

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147435.1 - phase: 0 /pseudo
(346 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
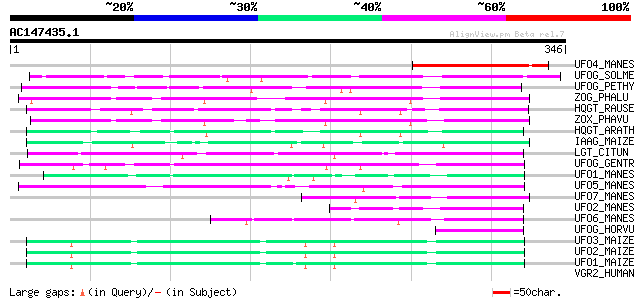
Sequences producing significant alignments: (bits) Value
UFO4_MANES (Q40286) Flavonol 3-O-glucosyltransferase 4 (EC 2.4.1... 106 1e-22
UFOG_SOLME (Q43641) Flavonol 3-O-glucosyltransferase (EC 2.4.1.9... 84 4e-16
UFOG_PETHY (Q43716) Flavonol 3-O-glucosyltransferase (EC 2.4.1.9... 84 5e-16
ZOG_PHALU (Q9ZSK5) Zeatin O-glucosyltransferase (EC 2.4.1.203) (... 81 3e-15
HQGT_RAUSE (Q9AR73) Hydroquinone glucosyltransferase (EC 2.4.1.2... 81 4e-15
ZOX_PHAVU (P56725) Zeatin O-xylosyltransferase (EC 2.4.2.40) (Ze... 80 1e-14
HQGT_ARATH (Q9M156) Probable hydroquinone glucosyltransferase (E... 78 4e-14
IAAG_MAIZE (Q41819) Indole-3-acetate beta-glucosyltransferase (E... 75 2e-13
LGT_CITUN (Q9MB73) Limonoid UDP-glucosyltransferase (EC 2.4.1.21... 70 8e-12
UFOG_GENTR (Q96493) Flavonol 3-O-glucosyltransferase (EC 2.4.1.9... 69 2e-11
UFO1_MANES (Q40284) Flavonol 3-O-glucosyltransferase 1 (EC 2.4.1... 69 2e-11
UFO5_MANES (Q40287) Flavonol 3-O-glucosyltransferase 5 (EC 2.4.1... 66 1e-10
UFO7_MANES (Q40289) Flavonol 3-O-glucosyltransferase 7 (EC 2.4.1... 64 7e-10
UFO2_MANES (Q40285) Flavonol 3-O-glucosyltransferase 2 (EC 2.4.1... 57 5e-08
UFO6_MANES (Q40288) Flavonol 3-O-glucosyltransferase 6 (EC 2.4.1... 55 3e-07
UFOG_HORVU (P14726) Flavonol 3-O-glucosyltransferase (EC 2.4.1.9... 51 4e-06
UFO3_MAIZE (P16167) Flavonol 3-O-glucosyltransferase (EC 2.4.1.9... 50 8e-06
UFO2_MAIZE (P16165) Flavonol 3-O-glucosyltransferase (EC 2.4.1.9... 49 1e-05
UFO1_MAIZE (P16166) Flavonol 3-O-glucosyltransferase (EC 2.4.1.9... 47 5e-05
VGR2_HUMAN (P35968) Vascular endothelial growth factor receptor ... 32 2.4
>UFO4_MANES (Q40286) Flavonol 3-O-glucosyltransferase 4 (EC
2.4.1.91) (UDP-glucose flavonoid 3-O-glucosyltransferase
4) (Fragment)
Length = 241
Score = 106 bits (264), Expect = 1e-22
Identities = 55/85 (64%), Positives = 62/85 (72%), Gaps = 1/85 (1%)
Query: 252 CNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALALEAT 311
CNK LDKA+RG AS+ LKWLDL +P SV+YACLGS+ L QL EL L LE+T
Sbjct: 1 CNKLKLDKAERGDKASVDNTELLKWLDLWEPGSVIYACLGSISGLTSWQLAELGLGLEST 60
Query: 312 NRPFIWVIREGNKSSEELEKWISEE 336
N+PFIWVIREG K SE LEKWI EE
Sbjct: 61 NQPFIWVIREGEK-SEGLEKWILEE 84
>UFOG_SOLME (Q43641) Flavonol 3-O-glucosyltransferase (EC 2.4.1.91)
(UDP-glucose flavonoid 3-O-glucosyltransferase)
Length = 433
Score = 84.3 bits (207), Expect = 4e-16
Identities = 87/338 (25%), Positives = 155/338 (45%), Gaps = 38/338 (11%)
Query: 13 FVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTL 72
F+ FP H P++ + + ++ TIF+ +S +S+ S+ V + IKI +
