

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147538.3 - phase: 0
(123 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
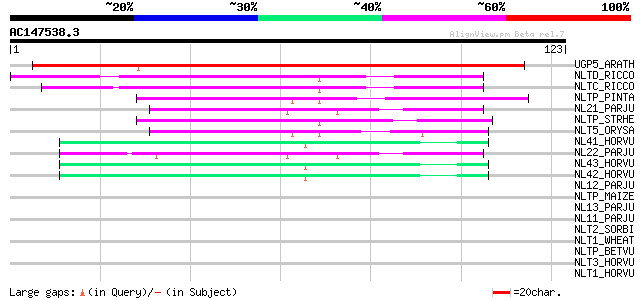
Score E
Sequences producing significant alignments: (bits) Value
UGP5_ARATH (Q9C7F7) Uncharacterized GPI-anchored protein At1g279... 112 2e-25
NLTD_RICCO (Q43119) Nonspecific lipid-transfer protein D, cotyle... 48 3e-06
NLTC_RICCO (P10975) Nonspecific lipid-transfer protein C, cotyle... 48 4e-06
NLTP_PINTA (Q41073) Nonspecific lipid-transfer protein precursor... 47 1e-05
NL21_PARJU (P55958) Probable nonspecific lipid-transfer protein ... 46 2e-05
NLTP_STRHE (O65888) Nonspecific lipid-transfer protein precursor... 44 6e-05
NLT5_ORYSA (O65091) Nonspecific lipid-transfer protein 5 precurs... 43 1e-04
NL41_HORVU (Q43767) Nonspecific lipid-transfer protein 4.1 precu... 42 3e-04
NL22_PARJU (O04403) Probable nonspecific lipid-transfer protein ... 41 4e-04
NL43_HORVU (Q42842) Nonspecific lipid-transfer protein 4.3 precu... 40 7e-04
NL42_HORVU (Q43875) Nonspecific lipid-transfer protein 4.2 precu... 40 7e-04
NL12_PARJU (O04404) Probable nonspecific lipid-transfer protein ... 40 0.001
NLTP_MAIZE (P19656) Nonspecific lipid-transfer protein precursor... 39 0.002
NL13_PARJU (Q40905) Probable nonspecific lipid-transfer protein ... 39 0.003
NL11_PARJU (P43217) Probable nonspecific lipid-transfer protein ... 39 0.003
NLT2_SORBI (Q43194) Nonspecific lipid-transfer protein 2 precurs... 38 0.004
NLT1_WHEAT (P24296) Nonspecific lipid-transfer protein precursor... 38 0.005
NLTP_BETVU (Q43748) Nonspecific lipid-transfer protein precursor... 37 0.006
NLT3_HORVU (Q43766) Nonspecific lipid-transfer protein 3 precurs... 37 0.008
NLT1_HORVU (P07597) Nonspecific lipid-transfer protein 1 precurs... 36 0.013
>UGP5_ARATH (Q9C7F7) Uncharacterized GPI-anchored protein At1g27950
precursor
Length = 193
Score = 112 bits (279), Expect = 2e-25
Identities = 52/111 (46%), Positives = 70/111 (62%), Gaps = 2/111 (1%)
Query: 6 HQMFMCLCVLALIIGGCNGAED--LASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANS 63
H + + + ++A I A LA +C QKV CLDFATGKA P K+CCDA
Sbjct: 7 HLVLVTMTIVASIAAAAPAAPGGALADECNQDFQKVTLCLDFATGKATIPSKKCCDAVED 66
Query: 64 IKATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCP 114
IK DP+CLC++IQQ G K +G+QEDKL+QLPT C ++ A+I++CP
Sbjct: 67 IKERDPKCLCFVIQQAKTGGQALKDLGVQEDKLIQLPTSCQLHNASITNCP 117
>NLTD_RICCO (Q43119) Nonspecific lipid-transfer protein D,
cotyledon-specific isoform precursor (NS-LTP D)
Length = 116
Score = 48.1 bits (113), Expect = 3e-06
Identities = 30/110 (27%), Positives = 48/110 (43%), Gaps = 15/110 (13%)
