

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147712.16 + phase: 0
(236 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
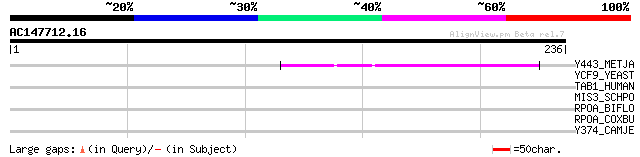
Score E
Sequences producing significant alignments: (bits) Value
Y443_METJA (Q57885) Hypothetical protein MJ0443 45 1e-04
YCF9_YEAST (P25586) Hypothetical 37.2 kDa protein in CHA1-PRD1 i... 41 0.002
TAB1_HUMAN (Q15750) Mitogen-activated protein kinase kinase kina... 36 0.070
MIS3_SCHPO (O74777) Ribosomal RNA assembly protein mis3 33 0.45
RPOA_BIFLO (Q8G3Z3) DNA-directed RNA polymerase alpha chain (EC ... 33 0.77
RPOA_COXBU (Q83EQ2) DNA-directed RNA polymerase alpha chain (EC ... 32 1.7
Y374_CAMJE (Q9PIC8) Hypothetical UPF0234 protein Cj0374 30 5.0
>Y443_METJA (Q57885) Hypothetical protein MJ0443
Length = 227
Score = 45.1 bits (105), Expect = 1e-04
Identities = 30/110 (27%), Positives = 57/110 (51%), Gaps = 2/110 (1%)
Query: 116 KCADFVHAFMLGFDVIDAIAILRLDELYVESFEIKDVKTLRGDHLSRAIGRLSGKGGKTK 175
K D V A GF+ A+ ++ DE +E +I+D + + + R GR+ GK GK++
Sbjct: 72 KARDIVRAIGRGFNPEIALKLVS-DEYVLEVIDIEDYAS-SDNSIRRLKGRVIGKEGKSR 129
Query: 176 FTIENASKTRIVIADTKIHILGSFANIKIARDSLCYLIMGSPAGKVYSKL 225
IE+ + + + + I+G ++IA++++ L+ G+ K Y L
Sbjct: 130 RYIESLTGANVSVYGNTVAIVGEHEPVQIAKEAVEMLLRGASHAKTYKFL 179
>YCF9_YEAST (P25586) Hypothetical 37.2 kDa protein in CHA1-PRD1
intergenic region
Length = 316
Score = 41.2 bits (95), Expect = 0.002
Identities = 34/142 (23%), Positives = 63/142 (43%), Gaps = 3/142 (2%)
Query: 65 PPHRYTPLKKVWMDIYTPVFEQMKIDIRMNLKGRKVELKTRHDTPDISNLQKCADFVHAF 124
P +R + LK +W D+ T ++ I ++L + +KT T D + + K D +
Sbjct: 46 PKYRESYLKTIWNDV-TRALDKHNIACVLDLVEGSMTVKTTRKTYDPAIILKARDLIKLL 104
Query: 125 MLGFDVIDAIAILRLDELYVESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKFTIENASKT 184
A+ IL+ D++ + +I + T + + R RL G G T +E +K
Sbjct: 105 ARSVPFPQAVKILQ-DDMACDVIKIGNFVTNKERFVKRR-QRLVGPNGNTLKALELLTKC 162
Query: 185 RIVIADTKIHILGSFANIKIAR 206
I++ + +G F +K R
Sbjct: 163 YILVQGNTVSAMGPFKGLKEVR 184
>TAB1_HUMAN (Q15750) Mitogen-activated protein kinase kinase kinase
7 interacting protein 1 (TAK1-binding protein 1)
Length = 504
Score = 36.2 bits (82), Expect = 0.070
Identities = 19/59 (32%), Positives = 35/59 (59%)
Query: 136 ILRLDELYVESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKFTIENASKTRIVIADTKIH 194
+ RL +L +++ +IK V + G +R IG K G T + +A+K++ +IA+ +IH
Sbjct: 215 LFRLSQLGLDAGKIKQVGIICGQESTRRIGDYKVKYGYTDIDLLSAAKSKPIIAEPEIH 273
>MIS3_SCHPO (O74777) Ribosomal RNA assembly protein mis3
Length = 327
Score = 33.5 bits (75), Expect = 0.45
Identities = 29/142 (20%), Positives = 63/142 (43%), Gaps = 3/142 (2%)
Query: 65 PPHRYTPLKKVWMDIYTPVFEQMKIDIRMNLKGRKVELKTRHDTPDISNLQKCADFVHAF 124
P +R L++VW + T ++ I ++L + +KT T D ++ D +