Sbjct: 10 FLAFPFGT--HATPLLTLVQKISPFLPSSTIFSFFNTSSSNSSIFSK-VPNQENIKIYNV 66
Query: 73 NFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFIP 132
G+ EG + +++I + + L+ K EE + K SCI SD F+
Sbjct: 67 -----WDGVKEGNDTPFGLEAIKL---FIQSTLLISKITEEAEEETGVKFSCIFSDAFL- 117
Query: 133 WT--IQIAENHKIPRISFH-GFSCFC---LHCMLKIQTSKILERVDSESEYFTVPGIPDQ 186
W +++ + P +++ G SC L+ L + ++ S ++ IP +
Sbjct: 118 WCFLVKLPKKMNAPGVAYWTGGSCSLAVHLYTDLIRSNKETSLKIPGFSSTLSINDIPPE 177
Query: 187 IQVTKEQIPGILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKN-GKVWF 245
VT E + G + L +H A+ ++N+F+EL++ + + +KN KV+
Sbjct: 178 --VTAEDLEGPMSSMLYNMALNLHKAD----AVVLNSFQELDRDPLINKDLQKNLQKVFN 231
Query: 246 VGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELA 305
+GP+ L + LD E C++WLD + KSVVY G++ L P+++ +A
Sbjct: 232 IGPLVLQSSRKLD-----------ESGCIQWLDKQKEKSVVYLSFGTVTTLPPNEIGSIA 280
Query: 306 LALEATNRPFIWVIREGNKSSEELEKWISEERNGYRRI 343
ALE PFIW +R N + L K E + +I
Sbjct: 281 EALETKKTPFIWSLR--NNGVKNLPKGFLERTKEFGKI 316
>UFOG_PETHY (Q43716) Flavonol 3-O-glucosyltransferase (EC 2.4.1.91)
(UDP-glucose flavonoid 3-O-glucosyltransferase)
(Anthocyanin rhamnosyl transferase)
Length = 473
Score = 84.0 bits (206), Expect = 5e-16
Identities = 85/328 (25%), Positives = 137/328 (40%), Gaps = 43/328 (13%)
Query: 8 NDVPHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQI 67
ND H V+FP A GHI P + +A L+ GV V+ FT NASR S+L+ A ++
Sbjct: 9 NDALHVVMFPFFAFGHISPFVQLANKLSSYGVKVSFFTASGNASRVKSMLNSAPTT---- 64
Query: 68 KIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIIS 127
IV L P + GLP G E+ + + L A+ L+Q + L L KP ++
Sbjct: 65 HIVPLTLPHVE-GLPPGAESTAELTPASAEL-LKVALDLMQPQIKTLLSHL--KPHFVLF 120
Query: 128 DFFIPWTIQIAENHKIPRISFH---GFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIP 184
DF W ++A I + + S L C ++ K ++ +
Sbjct: 121 DFAQEWLPKMANGLGIKTVYYSVVVALSTAFLTCPARVLEPKKYPSLEDMKK-------- 172
Query: 185 DQIQVTKEQIPGILKGELKEF---------GEKMHD---AEMKSYGEII-NTFEELEKAY 231
+ + + + E ++F G ++D + ++ I+ T ++E Y
Sbjct: 173 PPLGFPQTSVTSVRTFEARDFLYVFKSFHNGPTLYDRIQSGLRGCSAILAKTCSQMEGPY 232
Query: 232 VKDYKKEKNGKVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLG 291
+K + + N V+ +GPV G E WL+ + +V+Y G
Sbjct: 233 IKYVEAQFNKPVFLIGPVVPDPPSGK-----------LEEKWATWLNKFEGGTVIYCSFG 281
Query: 292 SLCNLIPSQLMELALALEATNRPFIWVI 319
S L Q+ ELAL LE T PF V+
Sbjct: 282 SETFLTDDQVKELALGLEQTGLPFFLVL 309
>ZOG_PHALU (Q9ZSK5) Zeatin O-glucosyltransferase (EC 2.4.1.203)
(Trans-zeatin O-beta-D-glucosyltransferase)
Length = 459
Score = 81.3 bits (199), Expect = 3e-15
Identities = 78/334 (23%), Positives = 138/334 (40%), Gaps = 50/334 (14%)
Query: 6 NINDVPH-----FVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRA 60
N +PH +L P AQGH+ + +++L+ + + V T + + T +
Sbjct: 4 NDKSIPHETKVVVLLIPFPAQGHLNQFLHLSRLIVAQNIPVHYVGTVTHIRQATLRYNNP 63
