

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
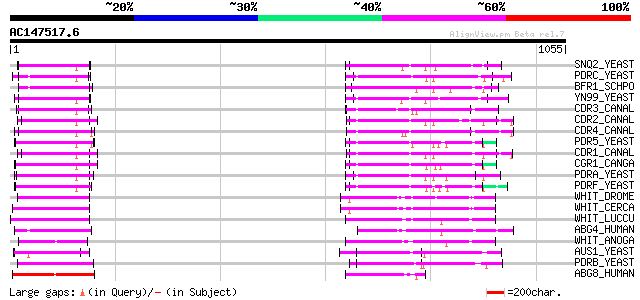
Query= AC147517.6 + phase: 0 /pseudo
(1055 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
SNQ2_YEAST (P32568) SNQ2 protein 168 8e-41
PDRC_YEAST (Q02785) ATP-dependent permease PDR12 163 3e-39
BFR1_SCHPO (P41820) Brefeldin A resistance protein 162 4e-39
YN99_YEAST (P53756) Probable ATP-dependent transporter YNR070W 160 2e-38
CDR3_CANAL (O42690) Opaque-specific ABC transporter CDR3 160 2e-38
CDR2_CANAL (P78595) Multidrug resistance protein CDR2 157 2e-37
CDR4_CANAL (O74676) ABC transporter CDR4 154 1e-36
PDR5_YEAST (P33302) Suppressor of toxicity of sporidesmin 154 2e-36
CDR1_CANAL (P43071) Multidrug resistance protein CDR1 154 2e-36
CGR1_CANGA (O74208) ATP-binding cassette transporter CGR1 (Pleom... 151 8e-36
PDRA_YEAST (P51533) ATP-dependent permease PDR10 150 2e-35
PDRF_YEAST (Q04182) ATP-dependent permease PDR15 149 4e-35
WHIT_DROME (P10090) White protein 117 2e-25
WHIT_CERCA (Q17320) White protein 117 2e-25
WHIT_LUCCU (Q05360) White protein 115 6e-25
ABG4_HUMAN (Q9H172) ATP-binding cassette, sub-family G, member 4 114 1e-24
WHIT_ANOGA (Q27256) White protein 107 2e-22
AUS1_YEAST (Q08409) ATP-dependent permease AUS1 105 6e-22
PDRB_YEAST (P40550) ATP-dependent permease PDR11 103 2e-21
ABG8_HUMAN (Q9H221) ATP-binding cassette, sub-family G, member 8... 102 5e-21
>SNQ2_YEAST (P32568) SNQ2 protein
Length = 1501
Score = 168 bits (425), Expect = 8e-41
Identities = 101/303 (33%), Positives = 172/303 (56%), Gaps = 11/303 (3%)
Query: 638 QKVMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSII-FMDEPTSGLDARA 696
+K++ ++ ++ ALVG G CGL++EQRK+L++ VELVA P ++ F+DEPTSGLD+++
Sbjct: 970 EKIIRVLGMEEYAEALVGEVG-CGLNVEQRKKLSIGVELVAKPDLLLFLDEPTSGLDSQS 1028
Query: 697 AAVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLIN 756
+ +++ +R G++++CTIHQPS +FE FD LLLL++GGQ +Y G +G NS+ ++N
Sbjct: 1029 SWAIIQLLRKLSKAGQSILCTIHQPSATLFEEFDRLLLLRKGGQTVYFGDIGKNSATILN 1088
Query: 757 HFEGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNSELYRINKA----LVKE 812
+FE G RK NPA ++LE + + D+ E + NS + K L+ +
Sbjct: 1089 YFER-NGARKCDSSENPAEYILEAIGAGATASVKEDWHEKWLNSVEFEQTKEKVQDLIND 1147
Query: 813 LSAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVAVLLGSM 872
LS S+ PS+Y+ S+ Q L + S+WR+ Y + + + +G
Sbjct: 1148 LSKQETKSEVGDKPSKYATSYAYQFRYVLIRTSTSFWRSLNYIMSKMMLMLVGGLYIGFT 1207
Query: 873 FWDLGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVERTVFYRESAAGNVFYFS 932
F+++G + L NAM + + ++IL N +Q R +F + N+F++S
Sbjct: 1208 FFNVG---KSYVGLQNAMFAAFISIIL-SAPAMNQIQGRAIASRELFEVRESQSNMFHWS 1263
Query: 933 ICL 935
+ L
Sbjct: 1264 LVL 1266
Score = 83.6 bits (205), Expect = 3e-15
Identities = 42/138 (30%), Positives = 68/138 (48%)
Query: 16 DYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDSSTT 75
D + GL +T VGN +R +SGG++KR++ E L D + GLD+ST
Sbjct: 285 DLYATIFGLRHTYNTKVGNDFVRGVSGGERKRVSIAEALAAKGSIYCWDNATRGLDASTA 344
Query: 76 FQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEFFESI 135
+ ++R ++LK T +++ Q Y FD + +L IY G +F +
Sbjct: 345 LEYAKAIRIMTNLLKSTAFVTIYQASENIYETFDKVTVLYSGKQIYFGLIHEAKPYFAKM 404
Query: 136 GFKCPNRKGVADFLQEVT 153
G+ CP R+ A+FL +T
Sbjct: 405 GYLCPPRQATAEFLTALT 422
Score = 69.7 bits (169), Expect = 4e-11
Identities = 73/289 (25%), Positives = 124/289 (42%), Gaps = 35/289 (12%)
Query: 646 LKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVR 705
L+ N VG V G+S +RKR+++A L A SI D T GLDA A + +R
Sbjct: 293 LRHTYNTKVGNDFVRGVSGGERKRVSIAEALAAKGSIYCWDNATRGLDASTALEYAKAIR 352
Query: 706 NTVDTGR-TVVCTIHQPSIDIFESFDELLLLKQGGQEIYVG----------------PLG 748
+ + T TI+Q S +I+E+FD++ +L G++IY G P
Sbjct: 353 IMTNLLKSTAFVTIYQASENIYETFDKVTVL-YSGKQIYFGLIHEAKPYFAKMGYLCPPR 411
Query: 749 HNSSNLINHFEGIQGVRKIKDGYNPATWMLEVTTSSKEREL----GIDFAELYKNSELY- 803
++ + G IK GY +V +++E E +FA++ K+ Y
Sbjct: 412 QATAEFLTALTDPNGFHLIKPGYEN-----KVPRTAEEFETYWLNSPEFAQMKKDIAAYK 466
Query: 804 -RINKALVKEL---SAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRF 859
++N KE+ S SK S Y+ S++ Q C + + N Y I
Sbjct: 467 EKVNTEKTKEVYDESMAQEKSKYTRKKSYYTVSYWEQVKLCTQRGFQRIYGNKSYTVINV 526
Query: 860 LYSTAVAVLLGSMFWDLGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSV 908
+ + + GS+F++ S F+ G +Y A++ +M ++
Sbjct: 527 CSAIIQSFITGSLFYNTPS---STSGAFSRGGVLYFALLYYSLMGLANI 572
Score = 55.1 bits (131), Expect = 1e-06
Identities = 40/141 (28%), Positives = 78/141 (54%), Gaps = 9/141 (6%)
Query: 18 VLRVLGLEICADTVVGNAMIRAISGGQKKRLTTG-EMLVGPTKALFMDEISTGLDSSTTF 76
++RVLG+E A+ +VG ++ Q+K+L+ G E++ P LF+DE ++GLDS +++
Sbjct: 972 IIRVLGMEEYAEALVGEVGC-GLNVEQRKKLSIGVELVAKPDLLLFLDEPTSGLDSQSSW 1030
Query: 77 QIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILL-SDSHIIYQGP----REHVLEF 131
I+ +R+ + +++ ++ QP + FD ++LL +Y G +L +
Sbjct: 1031 AIIQLLRK-LSKAGQSILCTIHQPSATLFEEFDRLLLLRKGGQTVYFGDIGKNSATILNY 1089
Query: 132 FESIGF-KCPNRKGVADFLQE 151
FE G KC + + A+++ E
Sbjct: 1090 FERNGARKCDSSENPAEYILE 1110
>PDRC_YEAST (Q02785) ATP-dependent permease PDR12
Length = 1511
Score = 163 bits (412), Expect = 3e-39
Identities = 103/326 (31%), Positives = 176/326 (53%), Gaps = 16/326 (4%)
Query: 638 QKVMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSII-FMDEPTSGLDARA 696
+K++ L+ ++ ALVG G GL++EQRK+L++ VELVA PS++ F+DEPTSGLD+++
Sbjct: 960 EKIITLLGMQNYAEALVGKTGR-GLNVEQRKKLSIGVELVAKPSLLLFLDEPTSGLDSQS 1018
Query: 697 AAVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLIN 756
A +++ +R D+G++++CTIHQPS +FE FD LLLLK+GG+ +Y G +G NS L+
Sbjct: 1019 AWSIVQFMRALADSGQSILCTIHQPSATLFEQFDRLLLLKKGGKMVYFGDIGPNSETLLK 1078
Query: 757 HFEGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNSELYRINKALVKELSAP 816
+FE G+ K NPA ++L + + D+ +L+ S +A V+EL
Sbjct: 1079 YFERQSGM-KCGVSENPAEYILNCIGAGATASVNSDWHDLWLASPECAAARAEVEELHRT 1137
Query: 817 AP---CSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVAVLLGSMF 873
P + D ++++ S+ TQ L + +WR+P Y +F A A+ +G +
Sbjct: 1138 LPGRAVNDDPELATRFAASYMTQIKCVLRRTALQFWRSPVYIRAKFFECVACALFVGLSY 1197
Query: 874 WDLGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVERTVFYRESAAGNVFYFSI 933
+ + + F+++ ++LI + N + R ++ AA N F++S+
Sbjct: 1198 VGVNHSVGGAIEAFSSI----FMLLLIALAMINQLHVFAYDSRELYEVREAASNTFHWSV 1253
Query: 934 CLW----AGKYPSPIC--FCTSCGLW 953
L + S +C C C W
Sbjct: 1254 LLLCHAAVENFWSTLCQFMCFICYYW 1279
Score = 81.3 bits (199), Expect = 1e-14
Identities = 45/148 (30%), Positives = 74/148 (49%)
Query: 6 TEGQKENLITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDE 65
T Q + I D V GL T VGN +R +SGG++KR++ E D
Sbjct: 262 TRKQYVDNIRDMWCTVFGLRHTYATKVGNDFVRGVSGGERKRVSLVEAQAMNASIYSWDN 321
Query: 66 ISTGLDSSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPR 125
+ GLD+ST + ++R +++ + ++++ Q Y LFD +L + IY GP
Sbjct: 322 ATRGLDASTALEFAQAIRTATNMVNNSAIVAIYQAGENIYELFDKTTVLYNGRQIYFGPA 381
Query: 126 EHVLEFFESIGFKCPNRKGVADFLQEVT 153
+ + +F+ +G+ PNR A+FL VT
Sbjct: 382 DKAVGYFQRMGWVKPNRMTSAEFLTSVT 409
Score = 59.3 bits (142), Expect = 5e-08
Identities = 71/302 (23%), Positives = 126/302 (41%), Gaps = 47/302 (15%)
Query: 654 VGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAA---AVVMRTVRNTVDT 710
VG V G+S +RKR+++ N SI D T GLDA A A +RT N V+
Sbjct: 288 VGNDFVRGVSGGERKRVSLVEAQAMNASIYSWDNATRGLDASTALEFAQAIRTATNMVN- 346
Query: 711 GRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHFEGIQGVRKIKDG 770
+ + I+Q +I+E FD+ +L G++IY GP + + +F+ + V+ +
Sbjct: 347 -NSAIVAIYQAGENIYELFDKTTVL-YNGRQIYFGP----ADKAVGYFQRMGWVK--PNR 398
Query: 771 YNPATWMLEVTTSSKERELGI-------------DFAELYKNSELYR------------- 804
A ++ VT + R L I +F E + NSE Y+
Sbjct: 399 MTSAEFLTSVTVDFENRTLDIKPGYEDKVPKSSSEFEEYWLNSEDYQELLRTYDDYQSRH 458
Query: 805 -INKALVK-ELSAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYS 862
+N+ + +++ + SQY +++TQ C+ + + Y +
Sbjct: 459 PVNETRDRLDVAKKQRLQQGQRENSQYVVNYWTQVYYCMIRGFQRVKGDSTYTKVYLSSF 518