Query: 1 MKNQQHQMFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDA 60
MKN +F L L + + A C +V K C+ FATGK P CC
Sbjct: 1 MKNIFFSVFFLLSFLLCLAN----VSEAAVPCSTVDMKAAACVGFATGKDSKPSSACCTG 56
Query: 61 ANSIKAT-----DPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHV 105
+ T D + +C ++ + SKS+GI++ L ++P C++
Sbjct: 57 LQQLAQTVKSVDDKKAICRCLKAS------SKSLGIKDQFLSKIPAACNI 100
>NLTC_RICCO (P10975) Nonspecific lipid-transfer protein C,
cotyledon-specific isoform precursor (NS-LTP C)
(Phospholipid transfer protein) (PLTP)
Length = 116
Score = 47.8 bits (112), Expect = 4e-06
Identities = 28/103 (27%), Positives = 49/103 (47%), Gaps = 12/103 (11%)
Query: 8 MFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKAT 67
+F L +L+ + N + A C +V K C+ FATGK P + CC + T
Sbjct: 5 VFSVLLLLSFLFCLAN-TNEAAVPCSTVDMKAAACVGFATGKDSKPSQACCTGLQQLAQT 63
Query: 68 -----DPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHV 105
D + +C ++ + SKS+GI++ L ++P C++
Sbjct: 64 VKTVDDKKAICRCLKAS------SKSLGIKDQFLSKIPAACNI 100
>NLTP_PINTA (Q41073) Nonspecific lipid-transfer protein precursor
(LTP)
Length = 123
Score = 46.6 bits (109), Expect = 1e-05
Identities = 27/92 (29%), Positives = 43/92 (46%), Gaps = 11/92 (11%)
Query: 29 ASKCGSVVQKVIPCLDFATGKAPTPKKECCDAA----NSIKAT-DPECLCYIIQQTHKGS 83
A C VV + PC + G A TP CC + + +KAT D + +C ++
Sbjct: 31 AISCNQVVSAMTPCATYLIGNAATPAATCCPSIRGLDSQVKATPDRQAVCNCLK------ 84
Query: 84 PESKSMGIQEDKLLQLPTVCHVNGANISDCPS 115
++KS G++ K LP +C V N+ P+
Sbjct: 85 TQAKSYGVKLGKAANLPGLCKVTDLNVPISPN 116
>NL21_PARJU (P55958) Probable nonspecific lipid-transfer protein 2
precursor (LTP 2) (Major pollen allergen Par j 2.0101)
(Par j II) (P2 protein)
Length = 133
Score = 45.8 bits (107), Expect = 2e-05
Identities = 25/79 (31%), Positives = 39/79 (48%), Gaps = 10/79 (12%)
Query: 32 CGSVVQKVIPCLDFATGKAPTPKKECCDA----ANSIKATDPEC-LCYIIQQTHKGSPES 86
CG VVQ ++PCL F G+ P KECC + +K T+ + C I + KG
Sbjct: 35 CGKVVQDIMPCLHFVKGEEKEPSKECCSGTKKLSEEVKTTEQKREACKCIVRATKG---- 90
Query: 87 KSMGIQEDKLLQLPTVCHV 105
GI+ + + ++P C +
Sbjct: 91 -ISGIKNELVAEVPKKCDI 108
>NLTP_STRHE (O65888) Nonspecific lipid-transfer protein precursor
(LTP) (Fragment)
Length = 118
Score = 43.9 bits (102), Expect = 6e-05
Identities = 25/84 (29%), Positives = 37/84 (43%), Gaps = 10/84 (11%)
Query: 29 ASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKAT-----DPECLCYIIQQTHKGS 83
A CG+V K C+ FATGK P CC + + T D + +C ++ K
Sbjct: 7 ALPCGTVDMKAASCISFATGKDKKPSAACCSGLHPLAQTVKTVEDNKAICRCLKTAIKNF 66
Query: 84 PESKSMGIQEDKLLQLPTVCHVNG 107
+Q+ L Q+PT C + G
Sbjct: 67 SR-----VQDRFLGQIPTPCKIKG 85
>NLT5_ORYSA (O65091) Nonspecific lipid-transfer protein 5 precursor
(LTP 5)
Length = 117