Query: 61 VSSGLQIKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTP 120
S+ I P + P ++F + F A L++P +L +L+
Sbjct: 64 TSN---IHFHAFQVPPFVSPPPNPEDDFP-----SHLIPSFEASAHLREPVGKLLQSLSS 115
Query: 121 --KPSCIISDFFIPWTIQIAEN-HKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEY 177
K +I+D + Q A N + +FH FS F ++ ++
Sbjct: 116 QAKRVVVINDSLMASVAQDAANISNVENYTFHSFSAF-----------------NTSGDF 158
Query: 178 FTVPGIPDQIQVTKEQIP---GILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKD 234
+ G P + P G + + K F ++ + G+I NT +E YV+
Sbjct: 159 WEEMGKPPVGDFHFPEFPSLEGCIAAQFKGFRTAQYEFRKFNNGDIYNTSRVIEGPYVEL 218
Query: 235 YKKEKNGK-VWFVGP---VSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACL 290
+ GK VW +GP +++ KD + H C++WLD +P SV+Y
Sbjct: 219 LELFNGGKKVWALGPFNPLAVEKKDSIG----------FRHPCMEWLDKQEPSSVIYISF 268
Query: 291 GSLCNLIPSQLMELALALEATNRPFIWVIREGNK 324
G+ L Q+ ++A LE + + FIWV+RE +K
Sbjct: 269 GTTTALRDEQIQQIATGLEQSKQKFIWVLREADK 302
>HQGT_RAUSE (Q9AR73) Hydroquinone glucosyltransferase (EC 2.4.1.218)
(Arbutin synthase)
Length = 470
Score = 80.9 bits (198), Expect = 4e-15
Identities = 82/333 (24%), Positives = 137/333 (40%), Gaps = 54/333 (16%)
Query: 11 PHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIV 70
PH + P GH+IP+++ AK L R F P + L +A S L
Sbjct: 5 PHIAMVPTPGMGHLIPLVEFAKRLVLRHNFGVTFIIPTDGP-----LPKAQKSFLDALPA 59
Query: 71 TLNF----PTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCII 126
+N+ P LP D+ + + + ++ ++ + L T K + ++
Sbjct: 60 GVNYVLLPPVSFDDLPA-----DVRIETRICLTITRSLPFVRDAVKTLL--ATTKLAALV 112
Query: 127 SDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQ 186
D F +A K+ F+ + CL + K+ + V E +P+
Sbjct: 113 VDLFGTDAFDVAIEFKVSPYIFYPTTAMCLSLFFHLP--KLDQMVSCEYR-----DVPEP 165
Query: 187 IQVTKEQIPGILKGELKEFGEKMHDAEMKSY--------------GEIINTFEELEKAYV 232
+Q IPG + K+F + D + +Y G ++NTF +LE +
Sbjct: 166 LQ-----IPGCIPIHGKDFLDPAQDRKNDAYKCLLHQAKRYRLAEGIMVNTFNDLEPGPL 220
Query: 233 KDYKKEKNGK--VWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACL 290
K ++E GK V+ +GP L D K + + CLKWLD SV++
Sbjct: 221 KALQEEDQGKPPVYPIGP--LIRADSSSK--------VDDCECLKWLDDQPRGSVLFISF 270
Query: 291 GSLCNLIPSQLMELALALEATNRPFIWVIREGN 323
GS + +Q +ELAL LE + + F+WV+R N
Sbjct: 271 GSGGAVSHNQFIELALGLEMSEQRFLWVVRSPN 303
>ZOX_PHAVU (P56725) Zeatin O-xylosyltransferase (EC 2.4.2.40)
(Zeatin O-beta-D-xylosyltransferase)
Length = 454
Score = 79.7 bits (195), Expect = 1e-14
Identities = 74/320 (23%), Positives = 134/320 (41%), Gaps = 43/320 (13%)
Query: 14 VLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTLN 73
+L P QGH+ P + ++ L+A + + V T + + A S+ I
Sbjct: 12 LLLPFPVQGHLNPFLQLSHLIAAQNIAVHYVGTVTHIRQAKLRYHNATSN---IHFHAFE 68
Query: 74 FPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTP--KPSCIISDFFI 131
