: 456 PKKEGTLRMVVDYKPLNKYVKPNIYPLPLIEQLLAKIQGSTIFTKLDLKSAYH 508
>RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protein
type 2
Length = 1333
Score = 70.9 bits (172), Expect = 1e-11
Identities = 34/113 (30%), Positives = 66/113 (58%), Gaps = 3/113 (2%)
Query: 686 PEREVEFSIDLVPGTKPVSMAPYRISASELKKQLEDL---LEKKFVRPSVSPWGASVLLV 742
P + +EF ++L + + Y + +++ +++ L+ +R S + V+ V
Sbjct: 396 PIKGLEFEVELTQENYRLPIRNYPLPPGKMQAMNDEINQGLKSGIIRESKAINACPVMFV 455
Query: 743 KKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGARVFSKIDLRSGYH 795
KK+G++R+ +DY+ LNK N YPLP I+ L+ ++ G+ +F+K+DL+S YH
Sbjct: 456 PKKEGTLRMVVDYKPLNKYVKPNIYPLPLIEQLLAKIQGSTIFTKLDLKSAYH 508
>RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protein
type 1
Length = 1333
Score = 70.9 bits (172), Expect = 1e-11
Identities = 34/113 (30%), Positives = 66/113 (58%), Gaps = 3/113 (2%)
Query: 686 PEREVEFSIDLVPGTKPVSMAPYRISASELKKQLEDL---LEKKFVRPSVSPWGASVLLV 742
P + +EF ++L + + Y + +++ +++ L+ +R S + V+ V
Sbjct: 396 PIKGLEFEVELTQENYRLPIRNYPLPPGKMQAMNDEINQGLKSGIIRESKAINACPVMFV 455
Query: 743 KKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGARVFSKIDLRSGYH 795
KK+G++R+ +DY+ LNK N YPLP I+ L+ ++ G+ +F+K+DL+S YH
Sbjct: 456 PKKEGTLRMVVDYKPLNKYVKPNIYPLPLIEQLLAKIQGSTIFTKLDLKSAYH 508
>POL2_DROME (P20825) Retrovirus-related Pol polyprotein from
transposon 297 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1059
Score = 69.7 bits (169), Expect = 3e-11
Identities = 32/88 (36%), Positives = 58/88 (65%), Gaps = 5/88 (5%)
Query: 714 ELKKQLEDLLEKKFVRPSVSPWGASVLLVKKKDGSM-----RLCIDYRQLNKVTIKNRYP 768
E++ Q++++L + +R S SP+ + +V KK + R+ IDYR+LN++TI +RYP
Sbjct: 221 EVENQVQEMLNQGLIRESNSPYNSPTWVVPKKPDASGANKYRVVIDYRKLNEITIPDRYP 280
Query: 769 LPRIDDLMDQLVGARVFSKIDLRSGYHQ 796
+P +D+++ +L + F+ IDL G+HQ
Sbjct: 281 IPNMDEILGKLGKCQYFTTIDLAKGFHQ 308
>POLY_DROME (P10401) Retrovirus-related Pol polyprotein from
transposon gypsy [Contains: Reverse transcriptase (EC
2.7.7.49); Endonuclease]
Length = 1035
Score = 67.4 bits (163), Expect = 1e-10
Identities = 54/211 (25%), Positives = 99/211 (46%), Gaps = 19/211 (9%)
Query: 594 LSGMDVILGMNWLEYNHVLINCFSKSVHFSSVEEESGAEFLSTKQLKQLERDGILMFSLM 653
L+ D I+G++ L V +N S+ + + E+ + S + F+ +
Sbjct: 85 LNAFDAIIGLDLLTQAGVKLNLAEDSLEYQGIAEK--LHYFSCPSVN---------FTDV 133
Query: 654 ATLSIENQAVIDRLPVVCDFPEVFPDEIPDVPPEREVEFSIDLVPGTKPVSMA-PYRISA 712
+ + + + + + F +P V +I V S A P +
Sbjct: 134 NDIVVPDSVKKEFKDTIIRRKKAFSTTNEALPFNTAVTATIRTVDNEPVYSRAYPTLMGV 193
Query: 713 SE-LKKQLEDLLEKKFVRPSVSPWGASVLLVKKK------DGSMRLCIDYRQLNKVTIKN 765
S+ + +++ LL+ +RPS SP+ + +V KK + + RL ID+R+LN+ TI +
Sbjct: 194 SDFVNNEVKQLLKDGIIRPSRSPYNSPTWVVDKKGTDAFGNPNKRLVIDFRKLNEKTIPD 253
Query: 766 RYPLPRIDDLMDQLVGARVFSKIDLRSGYHQ 796
RYP+P I ++ L A+ F+ +DL+SGYHQ
Sbjct: 254 RYPMPSIPMILANLGKAKFFTTLDLKSGYHQ 284
>GIS2_YEAST (P53849) Zinc-finger protein GIS2
Length = 153
Score = 65.5 bits (158), Expect = 5e-10