Query: 863 TAVAVLLGSMFWDLGSKIEKEQDLFNAMGSMYSAVILIG-------VMNCNSVQPVVAVE 915
A+++GSMF + K + + G M V+L + N S +PV+
Sbjct: 519 LIKALIIGSMFHKIDDKSQSTTAGAYSRGGMLFYVLLFASVTSLAEIGNSFSSRPVIVKH 578
Query: 916 RT 917
++
Sbjct: 579 KS 580
Score = 55.8 bits (133), Expect = 6e-07
Identities = 40/139 (28%), Positives = 78/139 (55%), Gaps = 9/139 (6%)
Query: 18 VLRVLGLEICADTVVGNAMIRAISGGQKKRLTTG-EMLVGPTKALFMDEISTGLDSSTTF 76
++ +LG++ A+ +VG R ++ Q+K+L+ G E++ P+ LF+DE ++GLDS + +
Sbjct: 962 IITLLGMQNYAEALVGKTG-RGLNVEQRKKLSIGVELVAKPSLLLFLDEPTSGLDSQSAW 1020
Query: 77 QIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILL-SDSHIIYQGP----REHVLEF 131
IV MR + +++ ++ QP + FD ++LL ++Y G E +L++
Sbjct: 1021 SIVQFMRALADSGQ-SILCTIHQPSATLFEQFDRLLLLKKGGKMVYFGDIGPNSETLLKY 1079
Query: 132 FE-SIGFKCPNRKGVADFL 149
FE G KC + A+++
Sbjct: 1080 FERQSGMKCGVSENPAEYI 1098
>BFR1_SCHPO (P41820) Brefeldin A resistance protein
Length = 1530
Score = 162 bits (410), Expect = 4e-39
Identities = 98/298 (32%), Positives = 165/298 (54%), Gaps = 12/298 (4%)
Query: 638 QKVMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSII-FMDEPTSGLDARA 696
+ V++L+E++ A++G PG GL++EQRKR T+ VEL A P+++ F+DEPTSGLD+++
Sbjct: 1000 ESVIKLLEMESYAEAIIGTPG-SGLNVEQRKRATIGVELAAKPALLLFLDEPTSGLDSQS 1058
Query: 697 AAVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLIN 756
A ++ +R D G+ ++CTIHQPS +F+ FD LLLL++GG+ +Y G +G +S L+N
Sbjct: 1059 AWSIVCFLRKLADAGQAILCTIHQPSAVLFDQFDRLLLLQKGGKTVYFGDIGEHSKTLLN 1118
Query: 757 HFEGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNSELYRINKALVKELSAP 816
+FE V DG NPA ++L+V + D+ E++ NSE + A + +++A
Sbjct: 1119 YFESHGAVHCPDDG-NPAEYILDVIGAGATATTNRDWHEVWNNSEERKAISAELDKINAS 1177
Query: 817 APCSKDLYFPSQYSRSFFT-----QCMACLWKQHWSYWRNPEYNAIRFLYSTAVAVLLGS 871
S+D S+ RS + Q + + SYWR P + + +G
Sbjct: 1178 FSNSEDKKTLSKEDRSTYAMPLWFQVKMVMTRNFQSYWREPSILMSKLALDIFAGLFIGF 1237
Query: 872 MFWDLGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVERTVFYRESAAGNVF 929
F++ G + Q++ N + +++ A +L V N +QP R VF N++
Sbjct: 1238 TFYNQGLGV---QNIQNKLFAVFMATVL-AVPLINGLQPKFIELRNVFEVREKPSNIY 1291
Score = 87.0 bits (214), Expect = 2e-16
Identities = 48/139 (34%), Positives = 70/139 (49%)
Query: 18 VLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDSSTTFQ 77
+ GL +T VGN +R +SGG++KR+T E D + GLDSST F+
Sbjct: 287 IATAFGLTHTFNTKVGNDFVRGVSGGERKRVTISEGFATRPTIACWDNSTRGLDSSTAFE 346
Query: 78 IVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEFFESIGF 137
VN +R + LK T ++ Q + Y LFD I +L IY GP + ++F +GF
Sbjct: 347 FVNVLRTCANELKMTSFVTAYQASEKIYKLFDRICVLYAGRQIYYGPADKAKQYFLDMGF 406
Query: 138 KCPNRKGVADFLQEVTSRK 156
C R+ DFL ++ K
Sbjct: 407 DCHPRETTPDFLTAISDPK 425
Score = 54.3 bits (129), Expect = 2e-06
Identities = 74/317 (23%), Positives = 126/317 (39%), Gaps = 60/317 (18%)
Query: 651 NALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAA---AVVMRTVRNT 707
N VG V G+S +RKR+T++ P+I D T GLD+ A V+RT N
Sbjct: 298 NTKVGNDFVRGVSGGERKRVTISEGFATRPTIACWDNSTRGLDSSTAFEFVNVLRTCAN- 356
Query: 708 VDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHFEGIQGVRKI 767
+ T T +Q S I++ FD + +L G++IY GP ++
Sbjct: 357 -ELKMTSFVTAYQASEKIYKLFDRICVL-YAGRQIYYGPADKAKQYFLDMGFDCHPRETT 414
Query: 768 KDGY----NPATWMLEVTTSSKERELGIDFAELYKNSELYR------------------- 804
D +P ++ +F ++++NS +Y
Sbjct: 415 PDFLTAISDPKARFPRKGFENRVPRTPDEFEQMWRNSSVYADLMAEMESYDKRWTETTPA 474
Query: 805 -------------INKALVKEL---SAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSY 848
I+ EL SA A SK + S Y+ +F Q CL + Y
Sbjct: 475 SSEAPEKDNFGSDISATTKHELYRQSAVAEKSKRVKDTSPYTVTFSQQLWYCLARSWERY 534
Query: 849 WRNPEY---NAIRFLYSTAVAVLLGSMFWDLGSKIEKEQDLFNAMGSMYSAVI------L 899
+P Y A FL+ ++++GS+F+D+ D+F+ G ++ +++ L
Sbjct: 535 INDPAYIGSMAFAFLFQ---SLIIGSIFYDMKL---NTVDVFSRGGVLFFSILFCALQSL 588
Query: 900 IGVMNCNSVQPVVAVER 916
+ N S +P++A R
Sbjct: 589 SEIANMFSQRPIIAKHR 605
Score = 53.1 bits (126), Expect = 4e-06
Identities = 42/142 (29%), Positives = 77/142 (53%), Gaps = 9/142 (6%)
Query: 18 VLRVLGLEICADTVVGNAMIRAISGGQKKRLTTG-EMLVGPTKALFMDEISTGLDSSTTF 76
V+++L +E A+ ++G ++ Q+KR T G E+ P LF+DE ++GLDS + +
Sbjct: 1002 VIKLLEMESYAEAIIGTPG-SGLNVEQRKRATIGVELAAKPALLLFLDEPTSGLDSQSAW 1060
Query: 77 QIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILL-SDSHIIYQGP-REH---VLEF 131
IV +R+ + ++ ++ QP ++ FD ++LL +Y G EH +L +
Sbjct: 1061 SIVCFLRKLADAGQ-AILCTIHQPSAVLFDQFDRLLLLQKGGKTVYFGDIGEHSKTLLNY 1119
Query: 132 FESIG-FKCPNRKGVADFLQEV 152
FES G CP+ A+++ +V
Sbjct: 1120 FESHGAVHCPDDGNPAEYILDV 1141
>YN99_YEAST (P53756) Probable ATP-dependent transporter YNR070W
Length = 1333
Score = 160 bits (405), Expect = 2e-38
Identities = 102/320 (31%), Positives = 177/320 (54%), Gaps = 17/320 (5%)
Query: 638 QKVMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSII-FMDEPTSGLDARA 696
+K++ ++E++ ALVG G GL++EQRK+L++ VELV P ++ F+DEPTSGLD+++
Sbjct: 846 EKIISILEMQEFSEALVGEIGY-GLNVEQRKKLSIGVELVGKPDLLLFLDEPTSGLDSQS 904
Query: 697 AAVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLIN 756
A V++ ++ G++++CTIHQPS +FE FD LLLL +GGQ IY G +G NSS++I
Sbjct: 905 AWAVVKMLKRLALAGQSILCTIHQPSATLFEQFDRLLLLGKGGQTIYFGEIGKNSSSVIK 964
Query: 757 HFEGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNSELY-RINKA---LVKE 812
+FE G RK + NPA ++LE + + ++ ++++ S Y IN+ ++K+
Sbjct: 965 YFEK-NGARKCQQNENPAEYILEAIGAGATASVQQNWPDIWQKSHEYANINEKINDMIKD 1023
Query: 813 LSAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVAVLLGSM 872
LS+ K S+Y+ S+ Q L + ++WRN Y + + + +G
Sbjct: 1024 LSS-TTLHKTATRASKYATSYSYQFHHVLKRSSLTFWRNLNYIMAKMMLLMISGLFIGFT 1082
Query: 873 FWDLG-SKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVERTVFYRESAAGNVFYF 931
F+ +G + I + LF I+I N +Q V + ++ + N+F++
Sbjct: 1083 FFHVGVNAIGLQNSLFACF-----MAIVISAPATNQIQERATVAKELYEVRESKSNMFHW 1137
Query: 932 SICL---WAGKYPSPICFCT 948
S+ L + + P + F T
Sbjct: 1138 SLLLITHYLNELPYHLLFST 1157
Score = 87.8 bits (216), Expect = 1e-16
Identities = 45/147 (30%), Positives = 76/147 (51%), Gaps = 3/147 (2%)
Query: 10 KENLIT---DYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEI 66
KE IT ++ ++ GL DT VGN I +SGG++KR++ E L D
Sbjct: 147 KEEYITANREFYAKIFGLTHTFDTKVGNDFISGVSGGERKRVSIAEALAAKGSIYCWDNA 206
Query: 67 STGLDSSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPRE 126
+ GLDSST + ++R ++L T ++++ Q Y FD + +L I+ G
Sbjct: 207 TRGLDSSTALEFARAIRTMTNLLGTTALVTVYQASENIYETFDKVTVLYAGRQIFCGKTT 266
Query: 127 HVLEFFESIGFKCPNRKGVADFLQEVT 153
++FE++G+ CP R+ A++L +T
Sbjct: 267 EAKDYFENMGYLCPPRQSTAEYLTAIT 293
Score = 63.9 bits (154), Expect = 2e-09
Identities = 65/280 (23%), Positives = 119/280 (42%), Gaps = 33/280 (11%)
Query: 654 VGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVRNTVD-TGR 712
VG + G+S +RKR+++A L A SI D T GLD+ A R +R + G
Sbjct: 172 VGNDFISGVSGGERKRVSIAEALAAKGSIYCWDNATRGLDSSTALEFARAIRTMTNLLGT 231
Query: 713 TVVCTIHQPSIDIFESFDELLLLKQGGQEI---------------YVGPLGHNSSNLINH 757
T + T++Q S +I+E+FD++ +L G Q Y+ P +++ +
Sbjct: 232 TALVTVYQASENIYETFDKVTVLYAGRQIFCGKTTEAKDYFENMGYLCPPRQSTAEYLTA 291
Query: 758 FEGIQGVRKIKDGYNPATWMLEVTTSSKEREL----GIDFAELYKNSELYR--INKALVK 811
G+ +IK G+ +V ++ E E ++A L + Y+ +N K
Sbjct: 292 ITDPNGLHEIKPGFE-----YQVPHTADEFEKYWLDSPEYARLKGEIQKYKHEVNTEWTK 346
Query: 812 EL---SAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVAVL 868
+ S SK S Y+ S++ Q C + + + Y I + A A +
Sbjct: 347 KTYNESMAQEKSKGTRKKSYYTVSYWEQIRLCTIRGFLRIYGDKSYTVINTCAAIAQAFI 406
Query: 869 LGSMFWDLGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSV 908
GS+F+ S F+ G ++ +++ +M ++
Sbjct: 407 TGSLFYQAPS---STLGAFSRSGVLFFSLLYYSLMGLANI 443
Score = 50.4 bits (119), Expect = 2e-05
Identities = 37/141 (26%), Positives = 76/141 (53%), Gaps = 9/141 (6%)
Query: 18 VLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVG-PTKALFMDEISTGLDSSTTF 76
++ +L ++ ++ +VG ++ Q+K+L+ G LVG P LF+DE ++GLDS + +