P + P ++F + F A L++P +L +L+ K +I+D +
Sbjct: 69 VPPYVSPPPNPEDDFP-----SHLIPSFEASAHLREPVGKLLQSLSSQAKRVVLINDSLM 123
Query: 132 PWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVTK 191
Q A N F +C ++ +++ +++ G P
Sbjct: 124 ASVAQDAAN-------FSNVERYCF---------QVFSALNTAGDFWEQMGKPPLADFHF 167
Query: 192 EQIP---GILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGK-VWFVG 247
IP G + + +F ++ + G+I NT +E YV+ ++ GK VW +G
Sbjct: 168 PDIPSLQGCISAQFTDFLTAQNEFRKFNNGDIYNTSRVIEGPYVELLERFNGGKEVWALG 227
Query: 248 P---VSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMEL 304
P +++ KD + + H C++WLD +P SV+Y G+ L Q+ EL
Sbjct: 228 PFTPLAVEKKDSIGFS----------HPCMEWLDKQEPSSVIYVSFGTTTALRDEQIQEL 277
Query: 305 ALALEATNRPFIWVIREGNK 324
A LE + + FIWV+R+ +K
Sbjct: 278 ATGLEQSKQKFIWVLRDADK 297
>HQGT_ARATH (Q9M156) Probable hydroquinone glucosyltransferase (EC
2.4.1.218) (Arbutin synthase)
Length = 480
Score = 77.8 bits (190), Expect = 4e-14
Identities = 85/330 (25%), Positives = 131/330 (38%), Gaps = 51/330 (15%)
Query: 11 PHFVLFPLIAQGHIIPMIDIAKLLAQ-RGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKI 69
PH + P GH+IP+++ AK L G+ VT + S R V L I
Sbjct: 7 PHVAIIPSPGMGHLIPLVEFAKRLVHLHGLTVTFVIAGEGPP---SKAQRTVLDSLPSSI 63
Query: 70 VTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPK---PSCII 126
++ P P + I+ R++L +T ++FD+ P+ ++
Sbjct: 64 SSVFLP------PVDLTDLSSSTRIESRISL--TVTRSNPELRKVFDSFVEGGRLPTALV 115
Query: 127 SDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQ 186
D F +A +P F+ + L L + K+ E V E T P +
Sbjct: 116 VDLFGTDAFDVAVEFHVPPYIFYPTTANVLSFFLHLP--KLDETVSCEFRELTEPLM--- 170
Query: 187 IQVTKEQIPGILKGELKEFGEKMHDAEMKSY--------------GEIINTFEELEKAYV 232
+PG + K+F + D + +Y G ++NTF ELE +
Sbjct: 171 -------LPGCVPVAGKDFLDPAQDRKDDAYKWLLHNTKRYKEAEGILVNTFFELEPNAI 223
Query: 233 KDYKKEKNGK--VWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACL 290
K ++ K V+ VGP+ K + + E CLKWLD SV+Y
Sbjct: 224 KALQEPGLDKPPVYPVGPLVNIGKQEAKQTE--------ESECLKWLDNQPLGSVLYVSF 275
Query: 291 GSLCNLIPSQLMELALALEATNRPFIWVIR 320
GS L QL ELAL L + + F+WVIR
Sbjct: 276 GSGGTLTCEQLNELALGLADSEQRFLWVIR 305
>IAAG_MAIZE (Q41819) Indole-3-acetate beta-glucosyltransferase (EC
2.4.1.121) (IAA-Glu synthetase) ((Uridine
5'-diphosphate-glucose:indol-3-ylacetyl)-beta-D-glucosyl
transferase)
Length = 471
Score = 75.5 bits (184), Expect = 2e-13
Identities = 83/343 (24%), Positives = 130/343 (37%), Gaps = 55/343 (16%)
Query: 11 PHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIV 70
PH ++ P QGH+ PM+ AK LA +GV T+ TT RF + + ++ +
Sbjct: 3 PHVLVVPFPGQGHMNPMVQFAKRLASKGVATTLVTT-----RFIQRTADVDAHPAMVEAI 57
Query: 71 TLNFP----TKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCII 126
+ AG+ E E S + +L++ A DA T C++