Score = 42.7 bits (99), Expect = 1e-04
Identities = 27/81 (33%), Positives = 41/81 (50%), Gaps = 12/81 (14%)
Query: 32 CGSVVQKVIPCLDFATGKAPTPKKECCDAA---NSIKAT--DPECLCYIIQQTHKGSPES 86
CG VV PC+ +A G+ P CCD NS AT D + C ++Q ++
Sbjct: 28 CGQVVSTWAPCIMYADGEGVAPTGGCCDGVRTLNSAAATTADRQTTCACLKQ------QT 81
Query: 87 KSMG-IQEDKLLQLPTVCHVN 106
K+MG ++ D + +P+ C VN
Sbjct: 82 KAMGRLRPDHVAGIPSKCGVN 102
>NL41_HORVU (Q43767) Nonspecific lipid-transfer protein 4.1
precursor (LTP 4.1) (CW21) (CW-21)
Length = 115
Score = 41.6 bits (96), Expect = 3e-04
Identities = 26/100 (26%), Positives = 39/100 (39%), Gaps = 13/100 (13%)
Query: 12 LCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSI-----KA 66
L ++AL+ A D A CG V + PC+ +A G P CC +
Sbjct: 9 LVLVALVAAMLLVAADAAISCGQVSSALSPCISYARGNGAKPPAACCSGVKRLAGAAQST 68
Query: 67 TDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
D + C I+ G K+ GI P++C V+
Sbjct: 69 ADKQAACKCIKSAAGGLNAGKAAGI--------PSMCGVS 100
>NL22_PARJU (O04403) Probable nonspecific lipid-transfer protein 2
precursor (LTP 2) (Major pollen allergen Par j 2.0102)
(Par j II) (P8 protein)
Length = 133
Score = 41.2 bits (95), Expect = 4e-04
Identities = 31/106 (29%), Positives = 47/106 (44%), Gaps = 18/106 (16%)
Query: 12 LCVLALIIGGCNGAEDLASK-------CGSVVQKVIPCLDFATGKAPTPKKECCDA---- 60
L V+A + + AE LAS CG VV ++PCL F G+ P K CC
Sbjct: 9 LVVIAAALAWTSSAE-LASAPAPGEGPCGKVVHHIMPCLKFVKGEEKEPSKSCCSGTKKL 67
Query: 61 ANSIKATDPEC-LCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHV 105
+ +K T+ + C I KG GI+ + + ++P C +
Sbjct: 68 SEEVKTTEQKREACKCIVAATKG-----ISGIKNELVAEVPKKCGI 108
>NL43_HORVU (Q42842) Nonspecific lipid-transfer protein 4.3
precursor (LTP 4.3)
Length = 115
Score = 40.4 bits (93), Expect = 7e-04
Identities = 25/100 (25%), Positives = 39/100 (39%), Gaps = 13/100 (13%)
Query: 12 LCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSI-----KA 66
L ++A++ A D A CG V + PC+ +A G P CC +
Sbjct: 9 LVLVAMVAAMLLVATDAAISCGQVSSALSPCISYARGNGAKPPVACCSGVKRLAGAAQST 68
Query: 67 TDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
D + C I+ G K+ GI P++C V+
Sbjct: 69 ADKQAACKCIKSAAGGLNAGKAAGI--------PSMCGVS 100
>NL42_HORVU (Q43875) Nonspecific lipid-transfer protein 4.2
precursor (LTP 4.2) (Low-temperature-responsive protein
4.9)
Length = 115
Score = 40.4 bits (93), Expect = 7e-04
Identities = 25/100 (25%), Positives = 39/100 (39%), Gaps = 13/100 (13%)
Query: 12 LCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSI-----KA 66
L ++A++ A D A CG V + PC+ +A G P CC +
Sbjct: 9 LVLVAMVAAMLIVATDAAISCGQVSSALSPCISYARGNGAKPPVACCSGVKRLAGAAQST 68
Query: 67 TDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