Sbjct: 848 IISILEMQEFSEALVGEIGY-GLNVEQRKKLSIGVELVGKPDLLLFLDEPTSGLDSQSAW 906
Query: 77 QIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILL-SDSHIIYQGP----REHVLEF 131
+V +++ + + +++ ++ QP + FD ++LL IY G V+++
Sbjct: 907 AVVKMLKR-LALAGQSILCTIHQPSATLFEQFDRLLLLGKGGQTIYFGEIGKNSSSVIKY 965
Query: 132 FESIGF-KCPNRKGVADFLQE 151
FE G KC + A+++ E
Sbjct: 966 FEKNGARKCQQNENPAEYILE 986
>CDR3_CANAL (O42690) Opaque-specific ABC transporter CDR3
Length = 1501
Score = 160 bits (404), Expect = 2e-38
Identities = 99/296 (33%), Positives = 162/296 (54%), Gaps = 11/296 (3%)
Query: 638 QKVMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSI-IFMDEPTSGLDARA 696
+K+++L+E++ +A+VG+PG GL++EQRKRLT+AVELVA P + +F+DEPTSGLD++
Sbjct: 959 EKIIDLLEMRTYVDAIVGVPGE-GLNVEQRKRLTIAVELVARPKLLVFLDEPTSGLDSQT 1017
Query: 697 AAVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLIN 756
A + + +R + G+ ++CTIHQPS + E FD LLLL Q G+ +Y G G N LI
Sbjct: 1018 AWSICKLIRKLANHGQAILCTIHQPSAILLEEFDRLLLL-QKGETVYFGEFGANCHTLIE 1076
Query: 757 HFEGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNSELYR-INKAL--VKEL 813
+FE G K NPA WML V ++ + D+ E ++NS YR + L ++E+
Sbjct: 1077 YFER-NGASKCPQHANPAEWMLGVIGAAPGTQANQDYFETWRNSPEYRAVQNELHRLEEM 1135
Query: 814 SAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVAVLLGSMF 873
A K+ Y+ SF+ Q + + + YWR P Y +F + ++ G +
Sbjct: 1136 PGLASGEKEPDTNQAYAASFWKQYIFVVHRLFQQYWRTPSYIYSKFAMAVLCSLFNGFTY 1195
Query: 874 WDLGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVERTVFYRESAAGNVF 929
+ + + Q L N M S++S +++ + P+ +R ++ F
Sbjct: 1196 YKSQNSM---QGLKNQMLSIFSMFVVLTTL-AQQYVPLFVTQRDLYEARERPSKTF 1247
Score = 94.0 bits (232), Expect = 2e-18
Identities = 46/141 (32%), Positives = 79/141 (55%)
Query: 14 ITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDSS 73
+ D V+ GL +T VGN IR ISGG++KRL+ E+ + D + GLD++
Sbjct: 268 VVDVVMATYGLSHTKNTKVGNDFIRGISGGERKRLSIAEVTLVQASIQCWDNSTRGLDAA 327
Query: 74 TTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEFFE 133
T + ++S++ IL T +I++ Q Y+LFD +I++ + + I+ G + +F+
Sbjct: 328 TALEFISSLKTSASILNDTPLIAIYQCSQNAYDLFDKVIVMYEGYQIFFGSSQRAAAYFK 387
Query: 134 SIGFKCPNRKGVADFLQEVTS 154
+GF C +R+ DFL +TS
Sbjct: 388 KMGFVCQDRQTTPDFLTSITS 408
Score = 48.9 bits (115), Expect = 7e-05
Identities = 37/138 (26%), Positives = 69/138 (49%), Gaps = 8/138 (5%)
Query: 18 VLRVLGLEICADTVVGNAMIRAISGGQKKRLTTG-EMLVGPTKALFMDEISTGLDSSTTF 76
++ +L + D +VG ++ Q+KRLT E++ P +F+DE ++GLDS T +
Sbjct: 961 IIDLLEMRTYVDAIVG-VPGEGLNVEQRKRLTIAVELVARPKLLVFLDEPTSGLDSQTAW 1019
Query: 77 QIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGP----REHVLEFF 132
I +R+ + + ++ ++ QP FD ++LL +Y G ++E+F
Sbjct: 1020 SICKLIRKLANHGQ-AILCTIHQPSAILLEEFDRLLLLQKGETVYFGEFGANCHTLIEYF 1078
Query: 133 ESIG-FKCPNRKGVADFL 149
E G KCP A+++
Sbjct: 1079 ERNGASKCPQHANPAEWM 1096
Score = 47.4 bits (111), Expect = 2e-04
Identities = 63/264 (23%), Positives = 114/264 (42%), Gaps = 35/264 (13%)
Query: 640 VMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAV 699
VM L +N VG + G+S +RKRL++A + SI D T GLDA A
Sbjct: 272 VMATYGLSHTKNTKVGNDFIRGISGGERKRLSIAEVTLVQASIQCWDNSTRGLDAATALE 331
Query: 700 VMRTVRNTVD-TGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHF 758
+ +++ + T + I+Q S + ++ FD+++++ +G Q I+ G +S +F
Sbjct: 332 FISSLKTSASILNDTPLIAIYQCSQNAYDLFDKVIVMYEGYQ-IFFG----SSQRAAAYF 386
Query: 759 EGIQGV-------------------RKIKDGY------NPATWMLEVTTSSKERELGIDF 793
+ + V R IK GY P + S + + L +
Sbjct: 387 KKMGFVCQDRQTTPDFLTSITSPAERIIKPGYERLVPRTPKEFYRYWRRSPERQALLEEI 446
Query: 794 AELYKNSELYRINKALVKELSAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPE 853
E N E Y + + + +A +K Y S Y+ S Q + + K++W R
Sbjct: 447 DEYLDNCENYDQKQKIFEANNAKK--AKHTYNKSSYTVSLPMQ-VRYIMKRYWDRMRGDI 503
Query: 854 YNAIRFLY-STAVAVLLGSMFWDL 876
+ + + A+A++L S+F++L
Sbjct: 504 IVPLSTVAGNIAMALILSSVFYNL 527
>CDR2_CANAL (P78595) Multidrug resistance protein CDR2
Length = 1499
Score = 157 bits (396), Expect = 2e-37
Identities = 108/330 (32%), Positives = 166/330 (49%), Gaps = 22/330 (6%)
Query: 640 VMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSII-FMDEPTSGLDARAAA 698
V++L+E+ +ALVG+ G GL++EQRKRLT+ VELVA P ++ F+DEPTSGLD++ A
Sbjct: 978 VIDLLEMTDYADALVGVAGE-GLNVEQRKRLTIGVELVAKPKLLLFLDEPTSGLDSQTAW 1036
Query: 699 VVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHF 758
+ + +R D G+ ++CTIHQPS I FD+LL L++GG+ Y G LG N +IN+F
Sbjct: 1037 SICKLMRKLADHGQAILCTIHQPSALIMAEFDKLLFLQKGGRTAYFGELGENCQTMINYF 1096
Query: 759 EGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNSELYRINKALVKELSA--- 815
E G NPA WML+V ++ D+ E+++NS Y+ + + + A
Sbjct: 1097 EK-YGADPCPKEANPAEWMLQVVGAAPGSHAKQDYFEVWRNSSEYQAVREEINRMEAELS 1155
Query: 816 PAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVAVLLGSMFWD 875
P D +Y+ + Q + W+ WR+P Y +YS + V+ S+F
Sbjct: 1156 KLPRDNDPEALLKYAAPLWKQYLLVSWRTIVQDWRSPGY-----IYSKLILVISSSLF-- 1208
Query: 876 LGSKIEKEQDLFNAMGSMYSAVILIGV---MNCNSVQPVVAVERTVFYRESAAGNVFYFS 932
+G K ++ + S AV + V + + P R V+ A F +
Sbjct: 1209 IGFSFFKSKNNLQGLQSQMLAVFMFFVPFTTFIDQMLPYFVKHRAVYEVREAPSRTFSW- 1267
Query: 933 ICLWAGKYPSPICFCTSCG-----LWHYSV 957
AG+ S I F G W+Y V
Sbjct: 1268 FAFIAGQITSEIPFQIVVGTISYFCWYYPV 1297
Score = 96.7 bits (239), Expect = 3e-19
Identities = 46/145 (31%), Positives = 84/145 (57%)
Query: 23 GLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDSSTTFQIVNSM 82
GL +T VGN +R +SGG++KR++ E + D + GLDS+T + + ++
Sbjct: 284 GLSHTRNTNVGNDFVRGVSGGERKRVSIAEASLSGANIQCWDNATRGLDSATALEFIRAL 343
Query: 83 RQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEFFESIGFKCPNR 142
+ IL T +I++ Q + Y LFD++++L + + I+ G E+FE++G+KCP R
Sbjct: 344 KTSATILDTTPLIAIYQCSQDAYELFDNVVVLYEGYQIFFGKASKAKEYFENMGWKCPQR 403
Query: 143 KGVADFLQEVTSRKDQEQYCEHKDR 167
+ ADFL +T+ ++E ++D+
Sbjct: 404 QTTADFLTSLTNPAEREPLPGYEDK 428
Score = 57.0 bits (136), Expect = 3e-07
Identities = 77/315 (24%), Positives = 131/315 (41%), Gaps = 49/315 (15%)
Query: 646 LKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVR 705
L RN VG V G+S +RKR+++A ++ +I D T GLD+ A +R ++
Sbjct: 285 LSHTRNTNVGNDFVRGVSGGERKRVSIAEASLSGANIQCWDNATRGLDSATALEFIRALK 344
Query: 706 NT---VDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHFEGIQ 762
+ +DT T + I+Q S D +E FD +++L +G Q I+ G +S +FE +
Sbjct: 345 TSATILDT--TPLIAIYQCSQDAYELFDNVVVLYEGYQ-IFFG----KASKAKEYFENMG 397
Query: 763 GVRKIKDGYNPATWMLEVTTSSKEREL----------GIDFAELYKNSELY--------- 803
K A ++ +T ++ L +F +KNS Y
Sbjct: 398 W--KCPQRQTTADFLTSLTNPAEREPLPGYEDKVPRTAQEFETFWKNSPEYAELTKEIDE 455
Query: 804 ------RINKALVKELSAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAI 857
R N S S + S Y+ SFF Q + + +P I
Sbjct: 456 YFVECERSNTGETYRESHVGKQSNNTRPSSPYTVSFFMQVRYVIARNFLRMKGDPSIPLI 515
Query: 858 RFLYSTAVAVLLGSMFWDLGSKIEKEQDLFNAM-GSMYSAVI------LIGVMNCNSVQP 910
L + ++L S+F++L K D F G+++ +V+ L+ +++ +P
Sbjct: 516 SILSQLVMGLILASVFFNL----RKSTDTFYFRGGALFFSVLFNAFSSLLEILSLYEARP 571
Query: 911 VVAVERT-VFYRESA 924
+V R YR SA
Sbjct: 572 IVEKHRKYALYRPSA 586
Score = 51.2 bits (121), Expect = 1e-05
Identities = 43/144 (29%), Positives = 72/144 (49%), Gaps = 9/144 (6%)
Query: 16 DYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTG-EMLVGPTKALFMDEISTGLDSST 74
DYV+ +L + AD +VG A ++ Q+KRLT G E++ P LF+DE ++GLDS T
Sbjct: 976 DYVIDLLEMTDYADALVGVAG-EGLNVEQRKRLTIGVELVAKPKLLLFLDEPTSGLDSQT 1034
Query: 75 TFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILL-SDSHIIYQGPR----EHVL 129
+ I MR+ + ++ ++ QP FD ++ L Y G + ++
Sbjct: 1035 AWSICKLMRKLADHGQ-AILCTIHQPSALIMAEFDKLLFLQKGGRTAYFGELGENCQTMI 1093
Query: 130 EFFESIGF-KCPNRKGVADFLQEV 152
+FE G CP A+++ +V
Sbjct: 1094 NYFEKYGADPCPKEANPAEWMLQV 1117
>CDR4_CANAL (O74676) ABC transporter CDR4
Length = 1490
Score = 154 bits (389), Expect = 1e-36
Identities = 103/327 (31%), Positives = 167/327 (50%), Gaps = 16/327 (4%)