Sbjct: 58 SDGHDEGGFASAAGVAEYLEKQAAAASASLA-------SLVEARASSA-DAFT----CVV 105
Query: 127 SDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSE------------ 174
D + W + +A +P + F SC ++ +
Sbjct: 106 YDSYEDWVLPVARRMGLPAVPFSTQSCAVSAVYYHFSQGRLAVPPGAAADGSDGGAGAAA 165
Query: 175 -SEYFTVPGIPDQIQVTKEQI-------PGILKGELKEFGEKMHDAEMKSYGEIINTFEE 226
SE F G+P+ + P I +K+F D + + N+FEE
Sbjct: 166 LSEAFL--GLPEMERSELPSFVFDHGPYPTIAMQAIKQFAHAGKDDWV-----LFNSFEE 218
Query: 227 LEKAYVKDYKKEKNGKVWFVGPVSLCNKDGLDKAQRGIIASIS-----EHHCLKWLDLHQ 281
LE + K + +GP G G I + E C KWLD
Sbjct: 219 LETEVLAGLTKYLKARA--IGPCVPLPTAGRTAGANGRITYGANLVKPEDACTKWLDTKP 276
Query: 282 PKSVVYACLGSLCNLIPSQLMELALALEATNRPFIWVIREGNK 324
+SV Y GSL +L +Q ELA L A +PF+WV+R ++
Sbjct: 277 DRSVAYVSFGSLASLGNAQKEELARGLLAAGKPFLWVVRASDE 319
>LGT_CITUN (Q9MB73) Limonoid UDP-glucosyltransferase (EC 2.4.1.210)
(Limonoid glucosyltransferase) (Limonoid GTase) (LGTase)
Length = 511
Score = 70.1 bits (170), Expect = 8e-12
Identities = 74/317 (23%), Positives = 128/317 (40%), Gaps = 21/317 (6%)
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
H +L GH+ P++ + +LLA +G +T+ TTP++ + +
Sbjct: 8 HVLLVSFPGHGHVNPLLRLGRLLASKGFFLTL-TTPESFGKQMRKAGNFTYEPTPVGDGF 66
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITL---LQKPAEELFDALTPKPSCIISD 128
+ F + G E + +D ++ L + ++K AEE SC+I++
Sbjct: 67 IRFEFFEDGWDEDDPRREDLDQYMAQLELIGKQVIPKIIKKSAEEYRPV-----SCLINN 121
Query: 129 FFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQ 188
FIPW +AE+ +P SC C L SE E +P
Sbjct: 122 PFIPWVSDVAESLGLPSAMLWVQSCACFAAYYHYFHG--LVPFPSEKEPEIDVQLPCMPL 179
Query: 189 VTKEQIPGILKGE-----LKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKV 243
+ +++P L L+ ++ K + +++TF ELEK + DY K+
Sbjct: 180 LKHDEMPSFLHPSTPYPFLRRAILGQYENLGKPFCILLDTFYELEKEII-DYM----AKI 234
Query: 244 WFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLME 303
+ PV K+ + C+ WLD P SVVY G++ L Q+ E
Sbjct: 235 CPIKPVGPLFKNPKAPTLTVRDDCMKPDECIDWLDKKPPSSVVYISFGTVVYLKQEQVEE 294
Query: 304 LALALEATNRPFIWVIR 320
+ AL + F+WV++
Sbjct: 295 IGYALLNSGISFLWVMK 311
>UFOG_GENTR (Q96493) Flavonol 3-O-glucosyltransferase (EC 2.4.1.91)
(UDP-glucose flavonoid 3-O-glucosyltransferase)
Length = 453
Score = 68.6 bits (166), Expect = 2e-11
Identities = 73/327 (22%), Positives = 136/327 (41%), Gaps = 32/327 (9%)
Query: 7 INDVPHFVLFPLIAQGHIIPMIDIAKLLAQRG--VIVTIFTTPKNASRFTSVLS--RAVS 62
++ V H + H P++ + LA +I + F+T +S T++ S +S
Sbjct: 1 MSPVSHVAVLAFPFGTHAAPLLTLVNRLAASAPDIIFSFFST---SSSITTIFSPTNLIS 57
Query: 63 SGLQIKIVTLNFPTKQAGLPEGCE-NFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPK 121
G IK + G PEG + + + I+ +N A K ++ +
Sbjct: 58 IGSNIKPYAV-----WDGSPEGFVFSGNPREPIEYFLNA--APDNFDKAMKKAVEDTGVN 110
Query: 122 PSCIISDFFIPWTIQIAENHKIPRIS-FHGFSC-FCLHCMLKIQTSKILERVDSESEYFT 179