Identities = 30/74 (40%), Positives = 42/74 (56%), Gaps = 6/74 (8%)
Query: 430 AEIVCFNCGEKGHKSNACPE----EIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISS 485
+E +C+NC + GH C E K+C CG+ GHV ++C T CFNCN GHIS
Sbjct: 21 SERLCYNCNKPGHVQTDCTMPRTVEFKQCYNCGETGHVRSEC--TVQRCFNCNQTGHISR 78
Query: 486 QCTQPKRAPTTGRV 499
+C +PK+ +V
Sbjct: 79 ECPEPKKTSRFSKV 92
Score = 57.4 bits (137), Expect = 1e-07
Identities = 27/69 (39%), Positives = 38/69 (54%), Gaps = 6/69 (8%)
Query: 424 PKKKDA-AEIVCFNCGEKGHKSNACPEEIK----KCVRCGKKGHVVADCNRTDIVCFNCN 478
PKK +++ C+ CG H + C +E KC CG+ GH+ DC + D +C+NCN
Sbjct: 83 PKKTSRFSKVSCYKCGGPNHMAKDCMKEDGISGLKCYTCGQAGHMSRDC-QNDRLCYNCN 141
Query: 479 GEGHISSQC 487
GHIS C
Sbjct: 142 ETGHISKDC 150
Score = 49.3 bits (116), Expect = 4e-05
Identities = 18/40 (45%), Positives = 27/40 (67%), Gaps = 1/40 (2%)
Query: 452 KKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPK 491
K C CGK GH+ DC+ ++ +C+NCN GH+ + CT P+
Sbjct: 4 KACYVCGKIGHLAEDCD-SERLCYNCNKPGHVQTDCTMPR 42
>CNBP_RAT (P62634) Cellular nucleic acid binding protein (CNBP)
(Zinc finger protein 9)
Length = 177
Score = 61.6 bits (148), Expect = 8e-09
Identities = 32/82 (39%), Positives = 41/82 (49%), Gaps = 7/82 (8%)
Query: 425 KKKDAAEIVCFNCGEKGHKSNACPEEIKK----CVRCGKKGHVVADCNRTD-IVCFNCNG 479
K D E C+NCG GH + C E ++ C CGK GH+ DC+ D C++C
Sbjct: 65 KDCDLQEDACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGE 124
Query: 480 EGHISSQCTQPK--RAPTTGRV 499
GHI CT+ K R TG V
Sbjct: 125 FGHIQKDCTKVKCYRCGETGHV 146
Score = 60.1 bits (144), Expect = 2e-08
Identities = 25/61 (40%), Positives = 38/61 (61%), Gaps = 3/61 (4%)
Query: 429 AAEIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRT-DIVCFNCNGEGHISSQC 487
A E C++CGE GH C + KC RCG+ GHV +C++T ++ C+ C GH++ +C
Sbjct: 114 ADEQKCYSCGEFGHIQKDCTKV--KCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 171
Query: 488 T 488
T
Sbjct: 172 T 172
Score = 58.5 bits (140), Expect = 7e-08
Identities = 25/71 (35%), Positives = 41/71 (57%), Gaps = 5/71 (7%)
Query: 420 DVRRPKKKDAAEIVCFNCGEKGHKSNACPE-EIKKCVRCGKKGHVVADCNRTDIVCFNCN 478
D + PK++ E C+NCG+ GH + C + +KC CG+ GH+ DC T + C+ C
Sbjct: 86 DCKEPKRE--REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDC--TKVKCYRCG 141
Query: 479 GEGHISSQCTQ 489
GH++ C++
Sbjct: 142 ETGHVAINCSK 152
Score = 56.6 bits (135), Expect = 3e-07
Identities = 26/87 (29%), Positives = 35/87 (39%), Gaps = 28/87 (32%)
Query: 434 CFNCGEKGHKSNACPEEIKK----------------------------CVRCGKKGHVVA 465
CF CG GH + CP + C RCG+ GH+
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGRGGFTSDRGFQFVSSSLPDICYRCGESGHLAK 65
Query: 466 DCNRTDIVCFNCNGEGHISSQCTQPKR 492
DC+ + C+NC GHI+ C +PKR
Sbjct: 66 DCDLQEDACYNCGRGGHIAKDCKEPKR 92
Score = 56.2 bits (134), Expect = 3e-07
Identities = 27/96 (28%), Positives = 45/96 (46%), Gaps = 13/96 (13%)
Query: 396 KGKGQQSRPKPYSAPADKGKQKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACPEEIKKCV 455