D + C I+ G K+ GI P++C V+
Sbjct: 69 ADKQAACKCIKSAAGGLNAGKAAGI--------PSMCGVS 100
>NL12_PARJU (O04404) Probable nonspecific lipid-transfer protein 1
precursor (LTP) (Major pollen allergen Par j 1.0102)
(Par j I) (P9 protein)
Length = 176
Score = 39.7 bits (91), Expect = 0.001
Identities = 15/33 (45%), Positives = 20/33 (60%)
Query: 32 CGSVVQKVIPCLDFATGKAPTPKKECCDAANSI 64
CG+VV+ ++PCL F GK P K CC A +
Sbjct: 41 CGTVVRALMPCLPFVQGKEKEPSKGCCSGAKRL 73
>NLTP_MAIZE (P19656) Nonspecific lipid-transfer protein precursor
(LTP) (Phospholipid transfer protein) (PLTP) (Allergen
Zea m 14)
Length = 120
Score = 38.9 bits (89), Expect = 0.002
Identities = 25/111 (22%), Positives = 41/111 (36%), Gaps = 11/111 (9%)
Query: 1 MKNQQHQMFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDA 60
M Q + V+AL++ +E A CG V + PC+ +A G+ P CC
Sbjct: 1 MARTQQLAVVATAVVALVLLAAATSE-AAISCGQVASAIAPCISYARGQGSGPSAGCCSG 59
Query: 61 ANSIK-----ATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
S+ D C ++ G G+ +P+ C V+
Sbjct: 60 VRSLNNAARTTADRRAACNCLKNAAAG-----VSGLNAGNAASIPSKCGVS 105
>NL13_PARJU (Q40905) Probable nonspecific lipid-transfer protein 1
precursor (LTP) (Major pollen allergen Par j 1.0201)
(Par j I) (P1 protein)
Length = 138
Score = 38.5 bits (88), Expect = 0.003
Identities = 15/33 (45%), Positives = 19/33 (57%)
Query: 32 CGSVVQKVIPCLDFATGKAPTPKKECCDAANSI 64
CG+VV ++PCL F GK P K CC A +
Sbjct: 40 CGTVVGALMPCLPFVQGKEKEPSKGCCSGAKRL 72
>NL11_PARJU (P43217) Probable nonspecific lipid-transfer protein
(LTP) (Major pollen allergen Par j 1.0101) (Par j I)
(P5 protein) (Fragment)
Length = 139
Score = 38.5 bits (88), Expect = 0.003
Identities = 14/33 (42%), Positives = 20/33 (60%)
Query: 32 CGSVVQKVIPCLDFATGKAPTPKKECCDAANSI 64
CG++V+ ++PCL F GK P K CC A +
Sbjct: 4 CGTMVRALMPCLPFVQGKEKEPSKGCCSGAKRL 36
>NLT2_SORBI (Q43194) Nonspecific lipid-transfer protein 2 precursor
(LTP 2)
Length = 122
Score = 38.1 bits (87), Expect = 0.004
Identities = 21/85 (24%), Positives = 34/85 (39%), Gaps = 10/85 (11%)
Query: 27 DLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKA-----TDPECLCYIIQQTHK 81
+ A CG V + PCL +A G+ P CC S+ + D C ++ +
Sbjct: 28 EAAVTCGQVSSAIGPCLSYARGQGSGPSAGCCSGVRSLNSAARTTADRRAACNCLKNAAR 87
Query: 82 GSPESKSMGIQEDKLLQLPTVCHVN 106
G G+ K +P+ C V+
Sbjct: 88 G-----IRGLNVGKAASIPSKCGVS 107
>NLT1_WHEAT (P24296) Nonspecific lipid-transfer protein precursor
(LTP) (Phospholipid transfer protein) (PLTP) (ns-LTP1)
(Fragment)
Length = 113
Score = 37.7 bits (86), Expect = 0.005
Identities = 27/105 (25%), Positives = 43/105 (40%), Gaps = 12/105 (11%)
Query: 7 QMFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSI-- 64