SC+++D F+ + +E +P I + SC CLH S+ E +E T
Sbjct: 111 ISCLLTDAFLWFAADFSEKIGVPWIPVWTAASCSLCLHVYTDEIRSRFAEFDIAEKAEKT 170
Query: 180 VPGIPDQIQVTKEQIPG--ILKGELKEFGEKMHDAEMKSY---GEIINTFEELEKAYVKD 234
+ IP ++ +P I++ F +H+ +K + +N+FEE++
Sbjct: 171 IDFIPGLSAISFSDLPEELIMEDSQSIFALTLHNMGLKLHKATAVAVNSFEEIDPIITNH 230
Query: 235 YKKEKNGKVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLC 294
+ + +GP+ + ++ CLKWL + SVVY G++
Sbjct: 231 LRSTNQLNILNIGPLQTLSSS----------IPPEDNECLKWLQTQKESSVVYLSFGTVI 280
Query: 295 NLIPSQLMELALALEATNRPFIWVIRE 321
N P+++ LA LE+ PF+W +R+
Sbjct: 281 NPPPNEMAALASTLESRKIPFLWSLRD 307
>UFO1_MANES (Q40284) Flavonol 3-O-glucosyltransferase 1 (EC
2.4.1.91) (UDP-glucose flavonoid 3-O-glucosyltransferase
1)
Length = 449
Score = 68.6 bits (166), Expect = 2e-11
Identities = 78/314 (24%), Positives = 127/314 (39%), Gaps = 42/314 (13%)
Query: 22 GHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTLNFPTKQAGL 81
GH++ ++ AKLL R ++I N S TS + V S + L F L
Sbjct: 2 GHLVSAVETAKLLLSRCHSLSITVLIFNNSVVTSKVHNYVDSQIASSSNRLRF----IYL 57
Query: 82 PEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFIPWTIQIAENH 141
P S+ + H + K E +P+ I D F I +A
Sbjct: 58 PRDETGISSFSSLIEKQKP-HVKESVMKITEFGSSVESPRLVGFIVDMFCTAMIDVANEF 116
Query: 142 KIPRISFHGFSCFCLHCMLKIQTSKILERVD------SESEYFTVPGIPDQIQ------- 188
+P F+ L+ ML +Q E + S+ E VPG+ +
Sbjct: 117 GVPSYIFYTSGAAFLNFMLHVQKIHDEENFNPTEFNASDGE-LQVPGLVNSFPSKAMPTA 175
Query: 189 -VTKEQIPGILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVG 247
++K+ P +L+ + +GE + G IINTF ELE ++ +K + ++ VG
Sbjct: 176 ILSKQWFPPLLENT-RRYGE--------AKGVIINTFFELESHAIESFK---DPPIYPVG 223
Query: 248 PVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALA 307
P+ +G + Q ++WLD P SVV+ C GS + Q+ E+A A
Sbjct: 224 PILDVRSNGRNTNQE----------IMQWLDDQPPSSVVFLCFGSNGSFSKDQVKEIACA 273
Query: 308 LEATNRPFIWVIRE 321
LE + F+W + +
Sbjct: 274 LEDSGHRFLWSLAD 287
>UFO5_MANES (Q40287) Flavonol 3-O-glucosyltransferase 5 (EC
2.4.1.91) (UDP-glucose flavonoid 3-O-glucosyltransferase
5)
Length = 487
Score = 66.2 bits (160), Expect = 1e-10
Identities = 79/332 (23%), Positives = 136/332 (40%), Gaps = 43/332 (12%)
Query: 6 NINDVPHFVLFPLIAQGHIIPMIDIAK-LLAQRGVIVTIFTTPKNASRFTSVLSRAVSSG 64
++N PH VL GH+IP++++ K ++ VTIF + S + R+ +
Sbjct: 5 DLNSKPHIVLLSSPGLGHLIPVLELGKRIVTLCNFDVTIFMVGSDTSAAEPQVLRSAMTP 64
Query: 65 LQIKIVTLNFPTKQAGL-PEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPS 123
+I+ L P + PE + LF + ++ AL +P+
Sbjct: 65 KLCEIIQLPPPNISCLIDPEAT----------VCTRLFVLMREIRPAFRAAVSALKFRPA 114
Query: 124 CIISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGI 183
II D F ++++A+ I + + + + L + + IL++ + E E+
Sbjct: 115 AIIVDLFGTESLEVAKELGIAKYVYIASNAWFLALTIYVP---ILDK-EVEGEFV----- 165
