

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147484.3 - phase: 0 /pseudo
(526 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
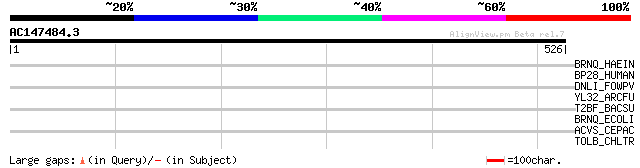
Score E
Sequences producing significant alignments: (bits) Value
BRNQ_HAEIN (P71345) Branched-chain amino acid transport system c... 35 0.62
BP28_HUMAN (Q9H583) Protein BAP28 33 1.4
DNLI_FOWPV (Q67480) DNA ligase (EC 6.5.1.1) (Polydeoxyribonucleo... 32 5.3
YL32_ARCFU (O28148) Hypothetical protein AF2132 31 6.9
T2BF_BACSU (P25217) Type II restriction enzyme BsuFI (EC 3.1.21.... 31 6.9
BRNQ_ECOLI (P37011) Branched-chain amino acid transport system I... 31 6.9
ACVS_CEPAC (P25464) N-(5-amino-5-carboxypentanoyl)-L-cysteinyl-D... 31 6.9
TOLB_CHLTR (O84604) TolB protein precursor 31 9.0
>BRNQ_HAEIN (P71345) Branched-chain amino acid transport system
carrier protein (Branched-chain amino acid uptake
carrier)
Length = 436
Score = 34.7 bits (78), Expect = 0.62
Identities = 17/72 (23%), Positives = 32/72 (43%)
Query: 336 LFEILRLSVPFHPVQYETLKLIYECISECPGAVSTSQMEELVLVLIKMLTKHSDGEMGMI 395
L + LR +P Y + L+ C S C + + E + L+K S+G ++
Sbjct: 356 LLQFLRKKLPSIKFTYNSTLLVTVCFSLCDSLNNVKMLPESINSLLKHFPLSSEGMAWLV 415
Query: 396 PETFIMACSVFV 407
P ++ S+F+
Sbjct: 416 PTLVMLVASIFI 427
>BP28_HUMAN (Q9H583) Protein BAP28
Length = 2144
Score = 33.5 bits (75), Expect = 1.4
Identities = 74/330 (22%), Positives = 132/330 (39%), Gaps = 42/330 (12%)
Query: 72 LHSIVTFHPHFLVSPLVAALSSFDDEPIAEQLVHLILNLSASSDPS-------VCR---- 120
L ++ F+ +VS LVAA D IA+ ++ L +S +C+
Sbjct: 213 LRVLLAFYASTIVSALVAA-EDVSDNIIAKLFPYIQKGLKSSLPDYRAATYMIICQISVK 271
Query: 121 -----EFVARVSDRVSSGALGWSSRQLHSLHCLGVLLNCEKEDDL----HVHIKDVYSLI 171
FV ++ ++ S L CL VLL +K + L H+ +V LI
Sbjct: 272 VTMENTFVNSLASQIIKTLTKIPSLIKDGLSCLIVLLQRQKPESLGKKPFPHLCNVPDLI 331
Query: 172 SVLVTGLQLPSEEIRGEVLFVLYKLFAL---HSTSDEGDGSDMLIPFCPKLLYLLGDVLL 228
++L G+ + ++ + ++L L H T +E +G D I + L +L + L
Sbjct: 332 TIL-HGIS-ETYDVSPLLRYMLPHLVVSIIHHVTGEETEGMDGQI-YKRHLEAILTKISL 388
Query: 229 KTQNDDVRLNCIALLTMLAQRQLLREEPAYDTGNMSLSGEVNFKETEDVPEGASLVNLFA 288
K D LL L L E +Y + S +V+ + +P L + +
Sbjct: 389 KNNLDH-------LLASL----LFEEYISYSSQEEMDSNKVSLLNEQFLPLIRLLESKYP 437
Query: 289 EAIKGPLLSSVSQVQIGALDLLFHYLSSVGTSGNQIQVLVEENIADYLFEILRLSVPFHP 348
+ L + ++ LFH S+ TSG + Q L + + + +L L+ P P
Sbjct: 438 RTLDVVLEEHLKEIADLKKQELFHQFVSLSTSGGKYQFLADSDTS----LMLSLNHPLAP 493
Query: 349 VQYETLKLIYECISECPGAVSTSQMEELVL 378
V+ + + + + V S ++E VL
Sbjct: 494 VRILAMNHLKKIMKTSKEGVDESFIKEAVL 523
>DNLI_FOWPV (Q67480) DNA ligase (EC 6.5.1.1)
(Polydeoxyribonucleotide synthase [ATP])
Length = 564
Score = 31.6 bits (70), Expect = 5.3
Identities = 11/47 (23%), Positives = 24/47 (50%)
Query: 322 NQIQVLVEENIADYLFEILRLSVPFHPVQYETLKLIYECISECPGAV 368
N ++ +V+ ++ D + ++ L VP P+ K E + +CP +