Query: 640 VMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSI-IFMDEPTSGLDARAAA 698
++ L+E++ +A+VG+ G GL++EQRKRL++ VELVA P + +F+DEPTSGLD++ A
Sbjct: 967 IIRLLEMEQYADAVVGVSGE-GLNVEQRKRLSIGVELVAKPKLLVFLDEPTSGLDSQTAW 1025
Query: 699 VVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHF 758
+ + +R D G+ ++CTIHQPS + FD LL L++GGQ +Y G LG N + LIN+F
Sbjct: 1026 SICKLIRKLADNGQAILCTIHQPSAILLAEFDRLLFLQRGGQTVYFGDLGKNFTTLINYF 1085
Query: 759 EGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELY-KNSELYRINKA--LVKELSA 815
E G K NPA WMLEV ++ + D+ +++ K+SE +N L+ E
Sbjct: 1086 EK-YGAPKCPPEANPAEWMLEVIGAAPGSKANQDYYDVWLKSSEFQEMNSELDLMSEELV 1144
Query: 816 PAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVAVLLGSMFWD 875
P D Y+ ++ Q + + WR P Y +FL ++ G F+
Sbjct: 1145 KKPLDDDPDRLKPYAAPYWEQYLFVTKRVFEQNWRTPSYLYSKFLLVVTSSLFNGFSFYK 1204
Query: 876 LGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVERTVFYRESAAGNVFYFSICL 935
+ Q L N M S++ ++++ + P +R ++ F + I
Sbjct: 1205 ADRSL---QGLQNQMFSVFMFLVILHTL-IQQYLPTFVSQRDLYEVRERPSKTFSW-ITF 1259
Query: 936 WAGKYPSPICFCTSCG-----LWHYSV 957
A + + I + CG W+Y V
Sbjct: 1260 IAAQVTAEIPWNIICGTLGYFCWYYPV 1286
Score = 92.4 bits (228), Expect = 6e-18
Identities = 43/137 (31%), Positives = 79/137 (57%)
Query: 18 VLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDSSTTFQ 77
V+ V GL +T VGN IR +SGG++KR++ E+ + D + GLDS+T +
Sbjct: 284 VMAVYGLSHTRNTKVGNDFIRGVSGGERKRVSIAEITLNNAMVQCWDNSTRGLDSATALE 343
Query: 78 IVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEFFESIGF 137
+ +++ I+ T ++++ Q + Y+LFD ++L+ + IY G + ++F +G+
Sbjct: 344 FIRALKASADIVHTTPLVAIYQCSQDAYDLFDKVVLMYQGYQIYFGSAKKAKQYFIDMGY 403
Query: 138 KCPNRKGVADFLQEVTS 154
+CP R+ ADFL +T+
Sbjct: 404 ECPQRQTTADFLTSLTN 420
Score = 57.8 bits (138), Expect = 2e-07
Identities = 68/260 (26%), Positives = 112/260 (42%), Gaps = 27/260 (10%)
Query: 640 VMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAV 699
VM + L RN VG + G+S +RKR+++A + N + D T GLD+ A
Sbjct: 284 VMAVYGLSHTRNTKVGNDFIRGVSGGERKRVSIAEITLNNAMVQCWDNSTRGLDSATALE 343
Query: 700 VMRTVRNTVDTGRTV-VCTIHQPSIDIFESFDELLLLKQGGQEIYVG----------PLG 748
+R ++ + D T + I+Q S D ++ FD+++L+ QG Q IY G +G
Sbjct: 344 FIRALKASADIVHTTPLVAIYQCSQDAYDLFDKVVLMYQGYQ-IYFGSAKKAKQYFIDMG 402
Query: 749 HNS----------SNLINHFEGIQGVRKIKDGYNPAT--WMLEVTTSSKERELGIDFAEL 796
+ ++L N E I VR+ +G P T E S E + + +
Sbjct: 403 YECPQRQTTADFLTSLTNPAERI--VRQGFEGKVPQTPQEFYEYWKKSPEGQQIVADVDQ 460
Query: 797 YKNSELYRINKALVKELSAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNA 856
Y K +KE + A S L S Y+ SFF Q + NP +
Sbjct: 461 YLTEHSSAAEKEAIKE-AHQARQSDHLKPASPYTVSFFMQVRYIAHRNILRIKGNPSIHL 519
Query: 857 IRFLYSTAVAVLLGSMFWDL 876
+ + ++ +L S+F++L
Sbjct: 520 FQIFGNIGMSFILSSIFYNL 539
Score = 57.0 bits (136), Expect = 3e-07
Identities = 49/165 (29%), Positives = 84/165 (50%), Gaps = 14/165 (8%)
Query: 9 QKENLITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKAL-FMDEIS 67
+++N DY++R+L +E AD VVG + ++ Q+KRL+ G LV K L F+DE +
Sbjct: 958 KEKNEYVDYIIRLLEMEQYADAVVGVSG-EGLNVEQRKRLSIGVELVAKPKLLVFLDEPT 1016
Query: 68 TGLDSSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLS-DSHIIYQGPR- 125
+GLDS T + I +R+ + ++ ++ QP FD ++ L +Y G
Sbjct: 1017 SGLDSQTAWSICKLIRKLADNGQ-AILCTIHQPSAILLAEFDRLLFLQRGGQTVYFGDLG 1075
Query: 126 ---EHVLEFFESIGF-KCPNRKGVADFLQEVT-----SRKDQEQY 161
++ +FE G KCP A+++ EV S+ +Q+ Y
Sbjct: 1076 KNFTTLINYFEKYGAPKCPPEANPAEWMLEVIGAAPGSKANQDYY 1120
>PDR5_YEAST (P33302) Suppressor of toxicity of sporidesmin
Length = 1511
Score = 154 bits (388), Expect = 2e-36
Identities = 89/271 (32%), Positives = 154/271 (55%), Gaps = 19/271 (7%)
Query: 638 QKVMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSI-IFMDEPTSGLDARA 696
++V++++E++ +A+VG+ G GL++EQRKRLT+ VEL A P + +F+DEPTSGLD++
Sbjct: 987 EEVIKILEMEKYADAVVGVAGE-GLNVEQRKRLTIGVELTAKPKLLVFLDEPTSGLDSQT 1045
Query: 697 AAVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLIN 756
A + + ++ + G+ ++CTIHQPS + + FD LL +++GG+ +Y G LG +I+
Sbjct: 1046 AWSICQLMKKLANHGQAILCTIHQPSAILMQEFDRLLFMQRGGKTVYFGDLGEGCKTMID 1105
Query: 757 HFEGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNSELYR--------INKA 808
+FE G K NPA WMLEV ++ D+ E+++NSE YR + +
Sbjct: 1106 YFES-HGAHKCPADANPAEWMLEVVGAAPGSHANQDYYEVWRNSEEYRAVQSELDWMERE 1164
Query: 809 LVKELSAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVAVL 868
L K+ S A K ++S+S Q + YWR+P+Y +F+ + +
Sbjct: 1165 LPKKGSITAAEDK-----HEFSQSIIYQTKLVSIRLFQQYWRSPDYLWSKFILTIFNQLF 1219
Query: 869 LGSMFWDLGSKIEKEQDLFNAMGSMYSAVIL 899
+G F+ G+ + Q L N M +++ ++
Sbjct: 1220 IGFTFFKAGTSL---QGLQNQMLAVFMFTVI 1247
Score = 99.8 bits (247), Expect = 4e-20
Identities = 45/147 (30%), Positives = 85/147 (57%)
Query: 12 NLITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLD 71
N + + + GL +T VGN ++R +SGG++KR++ E+ + +K D + GLD
Sbjct: 281 NHLAEVAMATYGLSHTRNTKVGNDIVRGVSGGERKRVSIAEVSICGSKFQCWDNATRGLD 340
Query: 72 SSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEF 131
S+T + + +++ I + +++ Q + Y+LF+ + +L D + IY GP + ++
Sbjct: 341 SATALEFIRALKTQADISNTSATVAIYQCSQDAYDLFNKVCVLDDGYQIYYGPADKAKKY 400
Query: 132 FESIGFKCPNRKGVADFLQEVTSRKDQ 158
FE +G+ CP+R+ ADFL VTS ++
Sbjct: 401 FEDMGYVCPSRQTTADFLTSVTSPSER 427
Score = 61.6 bits (148), Expect = 1e-08
Identities = 74/312 (23%), Positives = 122/312 (38%), Gaps = 39/312 (12%)
Query: 646 LKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVR 705
L RN VG V G+S +RKR+++A + D T GLD+ A +R ++
Sbjct: 293 LSHTRNTKVGNDIVRGVSGGERKRVSIAEVSICGSKFQCWDNATRGLDSATALEFIRALK 352
Query: 706 NTVDTGRT-VVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHFEGIQGV 764
D T I+Q S D ++ F+++ +L G Q IY GP + +FE + V
Sbjct: 353 TQADISNTSATVAIYQCSQDAYDLFNKVCVLDDGYQ-IYYGP----ADKAKKYFEDMGYV 407
Query: 765 ----RKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNSELYRINKALVKEL------- 813
+ D T E T + + GI + K Y + KEL
Sbjct: 408 CPSRQTTADFLTSVTSPSERTLNKDMLKKGIHIPQTPKEMNDYWVKSPNYKELMKEVDQR 467
Query: 814 -----SAPAPCSKDLYFPSQ---------YSRSFFTQCMACLWKQHWSYWRNPEYNAIRF 859
A K+ + Q Y+ S+ Q L + W N +
Sbjct: 468 LLNDDEASREAIKEAHIAKQSKRARPSSPYTVSYMMQVKYLLIRNMWRLRNNIGFTLFMI 527
Query: 860 LYSTAVAVLLGSMFWDLGSKIEKEQDLFNAMGSMYSAVI------LIGVMNCNSVQPVVA 913
L + ++A++LGSMF+ + K + F +M+ A++ L+ + + +P+
Sbjct: 528 LGNCSMALILGSMFFKIMKKGDTSTFYFRG-SAMFFAILFNAFSSLLEIFSLYEARPITE 586
Query: 914 VERTV-FYRESA 924
RT Y SA
Sbjct: 587 KHRTYSLYHPSA 598
Score = 57.0 bits (136), Expect = 3e-07
Identities = 47/165 (28%), Positives = 86/165 (51%), Gaps = 14/165 (8%)
Query: 9 QKENLITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTG-EMLVGPTKALFMDEIS 67
+++N + V+++L +E AD VVG A ++ Q+KRLT G E+ P +F+DE +
Sbjct: 980 EEKNRYVEEVIKILEMEKYADAVVGVAG-EGLNVEQRKRLTIGVELTAKPKLLVFLDEPT 1038
Query: 68 TGLDSSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLS-DSHIIYQGPR- 125
+GLDS T + I M++ + + ++ ++ QP FD ++ + +Y G
Sbjct: 1039 SGLDSQTAWSICQLMKKLANHGQ-AILCTIHQPSAILMQEFDRLLFMQRGGKTVYFGDLG 1097
Query: 126 ---EHVLEFFESIG-FKCPNRKGVADFLQEVT-----SRKDQEQY 161
+ ++++FES G KCP A+++ EV S +Q+ Y
Sbjct: 1098 EGCKTMIDYFESHGAHKCPADANPAEWMLEVVGAAPGSHANQDYY 1142
>CDR1_CANAL (P43071) Multidrug resistance protein CDR1
Length = 1501
Score = 154 bits (388), Expect = 2e-36
Identities = 104/325 (32%), Positives = 163/325 (50%), Gaps = 16/325 (4%)
Query: 640 VMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSII-FMDEPTSGLDARAAA 698
V++L+E+ +ALVG+ G GL++EQRKRLT+ VELVA P ++ F+DEPTSGLD++ A
Sbjct: 980 VIDLLEMTDYADALVGVAGE-GLNVEQRKRLTIGVELVAKPKLLLFLDEPTSGLDSQTAW 1038
Query: 699 VVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHF 758
+ + +R D G+ ++CTIHQPS I FD LL L++GG+ Y G LG N +IN+F
Sbjct: 1039 SICKLMRKLADHGQAILCTIHQPSALIMAEFDRLLFLQKGGRTAYFGELGENCQTMINYF 1098
Query: 759 EGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNSELYRINKALVKELSA--- 815
E G NPA WML+V ++ D+ E+++NS Y+ + + + A
Sbjct: 1099 EK-YGADPCPKEANPAEWMLQVVGAAPGSHAKQDYFEVWRNSSEYQAVREEINRMEAELS 1157
Query: 816 PAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVAVLLGSMFWD 875
P D +Y+ + Q + W+ WR+P Y + + A+ G F+
Sbjct: 1158 KLPRDNDPEALLKYAAPLWKQYLLVSWRTIVQDWRSPGYIYSKIFLVVSAALFNGFSFFK 1217
Query: 876 LGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVERTVFYRESAAGNVFYFSICL 935
+ + Q L N M S++ I + + P +R V+ A F +
Sbjct: 1218 AKNNM---QGLQNQMFSVFMFFIPFNTL-VQQMLPYFVKQRDVYEVREAPSRTFSW-FAF 1272
Query: 936 WAGKYPSPICFCTSCG-----LWHY 955
AG+ S I + + G W+Y
Sbjct: 1273 IAGQITSEIPYQVAVGTIAFFCWYY 1297
Score = 95.1 bits (235), Expect = 9e-19
Identities = 46/145 (31%), Positives = 83/145 (56%)
Query: 23 GLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDSSTTFQIVNSM 82
GL +T VGN +R +SGG++KR++ E + D + GLDS+T + + ++
Sbjct: 286 GLSHTRNTNVGNDFVRGVSGGERKRVSIAEASLSGANIQCWDNATRGLDSATALEFIRAL 345
Query: 83 RQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEFFESIGFKCPNR 142
+ IL T +I++ Q + Y+LFD +++L + + I+ G E+FE +G+KCP R
Sbjct: 346 KTSAVILDTTPLIAIYQCSQDAYDLFDKVVVLYEGYQIFFGKATKAKEYFEKMGWKCPQR 405
Query: 143 KGVADFLQEVTSRKDQEQYCEHKDR 167
+ ADFL +T+ ++E ++D+
Sbjct: 406 QTTADFLTSLTNPAEREPLPGYEDK 430
Score = 50.8 bits (120), Expect = 2e-05
Identities = 43/144 (29%), Positives = 72/144 (49%), Gaps = 9/144 (6%)
Query: 16 DYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTG-EMLVGPTKALFMDEISTGLDSST 74
DYV+ +L + AD +VG A ++ Q+KRLT G E++ P LF+DE ++GLDS T
Sbjct: 978 DYVIDLLEMTDYADALVGVAG-EGLNVEQRKRLTIGVELVAKPKLLLFLDEPTSGLDSQT 1036
Query: 75 TFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILL-SDSHIIYQGPR----EHVL 129
+ I MR+ + ++ ++ QP FD ++ L Y G + ++
Sbjct: 1037 AWSICKLMRKLADHGQ-AILCTIHQPSALIMAEFDRLLFLQKGGRTAYFGELGENCQTMI 1095
Query: 130 EFFESIGF-KCPNRKGVADFLQEV 152
+FE G CP A+++ +V
Sbjct: 1096 NYFEKYGADPCPKEANPAEWMLQV 1119
Score = 49.7 bits (117), Expect = 4e-05
Identities = 71/312 (22%), Positives = 128/312 (40%), Gaps = 43/312 (13%)
Query: 646 LKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVR 705
L RN VG V G+S +RKR+++A ++ +I D T GLD+ A +R ++
Sbjct: 287 LSHTRNTNVGNDFVRGVSGGERKRVSIAEASLSGANIQCWDNATRGLDSATALEFIRALK 346
Query: 706 -NTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHFEGIQGV 764
+ V T + I+Q S D ++ FD++++L +G Q I+ G ++ +FE +
Sbjct: 347 TSAVILDTTPLIAIYQCSQDAYDLFDKVVVLYEGYQ-IFFG----KATKAKEYFEKMGW- 400
Query: 765 RKIKDGYNPATWMLEVTTSSKEREL----------GIDFAELYKNSELY----------- 803
K A ++ +T ++ L +F +KNS Y
Sbjct: 401 -KCPQRQTTADFLTSLTNPAEREPLPGYEDKVPRTAQEFETYWKNSPEYAELTKEIDEYF 459
Query: 804 ----RINKALVKELSAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRF 859
R N S A S + S Y+ SFF Q + + +P
Sbjct: 460 VECERSNTRETYRESHVAKQSNNTRPASPYTVSFFMQVRYGVARNFLRMKGDPSIPIFSV 519
Query: 860 LYSTAVAVLLGSMFWDLGSKIEKEQDLFNAMGSMYSAVI------LIGVMNCNSVQPVVA 913
+ ++L S+F++L + + +M+ AV+ L+ +M+ +P+V
Sbjct: 520 FGQLVMGLILSSVFYNLS---QTTGSFYYRGAAMFFAVLFNAFSSLLEIMSLFEARPIVE 576
Query: 914 V-ERTVFYRESA 924
++ YR SA
Sbjct: 577 KHKKYALYRPSA 588
>CGR1_CANGA (O74208) ATP-binding cassette transporter CGR1
(Pleomorphic drug resistance homolog)
Length = 1542
Score = 151 bits (382), Expect = 8e-36
Identities = 89/267 (33%), Positives = 148/267 (55%), Gaps = 11/267 (4%)
Query: 638 QKVMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSI-IFMDEPTSGLDARA 696
+ V++++E++ +A+VG+PG GL++EQRKRLT+ VEL A P + +F+DEPTSGLD++
Sbjct: 1003 EAVIKILEMETYADAVVGVPGE-GLNVEQRKRLTIGVELAAKPKLLVFLDEPTSGLDSQT 1061
Query: 697 AAVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLIN 756
A + ++ + G+ ++CTIHQPS + + FD LL L++GGQ +Y G LG +I
Sbjct: 1062 AWATCQLMKKLANHGQAILCTIHQPSAMLMQEFDRLLFLQKGGQTVYFGDLGKGCKTMIK 1121
Query: 757 HFEGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNSELYRINK----ALVKE 812
+FE G K NPA WMLEV ++ D+ E+++NSE ++ K + KE
Sbjct: 1122 YFED-HGAHKCPPDANPAEWMLEVVGAAPGSHANQDYHEVWRNSEQFKQVKQELEQMEKE 1180
Query: 813 LSAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVAVLLGSM 872
LS D +++ S + Q + YWR P+Y +++ + + +G
Sbjct: 1181 LS-QKELDNDEDANKEFATSLWYQFQLVCVRLFQQYWRTPDYLWSKYILTIFNQLFIGFT 1239
Query: 873 FWDLGSKIEKEQDLFNAMGSMYSAVIL 899
F+ + Q L N M S++ ++
Sbjct: 1240 FFKADHTL---QGLQNQMLSIFMYTVI 1263
Score = 108 bits (270), Expect = 8e-23
Identities = 50/156 (32%), Positives = 89/156 (57%)
Query: 12 NLITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLD 71
N +TD + GL DT VGN ++R +SGG++KR++ E+ + +K D + GLD
Sbjct: 280 NHVTDVAMATYGLSHTRDTKVGNDLVRGVSGGERKRVSIAEVWICGSKFQCWDNATRGLD 339
Query: 72 SSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEF 131
S+T + V +++ HI K +++ Q + YNLF+ + +L + + IY G +H +
Sbjct: 340 SATALEFVRALKTQAHIAKNVATVAIYQCSQDAYNLFNKVSVLYEGYQIYFGDAQHAKVY 399
Query: 132 FESIGFKCPNRKGVADFLQEVTSRKDQEQYCEHKDR 167
F+ +G+ CP R+ + DFL +TS ++ E+ D+
Sbjct: 400 FQKMGYFCPKRQTIPDFLTSITSPAERRINKEYLDK 435
Score = 54.7 bits (130), Expect = 1e-06
Identities = 76/311 (24%), Positives = 122/311 (38%), Gaps = 37/311 (11%)
Query: 646 LKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVR 705
L R+ VG V G+S +RKR+++A + D T GLD+ A +R ++
Sbjct: 292 LSHTRDTKVGNDLVRGVSGGERKRVSIAEVWICGSKFQCWDNATRGLDSATALEFVRALK 351
Query: 706 NTVDTGRTV-VCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHFEGIQGV 764
+ V I+Q S D + F+++ +L +G Q IY G H
Sbjct: 352 TQAHIAKNVATVAIYQCSQDAYNLFNKVSVLYEGYQ-IYFGDAQHAKVYFQKMGYFCPKR 410
Query: 765 RKIKDGYNPATWMLEVTTSSKERELGI-------DFAELYKNSELYRINKALVKELSAP- 816
+ I D T E + + + GI D E + NSE Y+ + + E A
Sbjct: 411 QTIPDFLTSITSPAERRINKEYLDKGIKVPQTPLDMVEYWHNSEEYKQLREEIDETLAHQ 470
Query: 817 -------------APCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYST 863
A SK S Y S+ Q L + W + + ++
Sbjct: 471 SEDDKEEIKEAHIAKQSKRARPSSPYVVSYMMQVKYILIRNFWRIKNSASVTLFQVFGNS 530
Query: 864 AVAVLLGSMFWDLGSKIEK--EQDLFNAMG-SMYSAVI------LIGVMNCNSVQPVVAV 914
A+A +LGSMF+ KI+K D F G +M+ A++ L+ + + +P+
Sbjct: 531 AMAFILGSMFY----KIQKGSSADTFYFRGAAMFFAILFNAFSSLLEIFSLYEARPITEK 586
Query: 915 ERTV-FYRESA 924
RT Y SA
Sbjct: 587 HRTYSLYHPSA 597
Score = 52.8 bits (125), Expect = 5e-06
Identities = 42/151 (27%), Positives = 78/151 (50%), Gaps = 9/151 (5%)
Query: 9 QKENLITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTG-EMLVGPTKALFMDEIS 67
+++N + V+++L +E AD VVG ++ Q+KRLT G E+ P +F+DE +
Sbjct: 996 EEKNEYVEAVIKILEMETYADAVVG-VPGEGLNVEQRKRLTIGVELAAKPKLLVFLDEPT 1054
Query: 68 TGLDSSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILL-SDSHIIYQGPR- 125
+GLDS T + M++ + + ++ ++ QP FD ++ L +Y G
Sbjct: 1055 SGLDSQTAWATCQLMKKLANHGQ-AILCTIHQPSAMLMQEFDRLLFLQKGGQTVYFGDLG 1113
Query: 126 ---EHVLEFFESIG-FKCPNRKGVADFLQEV 152
+ ++++FE G KCP A+++ EV
Sbjct: 1114 KGCKTMIKYFEDHGAHKCPPDANPAEWMLEV 1144
>PDRA_YEAST (P51533) ATP-dependent permease PDR10
Length = 1564
Score = 150 bits (379), Expect = 2e-35
Identities = 87/253 (34%), Positives = 142/253 (55%), Gaps = 8/253 (3%)
Query: 638 QKVMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSI-IFMDEPTSGLDARA 696
++V+E++E+K +A+VG+PG GL++EQRKRLT+ VEL A P + +F+DEPTSGLD++
Sbjct: 1041 EEVIEVLEMKLYADAIVGVPGE-GLNVEQRKRLTIGVELAAKPKLLVFLDEPTSGLDSQT 1099
Query: 697 AAVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLIN 756
A + ++ G+ ++CTIHQPS + + FD LL L++GGQ +Y G LG +IN
Sbjct: 1100 AWSTCQLMKKLASRGQAILCTIHQPSALLMQEFDRLLFLQEGGQTVYFGELGKGCKTMIN 1159
Query: 757 HFEGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNSELYR-INKAL---VKE 812
+FE G K NPA WMLE+ ++ D+ ++++SE YR + K L +E
Sbjct: 1160 YFEA-HGAHKCPPDANPAEWMLEIVGAAPGTHASQDYFAIWRDSEEYREMQKELDWMERE 1218
Query: 813 LSAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVAVLLGSM 872
L S + +++ S Q ++ YWR P Y +F + + +G
Sbjct: 1219 LPKRTEGSSN-EEQKEFATSTLYQIKLVSYRLFHQYWRTPFYLWSKFFSTIVSELFIGFT 1277
Query: 873 FWDLGSKIEKEQD 885
F+ + ++ Q+
Sbjct: 1278 FFKANTSLQGLQN 1290
Score = 101 bits (251), Expect = 1e-20