Sbjct: 59 PKYREKYLREVWPHV-TRALDKFGITCVLDLVEGSMTVKTTRKTFDPYSILDARDLIKLL 117
Query: 125 MLGFDVIDAIAILRLDELYVESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKFTIENASKT 184
A+ I++ D + + +I ++ + + R RL G G+T +E ++
Sbjct: 118 ARSVPFPQAVKIMQ-DGVACDIIKIGNILRNKERFVKRR-QRLIGTNGQTLKALELLTQC 175
Query: 185 RIVIADTKIHILGSFANIKIAR 206
I++ T + ++G + +K R
Sbjct: 176 YILVQGTTVAVMGGYKGLKEVR 197
>RPOA_BIFLO (Q8G3Z3) DNA-directed RNA polymerase alpha chain (EC
2.7.7.6) (RNAP alpha subunit) (Transcriptase alpha
chain) (RNA polymerase alpha subunit)
Length = 331
Score = 32.7 bits (73), Expect = 0.77
Identities = 28/102 (27%), Positives = 47/102 (45%), Gaps = 28/102 (27%)
Query: 32 LSEQKVLPPKPKF--EPLKP---------------HEMPGAAVQFRKVSVPPHRYTPLKK 74
L+E+ + P + +F EPL+P +PGAAV ++S H +T L
Sbjct: 9 LTEESLNPQRSRFTIEPLEPGFGYTLGNSLRRTLLSSIPGAAVTSVRISGALHEFTTLPG 68
Query: 75 VWMDIYTPVFEQMKIDIRMNLKGRKVELKTRHDTPDISNLQK 116
V D+ +I +N+KG + L + +D P + L+K
Sbjct: 69 VQEDV---------TEILLNIKG--IVLTSEYDEPVVMYLRK 99
>RPOA_COXBU (Q83EQ2) DNA-directed RNA polymerase alpha chain (EC
2.7.7.6) (RNAP alpha subunit) (Transcriptase alpha
chain) (RNA polymerase alpha subunit)
Length = 327
Score = 31.6 bits (70), Expect = 1.7
Identities = 17/56 (30%), Positives = 30/56 (53%), Gaps = 9/56 (16%)
Query: 52 MPGAAVQFRKVSVPPHRYTPLKKVWMDIYTPVFEQMKIDIRMNLKGRKVELKTRHD 107
MPGAA+ ++ H Y+ ++ V D+ +D+ +NLKG ++L+ R D
Sbjct: 51 MPGAAIVQAEIDGVLHEYSSIEGVREDV---------VDVLLNLKGVAIKLEGRED 97
>Y374_CAMJE (Q9PIC8) Hypothetical UPF0234 protein Cj0374
Length = 163
Score = 30.0 bits (66), Expect = 5.0
Identities = 35/123 (28%), Positives = 53/123 (42%), Gaps = 15/123 (12%)
Query: 84 FEQMK--IDIRMNLKGRKVELKTRHDTPDISNLQKCADFVHAFMLGFDVIDAIAILRLDE 141
FEQ K +D R +LKG K E+ D+S + + DV+ I I +L +
Sbjct: 22 FEQAKKELDSRYDLKGIKCEI-------DLSEKESIFKLSSSSEGKLDVLKDIVISKLIK 74
Query: 142 LYVESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKFTIENASKTRIVIADTKIHILGSFAN 201
+ IK++ G A+ RL+ K ENA K VI D+K+ + S
Sbjct: 75 RGINPKAIKELSRESG-----AMFRLNLKANDA-IDSENAKKINKVIKDSKLKVNSSIRG 128
Query: 202 IKI 204
+I
Sbjct: 129 EEI 131
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.138 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,445,600
Number of Sequences: 164201
Number of extensions: 1054107
Number of successful extensions: 2559
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 2556
Number of HSP's gapped (non-prelim): 7
length of query: 236
length of database: 59,974,054
effective HSP length: 107
effective length of query: 129
effective length of database: 42,404,547
effective search space: 5470186563
effective search space used: 5470186563
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 64 (29.3 bits)
Medicago: description of AC147712.16