Sbjct: 185 NNLEQVVQRSLEDNIKPLIELMVPLQPMLASACKTFSEAVKKCPNGI 231
>YL32_ARCFU (O28148) Hypothetical protein AF2132
Length = 244
Score = 31.2 bits (69), Expect = 6.9
Identities = 14/33 (42%), Positives = 21/33 (63%)
Query: 310 LFHYLSSVGTSGNQIQVLVEENIADYLFEILRL 342
L +L G S Q+ V +E+NIAD+++E RL
Sbjct: 55 LSRFLKQHGYSLRQVHVALEQNIADFIYEFQRL 87
>T2BF_BACSU (P25217) Type II restriction enzyme BsuFI (EC 3.1.21.4)
(Endonuclease BsuFI) (R.BsuFI)
Length = 395
Score = 31.2 bits (69), Expect = 6.9
Identities = 20/66 (30%), Positives = 33/66 (49%), Gaps = 5/66 (7%)
Query: 44 LISNPSSPTIHVSYALS----QLSRSLSIPQFLHSIVTFHPHFLVSPLVAALSSFDDEPI 99
L+SN P VS + +L R++SI VT H VS L++ L + +P+
Sbjct: 227 LLSNRGKPKTDVSVTIKTNTKELIRNISIKNTREKTVTIHEGS-VSDLISRLKLSETDPL 285
Query: 100 AEQLVH 105
++ L+H
Sbjct: 286 SQALIH 291
>BRNQ_ECOLI (P37011) Branched-chain amino acid transport system II
carrier protein (LIV-II)
Length = 439
Score = 31.2 bits (69), Expect = 6.9
Identities = 14/51 (27%), Positives = 27/51 (52%)
Query: 35 TICLVCFSNLISNPSSPTIHVSYALSQLSRSLSIPQFLHSIVTFHPHFLVS 85
T+C + L + P + T+ ++ L+ ++P F++S+V F LVS
Sbjct: 87 TVCYLAVGPLFATPRTATVSFEVGIAPLTGDSALPLFIYSLVYFAIVILVS 137
>ACVS_CEPAC (P25464)
N-(5-amino-5-carboxypentanoyl)-L-cysteinyl-D-valine
synthase (EC 6.3.2.26)
(Delta-(L-alpha-aminoadipyl)-L-cysteinyl-D-valine
synthetase) (ACV synthetase) (ACVS)
Length = 3712
Score = 31.2 bits (69), Expect = 6.9
Identities = 21/73 (28%), Positives = 39/73 (52%), Gaps = 6/73 (8%)
Query: 420 DLSKSIEEAVKQAILACLYVS----ERNINQILQCFYLLKEAYAYSHDENSTHISKLELR 475
DL+ +++E+V + Y + E + ++ F+LL A H++ ST +SKL +
Sbjct: 2358 DLNVTVKESVNSLNVNFNYPTSLFEEETVQGFMETFHLLLRQLA--HNKASTSLSKLSVE 2415
Query: 476 SGILDICRTHLLP 488
G+L+ T+L P
Sbjct: 2416 DGVLNPEPTNLQP 2428
>TOLB_CHLTR (O84604) TolB protein precursor
Length = 431
Score = 30.8 bits (68), Expect = 9.0
Identities = 22/61 (36%), Positives = 31/61 (50%), Gaps = 7/61 (11%)
Query: 142 LHSLHCLGVLL----NCEKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLF 197
L SL CL LL +CE DL +H++ SL+ + V+ L P + + L L LF
Sbjct: 8 LRSLLCLLCLLPSTLHCE---DLEIHVRSESSLLPIAVSLLSSPKDSRQASYLASLRDLF 64
Query: 198 A 198
A
Sbjct: 65 A 65
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.137 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 59,713,678
Number of Sequences: 164201
Number of extensions: 2446683
Number of successful extensions: 5160
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 5154
Number of HSP's gapped (non-prelim): 12
length of query: 526
length of database: 59,974,054
effective HSP length: 115
effective length of query: 411
effective length of database: 41,090,939
effective search space: 16888375929
effective search space used: 16888375929
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Medicago: description of AC147484.3


