

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
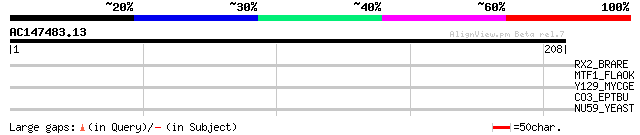
Query= AC147483.13 - phase: 0
(208 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
RX2_BRARE (O42357) Retinal homeobox protein Rx2 33 0.36
MTF1_FLAOK (P14871) Modification methylase FokI (EC 2.1.1.72) (A... 33 0.36
Y129_MYCGE (Q49397) Hypothetical protein MG129 33 0.62
CO3_EPTBU (P98094) Complement C3 [Contains: C3a anaphylatoxin] (... 29 6.8
NU59_YEAST (Q05166) Nucleoporin ASM4 (Nuclear pore protein NUP59) 29 8.9
>RX2_BRARE (O42357) Retinal homeobox protein Rx2
Length = 327
Score = 33.5 bits (75), Expect = 0.36
Identities = 34/113 (30%), Positives = 48/113 (42%), Gaps = 15/113 (13%)
Query: 36 DLVSLRELALKVKNQTGFRLQYGGLLTILRTNVEEKLVHTLVQFYDPSFRCFTFPDFQVV 95
D+ S ELA+KV N R+Q + +EK+ ++ +D R F P +
Sbjct: 161 DVYSREELAMKV-NLPEVRVQVWFQNRRAKWRRQEKMDTGTMKLHDSPIRSFNRPP--MA 217
Query: 96 PTLEAYSHLL------DSPTAEKTP------FTGPGTSLTPLVVAKDLYLKTS 136
P + S+ L SP + TP F GPG SL P A +L TS
Sbjct: 218 PNVGPMSNSLPLDPWLSSPLSSATPMHSIPGFMGPGQSLQPTYTAHPGFLNTS 270
>MTF1_FLAOK (P14871) Modification methylase FokI (EC 2.1.1.72)
(Adenine-specific methyltransferase FokI) (M.FokI)
Length = 647
Score = 33.5 bits (75), Expect = 0.36
Identities = 22/71 (30%), Positives = 33/71 (45%), Gaps = 3/71 (4%)
Query: 103 HLLDSPTAEKTPFTGPGTSLTPLVVAKDLYLKTSDVS---NHLTTKSHIRGFTSKYLLEQ 159
HLL++ P T L P V +K LY + +V NHL K++ R Y E
Sbjct: 231 HLLETIALYDYPEIYGKTGLRPYVESKSLYCQKKEVGNAFNHLIEKANFRHILVSYSSEG 290
Query: 160 ATCQDALEAIL 170
++ +E+IL
Sbjct: 291 LLLEEEIESIL 301
>Y129_MYCGE (Q49397) Hypothetical protein MG129
Length = 117
Score = 32.7 bits (73), Expect = 0.62
Identities = 23/64 (35%), Positives = 33/64 (50%), Gaps = 5/64 (7%)
Query: 13 GQIKNFVNITGMKKTKSYKFKEVDLVSLREL-ALKV----KNQTGFRLQYGGLLTILRTN 67
G +NFVNI FK+V+ VSL +L AL + KNQ FR G + L+
Sbjct: 53 GGKENFVNIKTTPTQLIVTFKDVNSVSLTKLNALNIKGINKNQNQFRFVLGNFVNELKKK 112
Query: 68 VEEK 71
+E++
Sbjct: 113 IEDE 116
>CO3_EPTBU (P98094) Complement C3 [Contains: C3a anaphylatoxin]
(Fragment)
Length = 1620
Score = 29.3 bits (64), Expect = 6.8
Identities = 16/59 (27%), Positives = 28/59 (47%), Gaps = 7/59 (11%)
Query: 48 KNQTGFRLQYGGLLTILRTNVEEKLVHTLVQFYDPSFRCFT--FPDFQVVPTLEAYSHL 104
KN GFRL + +N+ + T+ ++Y+P FRC P +V P + ++
Sbjct: 1418 KNCVGFRLNQ-----VFESNLVLPVTATVFEYYEPDFRCSKSYHPKMEVNPDASCHGNI 1471
>NU59_YEAST (Q05166) Nucleoporin ASM4 (Nuclear pore protein NUP59)
Length = 528
Score = 28.9 bits (63), Expect = 8.9
Identities = 19/80 (23%), Positives = 40/80 (49%), Gaps = 1/80 (1%)
Query: 91 DFQVVPTLEAYSHLLDSPTAEKTPFTGPGTSLTPLVVAKDLYLKTSDVSNHLTTKSHIRG 150
+F +P+L + D TA+KT + + TP V KD Y++ +++ + ++
Sbjct: 194 EFGSIPSLTT-QFVTDKYTAKKTNRSAYDSKNTPNVFDKDSYVRIANIEQNHLDNNYNTA 252
Query: 151 FTSKYLLEQATCQDALEAIL 170
T+ + E ++ +L AI+
Sbjct: 253 ETNNKVHETSSKSSSLSAII 272
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.139 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,071,146
Number of Sequences: 164201
Number of extensions: 977636
Number of successful extensions: 2046
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 2045
Number of HSP's gapped (non-prelim): 5
length of query: 208
length of database: 59,974,054
effective HSP length: 105
effective length of query: 103
effective length of database: 42,732,949
effective search space: 4401493747
effective search space used: 4401493747
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC147483.13