+G+G +SR + +D+G Q + + +C+ CGE GH + C + C
Sbjct: 25 RGRGMRSRGRG-GFTSDRGFQFV--------SSSLPDICYRCGESGHLAKDCDLQEDACY 75
Query: 456 RCGKKGHVVADC----NRTDIVCFNCNGEGHISSQC 487
CG+ GH+ DC + C+NC GH++ C
Sbjct: 76 NCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 111
Score = 43.9 bits (102), Expect = 0.002
Identities = 16/43 (37%), Positives = 26/43 (60%), Gaps = 1/43 (2%)
Query: 426 KKDAAEIVCFNCGEKGHKSNACPEEIK-KCVRCGKKGHVVADC 467
+KD ++ C+ CGE GH + C + + C RCG+ GH+ +C
Sbjct: 129 QKDCTKVKCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 171
>CNBP_HUMAN (P62633) Cellular nucleic acid binding protein (CNBP)
(Zinc finger protein 9)
Length = 177
Score = 61.6 bits (148), Expect = 8e-09
Identities = 32/82 (39%), Positives = 41/82 (49%), Gaps = 7/82 (8%)
Query: 425 KKKDAAEIVCFNCGEKGHKSNACPEEIKK----CVRCGKKGHVVADCNRTD-IVCFNCNG 479
K D E C+NCG GH + C E ++ C CGK GH+ DC+ D C++C
Sbjct: 65 KDCDLQEDACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGE 124
Query: 480 EGHISSQCTQPK--RAPTTGRV 499
GHI CT+ K R TG V
Sbjct: 125 FGHIQKDCTKVKCYRCGETGHV 146
Score = 60.1 bits (144), Expect = 2e-08
Identities = 25/61 (40%), Positives = 38/61 (61%), Gaps = 3/61 (4%)
Query: 429 AAEIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRT-DIVCFNCNGEGHISSQC 487
A E C++CGE GH C + KC RCG+ GHV +C++T ++ C+ C GH++ +C
Sbjct: 114 ADEQKCYSCGEFGHIQKDCTKV--KCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 171
Query: 488 T 488
T
Sbjct: 172 T 172
Score = 58.5 bits (140), Expect = 7e-08
Identities = 25/71 (35%), Positives = 41/71 (57%), Gaps = 5/71 (7%)
Query: 420 DVRRPKKKDAAEIVCFNCGEKGHKSNACPE-EIKKCVRCGKKGHVVADCNRTDIVCFNCN 478
D + PK++ E C+NCG+ GH + C + +KC CG+ GH+ DC T + C+ C
Sbjct: 86 DCKEPKRE--REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDC--TKVKCYRCG 141
Query: 479 GEGHISSQCTQ 489
GH++ C++
Sbjct: 142 ETGHVAINCSK 152
Score = 56.6 bits (135), Expect = 3e-07
Identities = 26/87 (29%), Positives = 35/87 (39%), Gaps = 28/87 (32%)
Query: 434 CFNCGEKGHKSNACPEEIKK----------------------------CVRCGKKGHVVA 465
CF CG GH + CP + C RCG+ GH+
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGRGGFTSDRGFQFVSSSLPDICYRCGESGHLAK 65
Query: 466 DCNRTDIVCFNCNGEGHISSQCTQPKR 492
DC+ + C+NC GHI+ C +PKR
Sbjct: 66 DCDLQEDACYNCGRGGHIAKDCKEPKR 92
Score = 56.2 bits (134), Expect = 3e-07
Identities = 27/96 (28%), Positives = 45/96 (46%), Gaps = 13/96 (13%)
Query: 396 KGKGQQSRPKPYSAPADKGKQKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACPEEIKKCV 455
+G+G +SR + +D+G Q + + +C+ CGE GH + C + C
Sbjct: 25 RGRGMRSRGRG-GFTSDRGFQFV--------SSSLPDICYRCGESGHLAKDCDLQEDACY 75
Query: 456 RCGKKGHVVADC----NRTDIVCFNCNGEGHISSQC 487
CG+ GH+ DC + C+NC GH++ C
Sbjct: 76 NCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 111
Score = 43.9 bits (102), Expect = 0.002
Identities = 16/43 (37%), Positives = 26/43 (60%), Gaps = 1/43 (2%)
Query: 426 KKDAAEIVCFNCGEKGHKSNACPEEIK-KCVRCGKKGHVVADC 467
+KD ++ C+ CGE GH + C + + C RCG+ GH+ +C