Identities = 45/145 (31%), Positives = 84/145 (57%)
Query: 14 ITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDSS 73
+T+ + GL ADT VGN +R +SGG++KR++ E+ + +K D + GLDS+
Sbjct: 303 MTEVAMATYGLSHTADTKVGNDFVRGVSGGERKRVSIAEVSICGSKFQCWDNATRGLDSA 362
Query: 74 TTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEFFE 133
T + + +++ I K +++ Q + Y+LFD + +L D + I+ GP + ++F+
Sbjct: 363 TALEFIKALKTQATITKSAATVAIYQCSKDAYDLFDKVCVLYDGYQIFFGPSKQAKKYFQ 422
Query: 134 SIGFKCPNRKGVADFLQEVTSRKDQ 158
+G+ CP R+ AD+L +TS ++
Sbjct: 423 RMGYVCPERQTTADYLTSITSPSER 447
Score = 56.2 bits (134), Expect = 4e-07
Identities = 69/313 (22%), Positives = 126/313 (40%), Gaps = 41/313 (13%)
Query: 654 VGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVRNTVD-TGR 712
VG V G+S +RKR+++A + D T GLD+ A ++ ++ T
Sbjct: 321 VGNDFVRGVSGGERKRVSIAEVSICGSKFQCWDNATRGLDSATALEFIKALKTQATITKS 380
Query: 713 TVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHFEGIQGVRKIKDGYN 772
I+Q S D ++ FD++ +L G Q I+ GP S +F+ + V +
Sbjct: 381 AATVAIYQCSKDAYDLFDKVCVLYDGYQ-IFFGP----SKQAKKYFQRMGYV--CPERQT 433
Query: 773 PATWMLEVTTSS---KEREL----------GIDFAELYKNSELYR-----INKALVKELS 814
A ++ +T+ S K++++ + + + SE Y+ +NK L + S
Sbjct: 434 TADYLTSITSPSERIKDKDMVKHGIMIPQTAYEMNQYWIQSEEYKQLQVQVNKHLDTDSS 493
Query: 815 APAPCSKDLYFPSQ---------YSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAV 865
K+ + Q Y+ SFF Q L + W +P L A+
Sbjct: 494 QQREQIKNAHIAKQSKRARPSSPYTVSFFLQVKYILIRDIWRIKNDPSIQLFTVLSHAAM 553
Query: 866 AVLLGSMFWDLGSKIEKEQDLFNAMGSMYSAVI-----LIGVMNCNSVQPVVAVERTV-F 919
A++LGSMF+++ + ++ + L+ + + +P+ +T
Sbjct: 554 ALILGSMFYEVMLSTTTTTFYYRGAAIFFAILFNAFSSLLEIFSLYETRPITEKHKTYSL 613
Query: 920 YRESAAGNVFYFS 932
YR SA FS
Sbjct: 614 YRPSADAFASTFS 626
Score = 50.8 bits (120), Expect = 2e-05
Identities = 40/152 (26%), Positives = 79/152 (51%), Gaps = 11/152 (7%)
Query: 9 QKENLITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTG-EMLVGPTKALFMDEIS 67
++++ + V+ VL +++ AD +VG ++ Q+KRLT G E+ P +F+DE +
Sbjct: 1034 EEKDKYVEEVIEVLEMKLYADAIVG-VPGEGLNVEQRKRLTIGVELAAKPKLLVFLDEPT 1092
Query: 68 TGLDSSTTFQIVNSMRQYVHILKGTVVI-SLLQPPPETYNLFDDIILLSD-SHIIYQGPR 125
+GLDS T + M++ +G ++ ++ QP FD ++ L + +Y G
Sbjct: 1093 SGLDSQTAWSTCQLMKKLAS--RGQAILCTIHQPSALLMQEFDRLLFLQEGGQTVYFGEL 1150
Query: 126 ----EHVLEFFESIG-FKCPNRKGVADFLQEV 152
+ ++ +FE+ G KCP A+++ E+
Sbjct: 1151 GKGCKTMINYFEAHGAHKCPPDANPAEWMLEI 1182
>PDRF_YEAST (Q04182) ATP-dependent permease PDR15
Length = 1529
Score = 149 bits (376), Expect = 4e-35
Identities = 90/272 (33%), Positives = 152/272 (55%), Gaps = 21/272 (7%)
Query: 638 QKVMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSI-IFMDEPTSGLDARA 696
++V++++E++ +A+VG+ G GL++EQRKRLT+ VEL A P + +F+DEPTSGLD++
Sbjct: 1002 EEVIKILEMQQYSDAVVGVAGE-GLNVEQRKRLTIGVELAARPKLLVFLDEPTSGLDSQT 1060
Query: 697 AAVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLIN 756
A + +R G+ ++CTIHQPS + + FD LL L++GGQ +Y G LG +I+
Sbjct: 1061 AWDTCQLMRKLATHGQAILCTIHQPSAILMQQFDRLLFLQKGGQTVYFGDLGEGCKTMID 1120
Query: 757 HFEGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNSELYRINKALVKELS-- 814
+FE +G K NPA WMLEV ++ D+ E+++NS+ Y KA+ +EL
Sbjct: 1121 YFES-KGAHKCPPDANPAEWMLEVVGAAPGSHATQDYNEVWRNSDEY---KAVQEELDWM 1176
Query: 815 -------APAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVAV 867
+ P +++ ++ S + Q + YWR+P+Y +F+ + V
Sbjct: 1177 EKNLPGRSKEPTAEE---HKPFAASLYYQFKMVTIRLFQQYWRSPDYLWSKFILTIFNQV 1233
Query: 868 LLGSMFWDLGSKIEKEQDLFNAMGSMYSAVIL 899
+G F+ + Q L N M S++ ++
Sbjct: 1234 FIGFTFFKADRSL---QGLQNQMLSIFMYTVI 1262
Score = 100 bits (250), Expect = 2e-20
Identities = 45/143 (31%), Positives = 82/143 (56%)
Query: 12 NLITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLD 71
N +T+ + GL DT VGN ++R +SGG++KR++ E+ + + D + GLD
Sbjct: 291 NHVTEVAMATYGLSHTRDTKVGNDLVRGVSGGERKRVSIAEVAICGARFQCWDNATRGLD 350
Query: 72 SSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEF 131
S+T + + +++ I K +++ Q + Y+LFD + +L D + +Y GP + ++
Sbjct: 351 SATALEFIRALKTQADIGKTAATVAIYQCSQDAYDLFDKVCVLDDGYQLYFGPAKDAKKY 410
Query: 132 FESIGFKCPNRKGVADFLQEVTS 154
F+ +G+ CP R+ ADFL +TS
Sbjct: 411 FQDMGYYCPPRQTTADFLTSITS 433
Score = 58.9 bits (141), Expect = 7e-08
Identities = 75/330 (22%), Positives = 129/330 (38%), Gaps = 33/330 (10%)
Query: 646 LKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVR 705
L R+ VG V G+S +RKR+++A + D T GLD+ A +R ++
Sbjct: 303 LSHTRDTKVGNDLVRGVSGGERKRVSIAEVAICGARFQCWDNATRGLDSATALEFIRALK 362
Query: 706 NTVDTGRT-VVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHFEGIQGV 764
D G+T I+Q S D ++ FD++ +L G Q +Y GP +
Sbjct: 363 TQADIGKTAATVAIYQCSQDAYDLFDKVCVLDDGYQ-LYFGPAKDAKKYFQDMGYYCPPR 421
Query: 765 RKIKDGYNPATWMLEVTTSSKERELGI-------DFAELYKNSELYR-----INKALVKE 812
+ D T E S + E G D AE + SE Y+ I+ L K
Sbjct: 422 QTTADFLTSITSPTERIISKEFIEKGTRVPQTPKDMAEYWLQSESYKNLIKDIDSTLEKN 481
Query: 813 LSAPAPCSKDLYFPSQYSR---------SFFTQCMACLWKQHWSYWRNPEYNAIRFLYST 863
+D + Q R ++ Q L + W ++ + + ++
Sbjct: 482 TDEARNIIRDAHHAKQAKRAPPSSPYVVNYGMQVKYLLIRNFWRMKQSASVTLWQVIGNS 541
Query: 864 AVAVLLGSMFWDLGSKIEKEQDLFNAMGSMYSAVI------LIGVMNCNSVQPVVAVERT 917
+A +LGSMF+ + K + F +M+ A++ L+ + + +P+ RT
Sbjct: 542 VMAFILGSMFYKVMKKNDTSTFYFRG-AAMFFAILFNAFSCLLEIFSLYETRPITEKHRT 600
Query: 918 V-FYRESAAGNVFYFSICLWAGKYPSPICF 946
Y SA + F + K + +CF
Sbjct: 601 YSLYHPSA--DAFASVLSEMPPKLITAVCF 628
Score = 53.9 bits (128), Expect = 2e-06
Identities = 43/151 (28%), Positives = 79/151 (51%), Gaps = 9/151 (5%)
Query: 9 QKENLITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTG-EMLVGPTKALFMDEIS 67
+++N + V+++L ++ +D VVG A ++ Q+KRLT G E+ P +F+DE +
Sbjct: 995 EEKNRYVEEVIKILEMQQYSDAVVGVAG-EGLNVEQRKRLTIGVELAARPKLLVFLDEPT 1053
Query: 68 TGLDSSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILL-SDSHIIYQGPR- 125
+GLDS T + MR+ + ++ ++ QP FD ++ L +Y G
Sbjct: 1054 SGLDSQTAWDTCQLMRK-LATHGQAILCTIHQPSAILMQQFDRLLFLQKGGQTVYFGDLG 1112
Query: 126 ---EHVLEFFESIG-FKCPNRKGVADFLQEV 152
+ ++++FES G KCP A+++ EV
Sbjct: 1113 EGCKTMIDYFESKGAHKCPPDANPAEWMLEV 1143
>WHIT_DROME (P10090) White protein
Length = 687
Score = 117 bits (293), Expect = 2e-25
Identities = 89/306 (29%), Positives = 157/306 (51%), Gaps = 23/306 (7%)
Query: 630 VPRHQC*NQKVMEL------VELKPLRNALVGLPG-VCGLSMEQRKRLTVAVELVANPSI 682
+PRH Q+V + + L ++ ++G+PG V GLS +RKRL A E + +P +
Sbjct: 202 MPRHLTYRQRVARVDQVIQELSLSKCQHTIIGVPGRVKGLSGGERKRLAFASEALTDPPL 261
Query: 683 IFMDEPTSGLDARAAAVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEI 742
+ DEPTSGLD+ A V++ ++ G+TV+ TIHQPS ++FE FD++LL+ + G+
Sbjct: 262 LICDEPTSGLDSFTAHSVVQVLKKLSQKGKTVILTIHQPSSELFELFDKILLMAE-GRVA 320
Query: 743 YVGPLGHNSSNLINHFEGIQGVRKIKDGYNPATWMLEV--TTSSKERELGIDFAELYKNS 800
++G S ++ F + + YNPA + ++V +E E A++ N
Sbjct: 321 FLG----TPSEAVDFFSYVGA--QCPTNYNPADFYVQVLAVVPGREIESRDRIAKICDNF 374
Query: 801 ELYRINKALVKELSAPAPCSKDLYFPSQ---YSRSFFTQCMACLWKQHWSYWRNPEYNAI 857
+ ++ + + ++L A K L P Y ++F Q A LW+ S + P +
Sbjct: 375 AISKVARDM-EQLLATKNLEKPLEQPENGYTYKATWFMQFRAVLWRSWLSVLKEPLLVKV 433
Query: 858 RFLYSTAVAVLLGSMFWDLGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVERT 917
R + +T VA+L+G +F LG ++ + + N G+++ + + N + V E
Sbjct: 434 RLIQTTMVAILIGLIF--LGQQL-TQVGVMNINGAIFLFLTNMTFQNVFATINVFTSELP 490
Query: 918 VFYRES 923
VF RE+
Sbjct: 491 VFMREA 496
Score = 82.4 bits (202), Expect = 6e-15
Identities = 43/138 (31%), Positives = 75/138 (54%), Gaps = 2/138 (1%)
Query: 16 DYVLRVLGLEICADTVVG-NAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDSST 74
D V++ L L C T++G ++ +SGG++KRL + L DE ++GLDS T
Sbjct: 216 DQVIQELSLSKCQHTIIGVPGRVKGLSGGERKRLAFASEALTDPPLLICDEPTSGLDSFT 275
Query: 75 TFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEFFES 134
+V +++ K TV++++ QP E + LFD I+L+++ + + G ++FF
Sbjct: 276 AHSVVQVLKKLSQKGK-TVILTIHQPSSELFELFDKILLMAEGRVAFLGTPSEAVDFFSY 334
Query: 135 IGFKCPNRKGVADFLQEV 152