Query: 184 PDQIQVTKEQIPGILKGELKEFGEKMHDAEMKSYGE--------------IINTFEELEK 229
+Q +IPG +E + M D + Y E ++NT+E LE
Sbjct: 166 ---LQKEPMKIPGCRPVRTEEVVDPMLDRTNQQYSEYFRLGIEIPTADGILMNTWEALEP 222
Query: 230 AYVKDYKKEKNGKVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYAC 289
+ K F+G V+ + +R S L WLD +SVVY
Sbjct: 223 TTFGALRDVK-----FLGRVAKVPVFPIGPLRRQAGPCGSNCELLDWLDQQPKESVVYVS 277
Query: 290 LGSLCNLIPSQLMELALALEATNRPFIWVIRE 321
GS L Q++ELA LE + + FIWV+R+
Sbjct: 278 FGSGGTLSLEQMIELAWGLERSQQRFIWVVRQ 309
>UFO7_MANES (Q40289) Flavonol 3-O-glucosyltransferase 7 (EC
2.4.1.91) (UDP-glucose flavonoid 3-O-glucosyltransferase
7) (Fragment)
Length = 287
Score = 63.5 bits (153), Expect = 7e-10
Identities = 43/147 (29%), Positives = 73/147 (49%), Gaps = 15/147 (10%)
Query: 183 IPDQIQVTKEQIP-GILKGELKE-FGEKMHDAEM---KSYGEIINTFEELEKAYVKDYKK 237
IP ++ +P G+L G L+ F + +H+ ++ ++N+FEEL+ V D
Sbjct: 5 IPGMSKIQIRDLPEGVLFGNLESLFSQMLHNMGRMLPRAAAVLMNSFEELDPTIVSDLNS 64
Query: 238 EKNGKVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLI 297
+ N + +GP +L + G C+ WLD +P SV Y GS+
Sbjct: 65 KFNN-ILCIGPFNLVSPPPPVPDTYG---------CMAWLDKQKPASVAYISFGSVATPP 114
Query: 298 PSQLMELALALEATNRPFIWVIREGNK 324
P +L+ LA ALEA+ PF+W +++ +K
Sbjct: 115 PHELVALAEALEASKVPFLWSLKDHSK 141
>UFO2_MANES (Q40285) Flavonol 3-O-glucosyltransferase 2 (EC
2.4.1.91) (UDP-glucose flavonoid 3-O-glucosyltransferase
2) (Fragment)
Length = 346
Score = 57.4 bits (137), Expect = 5e-08
Identities = 38/121 (31%), Positives = 58/121 (47%), Gaps = 17/121 (14%)
Query: 200 GELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVGPVSLCNKDGLDK 259
G+L +K A+ G I+NTF ELE ++ +K ++ VGP+ DG +
Sbjct: 79 GQLLAIAKKFRQAK----GIIVNTFLELESRAIESFKVPP---LYHVGPILDVKSDGRN- 130
Query: 260 AQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALALEATNRPFIWVI 319
+ ++WLD SVV+ C GS+ + QL E+A ALE + F+W I
Sbjct: 131 ---------THPEIMQWLDDQPEGSVVFLCFGSMGSFSEDQLKEIAYALENSGHRFLWSI 181
Query: 320 R 320
R
Sbjct: 182 R 182
>UFO6_MANES (Q40288) Flavonol 3-O-glucosyltransferase 6 (EC
2.4.1.91) (UDP-glucose flavonoid 3-O-glucosyltransferase
6) (Fragment)
Length = 394
Score = 55.1 bits (131), Expect = 3e-07
Identities = 49/204 (24%), Positives = 88/204 (43%), Gaps = 19/204 (9%)
Query: 126 ISDFFIPWTIQIAENHKIPRI-------SFHGFSCFCLHCMLKIQTSKILERVDSESEYF 178
+ D F I +A+ +P +F GF F + + Q + + + DS++E
Sbjct: 35 VLDMFCTSMIDVAKELGVPYYIFFTSGAAFLGF-LFYVQLIHDEQDADLTQFKDSDAE-L 92
Query: 179 TVPGIPDQIQVTKEQIPGILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKE 238
+VP + + + ++K F + ++ G ++NTF ELE + K +
Sbjct: 93 SVPSLANSLPARVLPASMLVKDRFYAFIRIIRGLR-EAKGIMVNTFMELESHALNSLKDD 151
Query: 239 KNG--KVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNL 296
++ ++ VGP+ + D G ++WLD P SVV+ C GS+