Sbjct: 129 QKDCTKVKCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 171
>HEXP_LEIMA (Q04832) DNA-binding protein HEXBP (Hexamer-binding
protein)
Length = 271
Score = 61.2 bits (147), Expect = 1e-08
Identities = 28/76 (36%), Positives = 35/76 (45%), Gaps = 14/76 (18%)
Query: 426 KKDAAEIVCFNCGEKGHKSNACPEEIKK-------CVRCGKKGHVVADCNRT-------D 471
K D CF CGE+GH S CP E + C RCG+ GH+ DC +
Sbjct: 37 KGDERSTTCFRCGEEGHMSRECPNEARSGAAGAMTCFRCGEAGHMSRDCPNSAKPGAAKG 96
Query: 472 IVCFNCNGEGHISSQC 487
C+ C EGH+S C
Sbjct: 97 FECYKCGQEGHLSRDC 112
Score = 57.8 bits (138), Expect = 1e-07
Identities = 28/82 (34%), Positives = 41/82 (49%), Gaps = 16/82 (19%)
Query: 420 DVRRPKKKDAAEIVCFNCGEKGHKSNACPEEIKK-------CVRCGKKGHVVADCNRT-- 470
DV+RP+ + + C NCG++GH + CPE K C RCG++GH+ +C
Sbjct: 6 DVKRPRTESSTS--CRNCGKEGHYARECPEADSKGDERSTTCFRCGEEGHMSRECPNEAR 63
Query: 471 -----DIVCFNCNGEGHISSQC 487
+ CF C GH+S C
Sbjct: 64 SGAAGAMTCFRCGEAGHMSRDC 85
Score = 53.1 bits (126), Expect = 3e-06
Identities = 24/75 (32%), Positives = 35/75 (46%), Gaps = 14/75 (18%)
Query: 429 AAEIVCFNCGEKGHKSNACPE--------EIKKCVRCGKKGHVVADC------NRTDIVC 474
A + C+ CG+ GH S CP +KC +CG+ GH+ +C D C
Sbjct: 165 AGDRTCYKCGDAGHISRDCPNGQGGYSGAGDRKCYKCGESGHMSRECPSAGSTGSGDRAC 224
Query: 475 FNCNGEGHISSQCTQ 489
+ C GHIS +C +
Sbjct: 225 YKCGKPGHISRECPE 239
Score = 53.1 bits (126), Expect = 3e-06
Identities = 25/76 (32%), Positives = 32/76 (41%), Gaps = 17/76 (22%)
Query: 429 AAEIVCFNCGEKGHKSNACPEE------IKKCVRCGKKGHVVADCNRT-----------D 471
A + C+ CGE GH S CP + C +CGK GH+ +C D
Sbjct: 193 AGDRKCYKCGESGHMSRECPSAGSTGSGDRACYKCGKPGHISRECPEAGGSYGGSRGGGD 252
Query: 472 IVCFNCNGEGHISSQC 487
C+ C GHIS C
Sbjct: 253 RTCYKCGEAGHISRDC 268
Score = 45.4 bits (106), Expect = 6e-04
Identities = 23/85 (27%), Positives = 30/85 (35%), Gaps = 31/85 (36%)
Query: 434 CFNCGEKGHKSNACPEE-----------------------IKKCVRCGKKGHVVADC--- 467
C+ CG++GH S CP + C +CG GH+ DC
Sbjct: 99 CYKCGQEGHLSRDCPSSQGGSRGGYGQKRGRSGAQGGYSGDRTCYKCGDAGHISRDCPNG 158
Query: 468 -----NRTDIVCFNCNGEGHISSQC 487
D C+ C GHIS C
Sbjct: 159 QGGYSGAGDRTCYKCGDAGHISRDC 183
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.138 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 93,643,851
Number of Sequences: 164201
Number of extensions: 4064217
Number of successful extensions: 10730
Number of sequences better than 10.0: 200
Number of HSP's better than 10.0 without gapping: 96
Number of HSP's successfully gapped in prelim test: 104
Number of HSP's that attempted gapping in prelim test: 10066
Number of HSP's gapped (non-prelim): 520
length of query: 805
length of database: 59,974,054
effective HSP length: 118
effective length of query: 687
effective length of database: 40,598,336
effective search space: 27891056832
effective search space used: 27891056832
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 70 (31.6 bits)
Medicago: description of AC147714.1