+G +CP ADF +V
Sbjct: 335 VGAQCPTNYNPADFYVQV 352
>WHIT_CERCA (Q17320) White protein
Length = 679
Score = 117 bits (293), Expect = 2e-25
Identities = 92/306 (30%), Positives = 154/306 (50%), Gaps = 22/306 (7%)
Query: 630 VPRHQC*NQKVMEL------VELKPLRNALVGLPG-VCGLSMEQRKRLTVAVELVANPSI 682
+PRH QKV + + L +N L+G+PG V GLS +RKRL A E + +P +
Sbjct: 193 MPRHMTQKQKVQRVDQVIQDLSLGKCQNTLIGVPGRVKGLSGGERKRLAFASEALTDPPL 252
Query: 683 IFMDEPTSGLDARAAAVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEI 742
+ DEPTSGLD+ A V++ ++ G+TV+ TIHQPS ++FE FD++LL+ + G+
Sbjct: 253 LICDEPTSGLDSFMAHSVVQVLKKLSQKGKTVILTIHQPSSELFELFDKILLMAE-GRVA 311
Query: 743 YVGPLGHNSSNLINHFEGIQGVRKIKDGYNPATWMLEV--TTSSKERELGIDFAELYKNS 800
++G G ++ F I Y PA + ++V +E E A++ N
Sbjct: 312 FLGTPG----EAVDFFSYIGAT--CPTNYTPADFYVQVLAVVPGREVESRDRVAKICDNF 365
Query: 801 ELYRINKAL---VKELSAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAI 857
+ ++++ + ++L K+ Y S+F Q A LW+ S + P +
Sbjct: 366 AVGKVSREMEQNFQKLVKSNGFGKEDENEYTYKASWFMQFRAVLWRSWLSVLKEPLLVKV 425
Query: 858 RFLYSTAVAVLLGSMFWDLGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVERT 917
R L +T VAVL+G +F LG ++ + + N G+++ + + N + V E
Sbjct: 426 RLLQTTMVAVLIGLIF--LGQQL-TQVGVMNINGAIFLFLTNMTFQNSFATITVFTTELP 482
Query: 918 VFYRES 923
VF RE+
Sbjct: 483 VFMRET 488
Score = 77.8 bits (190), Expect = 1e-13
Identities = 46/148 (31%), Positives = 79/148 (53%), Gaps = 3/148 (2%)
Query: 6 TEGQKENLITDYVLRVLGLEICADTVVG-NAMIRAISGGQKKRLTTGEMLVGPTKALFMD 64
T+ QK + D V++ L L C +T++G ++ +SGG++KRL + L D
Sbjct: 198 TQKQKVQRV-DQVIQDLSLGKCQNTLIGVPGRVKGLSGGERKRLAFASEALTDPPLLICD 256
Query: 65 EISTGLDSSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGP 124
E ++GLDS +V +++ K TV++++ QP E + LFD I+L+++ + + G
Sbjct: 257 EPTSGLDSFMAHSVVQVLKKLSQKGK-TVILTIHQPSSELFELFDKILLMAEGRVAFLGT 315
Query: 125 REHVLEFFESIGFKCPNRKGVADFLQEV 152
++FF IG CP ADF +V
Sbjct: 316 PGEAVDFFSYIGATCPTNYTPADFYVQV 343
>WHIT_LUCCU (Q05360) White protein
Length = 677
Score = 115 bits (288), Expect = 6e-25
Identities = 85/292 (29%), Positives = 149/292 (50%), Gaps = 17/292 (5%)
Query: 639 KVMELVELKPLRNALVGLPG-VCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAA 697
+V++ + L +N ++G+PG V GLS +RKRL A E + +P ++ DEPTSGLD+ A
Sbjct: 206 QVIQDLSLIKCQNTIIGVPGRVKGLSGGERKRLAFASEALTDPPLLICDEPTSGLDSFMA 265
Query: 698 AVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINH 757
A V++ ++ G+TV+ TIHQPS ++FE FD++LL+ +G P+ ++
Sbjct: 266 ASVVQVLKKLSQRGKTVILTIHQPSSELFELFDKILLMAEGRVAFLGTPV-----EAVDF 320
Query: 758 FEGIQGVRKIKDGYNPATWMLEV--TTSSKERELGIDFAELYKNSELYRINKALVKELSA 815
F I + YNPA + ++V +E E +++ N + ++++ + +
Sbjct: 321 FSFIGA--QCPTNYNPADFYVQVLAVVPGREIESRDRISKICDNFAVGKVSREMEQNFQK 378
Query: 816 PAP----CSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVAVLLGS 871
A KD Y S+FTQ A +W+ S + P +R + +T VAVL+G
Sbjct: 379 IAAKTDGLQKDDETTILYKASWFTQFRAIMWRSWISTLKEPLLVKVRLIQTTMVAVLIGL 438
Query: 872 MFWDLGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVERTVFYRES 923
+F ++ + + N G+++ + + N +V V E VF RE+
Sbjct: 439 IFL---NQPMTQVGVMNINGAIFLFLTNMTFQNVFAVINVFTSELPVFMRET 487
Score = 79.7 bits (195), Expect = 4e-14
Identities = 47/153 (30%), Positives = 81/153 (52%), Gaps = 3/153 (1%)
Query: 1 MKAVATEGQKENLITDYVLRVLGLEICADTVVG-NAMIRAISGGQKKRLTTGEMLVGPTK 59
M T+ QK + D V++ L L C +T++G ++ +SGG++KRL +
Sbjct: 191 MPRTMTQKQKLQRV-DQVIQDLSLIKCQNTIIGVPGRVKGLSGGERKRLAFASEALTDPP 249
Query: 60 ALFMDEISTGLDSSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHI 119
L DE ++GLDS +V +++ K TV++++ QP E + LFD I+L+++ +
Sbjct: 250 LLICDEPTSGLDSFMAASVVQVLKKLSQRGK-TVILTIHQPSSELFELFDKILLMAEGRV 308
Query: 120 IYQGPREHVLEFFESIGFKCPNRKGVADFLQEV 152
+ G ++FF IG +CP ADF +V
Sbjct: 309 AFLGTPVEAVDFFSFIGAQCPTNYNPADFYVQV 341
>ABG4_HUMAN (Q9H172) ATP-binding cassette, sub-family G, member 4
Length = 646
Score = 114 bits (286), Expect = 1e-24
Identities = 87/301 (28%), Positives = 145/301 (47%), Gaps = 17/301 (5%)
Query: 662 LSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVRNTVDTGRTVVCTIHQP 721
LS QRKRL +A+ELV NP ++F DEPTSGLD+ + V+ +++ GRT++CTIHQP
Sbjct: 201 LSGGQRKRLAIALELVNNPPVMFFDEPTSGLDSASCFQVVSLMKSLAQGGRTIICTIHQP 260
Query: 722 SIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHFEGIQGVRKIKDGYNPATWMLEVT 781
S +FE FD+L +L Q GQ I+ G + +NLI + +G+ G+ +NPA +++EV
Sbjct: 261 SAKLFEMFDKLYILSQ-GQCIFKGVV----TNLIPYLKGL-GLH-CPTYHNPADFIIEVA 313
Query: 782 TSSKERELGIDFAELYKNSELYRINKALVKELSAPAPC-----SKDLYFPSQYSRSFFTQ 836
+ + F + K+ ++ PAPC D ++ S TQ
Sbjct: 314 SGEYGDLNPMLFRAVQNGLCAMAEKKSSPEKNEVPAPCPPCPPEVDPIESHTFATSTLTQ 373
Query: 837 CMACLWKQHWSYWRNPEYNAIRFLYSTAVAVLLGSMFWDLGSKIEKEQDLFNAMGSMYSA 896
+ S R+ +RF+ + VL+G ++ +G K +FN G ++ +
Sbjct: 374 FCILFKRTFLSILRDTVLTHLRFMSHVVIGVLIGLLYLHIGDDASK---VFNNTGCLFFS 430
Query: 897 VILIGVMNCNSVQPVVAVERTVFYRESAAGNVFYFSICLWAGKYPSPICFCTSCGLWHYS 956
++ + +E VF RE N +Y + K + + F C + + S
Sbjct: 431 MLFLMFAALMPTVLTFPLEMAVFMREHL--NYWYSLKAYYLAKTMADVPFQVVCPVVYCS 488
Query: 957 V 957
+
Sbjct: 489 I 489
Score = 84.0 bits (206), Expect = 2e-15
Identities = 47/145 (32%), Positives = 81/145 (55%), Gaps = 7/145 (4%)
Query: 10 KENLITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTG 69
K+ L+T+ +L LGL C+ T +SGGQ+KRL LV +F DE ++G
Sbjct: 177 KKELVTE-ILTALGLMSCSHTRTA-----LLSGGQRKRLAIALELVNNPPVMFFDEPTSG 230
Query: 70 LDSSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVL 129
LDS++ FQ+V+ M+ + T++ ++ QP + + +FD + +LS I++G +++
Sbjct: 231 LDSASCFQVVSLMKSLAQGGR-TIICTIHQPSAKLFEMFDKLYILSQGQCIFKGVVTNLI 289
Query: 130 EFFESIGFKCPNRKGVADFLQEVTS 154
+ + +G CP ADF+ EV S
Sbjct: 290 PYLKGLGLHCPTYHNPADFIIEVAS 314
>WHIT_ANOGA (Q27256) White protein
Length = 695
Score = 107 bits (267), Expect = 2e-22
Identities = 80/296 (27%), Positives = 146/296 (49%), Gaps = 21/296 (7%)
Query: 638 QKVMELVELKPLRNALVGLPG-VCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARA 696
Q+V++ + L + ++G PG + GLS +RKRL A E + +P ++ DEPTSGLD+
Sbjct: 219 QEVLQELSLVKCADTIIGAPGRIKGLSGGERKRLAFASETLTDPHLLLCDEPTSGLDSFM 278
Query: 697 AAVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLIN 756
A V++ ++ G+T++ TIHQPS +++ FD++LL+ +G P + S+ +
Sbjct: 279 AHSVLQVLKGMAMKGKTIILTIHQPSSELYCLFDKILLVAEGRVAFLGSP--YQSAEFFS 336
Query: 757 HFEGIQGVRKIKDGYNPATWMLEV--TTSSKERELGIDFAELYKNSELYRINKALVKELS 814
GI YNPA + +++ +KE E ++ + + I + +++ S
Sbjct: 337 QL-GI----PCPPNYNPADFYVQMLAIAPAKEAECRDMIKKICDSFAVSPIAREVLETAS 391
Query: 815 APAPCSKDLYFPSQ--------YSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVA 866
+ Y Q Y S++TQ LW+ S ++P +R L + VA
Sbjct: 392 VAGKGMDEPYMLQQVEGVGSTGYRSSWWTQFYCILWRSWLSVLKDPMLVKVRLLQTAMVA 451
Query: 867 VLLGSMFWDLGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVERTVFYRE 922
L+GS+++ ++ + + N GS++ + + N +V V + E VF RE
Sbjct: 452 TLIGSIYF---GQVLDQDGVMNINGSLFLFLTNMTFQNVFAVINVFSAELPVFLRE 504
Score = 79.3 bits (194), Expect = 5e-14
Identities = 45/133 (33%), Positives = 72/133 (53%), Gaps = 4/133 (3%)
Query: 18 VLRVLGLEICADTVVGN-AMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDSSTTF 76
VL+ L L CADT++G I+ +SGG++KRL + L DE ++GLDS
Sbjct: 221 VLQELSLVKCADTIIGAPGRIKGLSGGERKRLAFASETLTDPHLLLCDEPTSGLDSFMAH 280
Query: 77 QIVNSMRQYVHILKG-TVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEFFESI 135
++ ++ +KG T+++++ QP E Y LFD I+L+++ + + G EFF +
Sbjct: 281 SVLQVLKGMA--MKGKTIILTIHQPSSELYCLFDKILLVAEGRVAFLGSPYQSAEFFSQL 338
Query: 136 GFKCPNRKGVADF 148
G CP ADF
Sbjct: 339 GIPCPPNYNPADF 351
>AUS1_YEAST (Q08409) ATP-dependent permease AUS1
Length = 1394
Score = 105 bits (262), Expect = 6e-22
Identities = 77/294 (26%), Positives = 139/294 (47%), Gaps = 26/294 (8%)
Query: 659 VCGLSMEQRKRLTVAVELVANPSII-FMDEPTSGLDARAAAVVMRTVRNTVDTGRTVVCT 717
V L+ QRK L++ VELV PS++ F+DEPTSGLDA AA +++ ++ G+ + CT
Sbjct: 874 VADLNPTQRKLLSIGVELVTKPSLLLFLDEPTSGLDAEAALTIVKFLKQLSLQGQAIFCT 933