Q+ + L L++ A +A CG V V PCL + G P P +CCD ++
Sbjct: 2 QVMLMAVALVLMLAAVPRAA-VAIDCGHVDSLVRPCLSYVQG-GPGPSGQCCDGVKNLHN 59
Query: 65 ---KATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
+D + C ++ +G + ED +P C VN
Sbjct: 60 QARSQSDRQSACNCLKGIARG-----IHNLNEDNARSIPPKCGVN 99
>NLTP_BETVU (Q43748) Nonspecific lipid-transfer protein precursor
(LTP)
Length = 117
Score = 37.4 bits (85), Expect = 0.006
Identities = 27/103 (26%), Positives = 41/103 (39%), Gaps = 14/103 (13%)
Query: 9 FMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKA-- 66
F C V+ +++ A CG V K+ PC+ + G AP P CC S+ +
Sbjct: 9 FTCALVMCMMVAAPLAE---AITCGLVASKLAPCIGYLQG-APGPSAACCGGIKSLNSAA 64
Query: 67 ---TDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
D + C ++ S + GI K LP C V+
Sbjct: 65 ASPADRKTACTCLK-----SAATSIKGINYGKAASLPRQCGVS 102
>NLT3_HORVU (Q43766) Nonspecific lipid-transfer protein 3 precursor
(LTP 3) (CW20) (CW-20) (CW-19)
Length = 118
Score = 37.0 bits (84), Expect = 0.008
Identities = 24/100 (24%), Positives = 39/100 (39%), Gaps = 10/100 (10%)
Query: 12 LCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSI-----KA 66
L ++A++ A D A CG V + PC+ +A G P CC +
Sbjct: 9 LVLVAMVAAMLLVATDAAISCGQVSSALSPCISYARGNGAKPPVACCSGVKRLAGAAQST 68
Query: 67 TDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
D + C ++ S + GI K+ +P C V+
Sbjct: 69 ADKQAACRCLK-----SLATSIKGINMGKVSGVPGKCGVS 103
>NLT1_HORVU (P07597) Nonspecific lipid-transfer protein 1 precursor
(LTP 1) (Probable amylase/protease inhibitor)
Length = 117
Score = 36.2 bits (82), Expect = 0.013
Identities = 25/105 (23%), Positives = 45/105 (42%), Gaps = 12/105 (11%)
Query: 7 QMFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSI-- 64
Q+ + L L++ A +A CG V K+ PCL + G P P ECC+ +
Sbjct: 5 QVLLMAAALVLMLTAAPRAA-VALNCGQVDSKMKPCLTYVQG-GPGPSGECCNGVRDLHN 62
Query: 65 ---KATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
+ D + +C ++ +G + + +P+ C+VN
Sbjct: 63 QAQSSGDRQTVCNCLKGIARG-----IHNLNLNNAASIPSKCNVN 102
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,511,428
Number of Sequences: 164201
Number of extensions: 545131
Number of successful extensions: 1253
Number of sequences better than 10.0: 85
Number of HSP's better than 10.0 without gapping: 33
Number of HSP's successfully gapped in prelim test: 52
Number of HSP's that attempted gapping in prelim test: 1210
Number of HSP's gapped (non-prelim): 91
length of query: 123
length of database: 59,974,054
effective HSP length: 99
effective length of query: 24
effective length of database: 43,718,155
effective search space: 1049235720
effective search space used: 1049235720
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC147538.3