Query: 718 IHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHFEGIQGVRKIKDGYNPATWM 777
IHQPS + FD + LLK+GG+ ++ GP+ ++H + K+ NPA ++
Sbjct: 934 IHQPSKSVISHFDNIFLLKRGGECVFFGPMDDACGYFMSHDNTLV---YDKEHDNPADFV 990
Query: 778 LEVTTSSKE-------------RELGIDFAELYKNSELYRINKALVKELSAPAPCSKDLY 824
++ +S + ID++ L+++S ++ K L A S Y
Sbjct: 991 IDAVGNSNSSAGKDTAEEALTLNKEAIDWSALWESSVEKKLVKKETARLEDDARASGVDY 1050
Query: 825 FPSQYSRSFFTQCMACLW-KQHWSYWRNPEYNAIRFLYSTAVAVLLGSMFWDLGSKIEKE 883
S + + + Q +A + +Q+ R+ Y ++ + + +G FW + I
Sbjct: 1051 TTSLWKQPSYLQQLALITRRQYICTKRDMTYVMAKYCLNGGAGLFIGFSFWHIKHNIIGL 1110
Query: 884 QD--LFNAMGSMYSAVILIGVMNCNSVQPVVAVERTVFYRESAAGNVFYFSICL 935
QD F M S+ ++ N +Q + V+ A N +++++ L
Sbjct: 1111 QDSIFFCFMALCVSSPLI------NQIQDKALKTKEVYVAREARSNTYHWTVLL 1158
Score = 68.9 bits (167), Expect = 7e-11
Identities = 37/147 (25%), Positives = 76/147 (51%), Gaps = 2/147 (1%)
Query: 7 EGQKENLITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEI 66
+G+++ I + +LR GL T+VGN R +SGG++KR++ E + D
Sbjct: 140 KGERDQ-IRNELLREFGLSHVLKTIVGNDFFRGVSGGERKRISIIETFIANGSVYLWDNS 198
Query: 67 STGLDSSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPRE 126
+ GLDS+T + +R+ + ++ + Q + + FD I++LSDS+ ++ G +
Sbjct: 199 TKGLDSATALDFLEILRKMAKATRSVNLVRISQASDKIVDKFDKILMLSDSYQLFYGTVD 258
Query: 127 HVLEFF-ESIGFKCPNRKGVADFLQEV 152
L +F +++G + + ++L +
Sbjct: 259 ECLTYFRDTLGIEKDPNDCIIEYLTSI 285
Score = 48.5 bits (114), Expect = 9e-05
Identities = 42/157 (26%), Positives = 68/157 (42%), Gaps = 12/157 (7%)
Query: 628 PVVPRHQC*NQKVMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDE 687
P R Q N+ + E L + +VG G+S +RKR+++ +AN S+ D
Sbjct: 139 PKGERDQIRNELLREF-GLSHVLKTIVGNDFFRGVSGGERKRISIIETFIANGSVYLWDN 197
Query: 688 PTSGLDARAAAVVMRTVRNTVDTGRTV-VCTIHQPSIDIFESFDELLLLKQGGQEIYVGP 746
T GLD+ A + +R R+V + I Q S I + FD++L+L Q Y
Sbjct: 198 STKGLDSATALDFLEILRKMAKATRSVNLVRISQASDKIVDKFDKILMLSDSYQLFY--- 254
Query: 747 LGHNSSNLINHFEGIQGVRKIKDGYNPATWMLEVTTS 783
+ +F G+ K +P ++E TS
Sbjct: 255 --GTVDECLTYFRDTLGIEK-----DPNDCIIEYLTS 284
Score = 45.4 bits (106), Expect = 8e-04
Identities = 36/133 (27%), Positives = 65/133 (48%), Gaps = 9/133 (6%)
Query: 10 KENLITDYVLRVLGLEICADTVVG------NAMIRAISGGQKKRLTTG-EMLVGPTKALF 62
KE+L +LR G DTV + ++ ++ Q+K L+ G E++ P+ LF
Sbjct: 841 KESLEISCLLRGDGDRAYLDTVSNLLKLPSDILVADLNPTQRKLLSIGVELVTKPSLLLF 900
Query: 63 MDEISTGLDSSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLS-DSHIIY 121
+DE ++GLD+ IV ++Q + + + ++ QP + FD+I LL ++
Sbjct: 901 LDEPTSGLDAEAALTIVKFLKQ-LSLQGQAIFCTIHQPSKSVISHFDNIFLLKRGGECVF 959
Query: 122 QGPREHVLEFFES 134
GP + +F S
Sbjct: 960 FGPMDDACGYFMS 972
>PDRB_YEAST (P40550) ATP-dependent permease PDR11
Length = 1410
Score = 103 bits (257), Expect = 2e-21
Identities = 85/310 (27%), Positives = 139/310 (44%), Gaps = 44/310 (14%)
Query: 659 VCGLSMEQRKRLTVAVELVANPSII-FMDEPTSGLDARAAAVVMRTVRNTVDTGRTVVCT 717
V LS QRK L++ VELV PS++ F+DEPTSGLDA AA +++ ++ G+ ++CT
Sbjct: 873 VADLSPTQRKLLSIGVELVTKPSLLLFLDEPTSGLDAEAALTIVQFLKKLSMQGQAILCT 932
Query: 718 IHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHFEGIQGVRKIKDGYNPATWM 777
IHQPS + FD + LLK+GG+ +Y G L + + H + R++ NPA ++
Sbjct: 933 IHQPSKSVISYFDNIYLLKRGGECVYFGSLPNACDYFVAHDRRLTFDREMD---NPADFV 989
Query: 778 LEVT---------------TSSK--------ERELGIDFAELYKNS-ELYRINKALVKEL 813
++V TSSK ++ I++AEL+++S E R+ L+
Sbjct: 990 IDVVGSGSTNIPMDDAEKPTSSKIDEPVSYHKQSDSINWAELWQSSPEKVRVADDLLLLE 1049
Query: 814 SAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVAVLLGSMF 873
D S+ Q +Q+ R+ Y ++ + + +G F
Sbjct: 1050 EEARKSGVDFTTSVWSPPSYMEQIKLITKRQYICTKRDMTYVFAKYALNAGAGLFIGFSF 1109
Query: 874 WDLGSKIEKEQDLFNAMGSMYSAVILIGVMNC------NSVQPVVAVERTVFYRESAAGN 927
W I QD A+ L +M C N VQ + V+ A N
Sbjct: 1110 WRTKHNINGLQD----------AIFLCFMMLCVSSPLINQVQDKALQSKEVYIAREARSN 1159
Query: 928 VFYFSICLWA 937
+++++ L A
Sbjct: 1160 TYHWTVLLIA 1169
Score = 63.9 bits (154), Expect = 2e-09
Identities = 36/145 (24%), Positives = 70/145 (47%), Gaps = 1/145 (0%)
Query: 16 DYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDSSTT 75
D +L+ GL T VGN +R +SGG++KR++ E + D + GLDS+T
Sbjct: 147 DELLKEFGLSHVKKTYVGNDYVRGVSGGERKRISIIETFIANGSVYLWDNSTKGLDSATA 206
Query: 76 FQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVL-EFFES 134
+ ++ ++ + + + Q + + FD I++L DS ++ G E L F ++
Sbjct: 207 LEFLSITQKMAKATRSVNFVKISQASDKIVSKFDKILMLGDSFQVFYGTMEECLTHFHDT 266
Query: 135 IGFKCPNRKGVADFLQEVTSRKDQE 159
+ K + ++L + + K +E
Sbjct: 267 LQIKKNPNDCIIEYLTSILNFKFKE 291
Score = 48.5 bits (114), Expect = 9e-05
Identities = 38/139 (27%), Positives = 59/139 (42%), Gaps = 11/139 (7%)
Query: 646 LKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVR 705
L ++ VG V G+S +RKR+++ +AN S+ D T GLD+ A + +
Sbjct: 155 LSHVKKTYVGNDYVRGVSGGERKRISIIETFIANGSVYLWDNSTKGLDSATALEFLSITQ 214
Query: 706 NTVDTGRTV-VCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHFEGIQGV 764
R+V I Q S I FD++L+L Q Y + HF +
Sbjct: 215 KMAKATRSVNFVKISQASDKIVSKFDKILMLGDSFQVFY-----GTMEECLTHFHDTLQI 269
Query: 765 RKIKDGYNPATWMLEVTTS 783
+K NP ++E TS
Sbjct: 270 KK-----NPNDCIIEYLTS 283
Score = 42.0 bits (97), Expect = 0.009
Identities = 26/99 (26%), Positives = 53/99 (53%), Gaps = 3/99 (3%)
Query: 36 MIRAISGGQKKRLTTG-EMLVGPTKALFMDEISTGLDSSTTFQIVNSMRQYVHILKGTVV 94
++ +S Q+K L+ G E++ P+ LF+DE ++GLD+ IV +++ + + ++
Sbjct: 872 LVADLSPTQRKLLSIGVELVTKPSLLLFLDEPTSGLDAEAALTIVQFLKK-LSMQGQAIL 930
Query: 95 ISLLQPPPETYNLFDDIILLS-DSHIIYQGPREHVLEFF 132
++ QP + FD+I LL +Y G + ++F
Sbjct: 931 CTIHQPSKSVISYFDNIYLLKRGGECVYFGSLPNACDYF 969
>ABG8_HUMAN (Q9H221) ATP-binding cassette, sub-family G, member 8
(Sterolin-2)
Length = 673
Score = 102 bits (254), Expect = 5e-21
Identities = 57/159 (35%), Positives = 97/159 (60%), Gaps = 8/159 (5%)
Query: 6 TEGQKENLITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDE 65
++ Q++ + D V+ L L CADT VGN +R +SGG+++R++ G L+ L +DE
Sbjct: 180 SQAQRDKRVED-VIAELRLRQCADTRVGNMYVRGLSGGERRRVSIGVQLLWNPGILILDE 238
Query: 66 ISTGLDSSTTFQIVNSMRQYVHILKGT--VVISLLQPPPETYNLFDDIILLSDSHIIYQG 123
++GLDS T +V ++ + + KG V+ISL QP + + LFD ++L++ IY G
Sbjct: 239 PTSGLDSFTAHNLVKTLSR---LAKGNRLVLISLHQPRSDIFRLFDLVLLMTSGTPIYLG 295
Query: 124 PREHVLEFFESIGFKCPNRKGVADFLQEVTS--RKDQEQ 160
+H++++F +IG+ CP ADF ++TS R+ +EQ
Sbjct: 296 AAQHMVQYFTAIGYPCPRYSNPADFYVDLTSIDRRSREQ 334
Score = 81.3 bits (199), Expect = 1e-14
Identities = 55/161 (34%), Positives = 89/161 (55%), Gaps = 22/161 (13%)
Query: 638 QKVMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAA 697
+ V+ + L+ + VG V GLS +R+R+++ V+L+ NP I+ +DEPTSGLD+ A
Sbjct: 189 EDVIAELRLRQCADTRVGNMYVRGLSGGERRRVSIGVQLLWNPGILILDEPTSGLDSFTA 248
Query: 698 AVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINH 757
+++T+ R V+ ++HQP DIF FD L+LL G IY+G H ++ +
Sbjct: 249 HNLVKTLSRLAKGNRLVLISLHQPRSDIFRLFD-LVLLMTSGTPIYLGAAQH----MVQY 303
Query: 758 FEGIQGVRKIKDGY------NPATWMLEVTT---SSKEREL 789
F I GY NPA + +++T+ S+E+EL
Sbjct: 304 FTAI--------GYPCPRYSNPADFYVDLTSIDRRSREQEL 336
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.341 0.149 0.502
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 108,675,347
Number of Sequences: 164201
Number of extensions: 4220417
Number of successful extensions: 15012
Number of sequences better than 10.0: 786
Number of HSP's better than 10.0 without gapping: 595
Number of HSP's successfully gapped in prelim test: 191
Number of HSP's that attempted gapping in prelim test: 13316
Number of HSP's gapped (non-prelim): 1794
length of query: 1055
length of database: 59,974,054
effective HSP length: 121
effective length of query: 934
effective length of database: 40,105,733
effective search space: 37458754622
effective search space used: 37458754622
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 71 (32.0 bits)
Medicago: description of AC147517.6