Sbjct: 152 QSKIPPIYPVGPILKLSNQENDVGPEG-------SEIIEWLDDQPPSSVVFLCFGSMGGF 204
Query: 297 IPSQLMELALALEATNRPFIWVIR 320
Q E+A ALE + F+W +R
Sbjct: 205 DMDQAKEIACALEQSRHRFLWSLR 228
>UFOG_HORVU (P14726) Flavonol 3-O-glucosyltransferase (EC 2.4.1.91)
(UDP-glucose flavonoid 3-O-glucosyltransferase)
(Bronze-1)
Length = 455
Score = 51.2 bits (121), Expect = 4e-06
Identities = 23/55 (41%), Positives = 29/55 (51%)
Query: 266 ASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALALEATNRPFIWVIR 320
A H CL WLD +SV Y G+ P +L ELA LEA+ PF+W +R
Sbjct: 260 APADPHGCLAWLDRRPARSVAYVSFGTNATARPDELQELAAGLEASGAPFLWSLR 314
>UFO3_MAIZE (P16167) Flavonol 3-O-glucosyltransferase (EC 2.4.1.91)
(UDP-glucose flavonoid 3-O-glucosyltransferase)
(Bronze-1) (Bz-W22 allele)
Length = 471
Score = 50.1 bits (118), Expect = 8e-06
Identities = 74/326 (22%), Positives = 121/326 (36%), Gaps = 26/326 (7%)
Query: 11 PHFVLFPLIAQGHIIPMIDIAKLLAQR----GVIVTIFTTPKNASRFTSVLSRAVSSGLQ 66
PH + H ++ IA+ LA G ++ +T + ++ S + GL
Sbjct: 12 PHVAVVAFPFSSHAAVLLSIARALAAAAAPSGATLSFLSTASSLAQLRKASSASAGHGLP 71
Query: 67 IKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCII 126
+ + P G P E+ + + + M A + A + +C++
Sbjct: 72 GNLRFVEVPD---GAPAAEESVPVPRQMQLFMEAAEAGGVKAWLEAARAAAGGARVTCVV 128
Query: 127 SDFFIPWTIQIAENHKIPRIS-FHGFSCFCLHCMLKIQTSKILERV-DSESEYFTVPGI- 183
D F+ A + P + + SC L I+T + E V D + P I
Sbjct: 129 GDAFVWPAADAAASAGAPWVPVWTAASCALL---AHIRTDALREDVGDQAANRVDEPLIS 185
Query: 184 -PDQIQVTKEQIP-GILKGE----LKEFGEKMHDAEMKSYGEI-INTFEELEKAYVKDYK 236
P +P G++ G+ + +M +S + +NTF L+ V
Sbjct: 186 HPGLASYRVRDLPDGVVSGDFNYVINLLVHRMGQCLPRSAAAVALNTFPGLDPPDVTAAL 245
Query: 237 KEKNGKVWFVGPVSLC-NKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCN 295
E GP L +D D A A H CL WL + V Y G++
Sbjct: 246 AEILPNCVPFGPYHLLLAEDDADTA-----APADPHGCLAWLGRQPARGVAYVSFGTVAC 300
Query: 296 LIPSQLMELALALEATNRPFIWVIRE 321
P +L ELA LEA+ PF+W +RE
Sbjct: 301 PRPDELRELAAGLEASGAPFLWSLRE 326
>UFO2_MAIZE (P16165) Flavonol 3-O-glucosyltransferase (EC 2.4.1.91)
(UDP-glucose flavonoid 3-O-glucosyltransferase)
(Bronze-1) (Bz-Mc2 allele)
Length = 471
Score = 49.3 bits (116), Expect = 1e-05
Identities = 74/326 (22%), Positives = 120/326 (36%), Gaps = 26/326 (7%)
Query: 11 PHFVLFPLIAQGHIIPMIDIAKLLAQR----GVIVTIFTTPKNASRFTSVLSRAVSSGLQ 66
PH + H ++ IA+ LA G ++ +T + ++ S + GL
Sbjct: 12 PHVAVVAFPFSSHAAVLLSIARALAAAAAPSGATLSFLSTASSLAQLRKASSASAGHGLP 71
Query: 67 IKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCII 126
+ + P G P E + + + M A + A + +C++
Sbjct: 72 GNLRFVEVPD---GAPAAEETVPVPRQMQLFMEAAEAGGVKAWLEAARAAAGGARVTCVV 128
Query: 127 SDFFIPWTIQIAENHKIPRIS-FHGFSCFCLHCMLKIQTSKILERV-DSESEYFTVPGI- 183
D F+ A + P + + SC L I+T + E V D + P I
Sbjct: 129 GDAFVWPAADAAASAGAPWVPVWTAASCALL---AHIRTDSLREDVGDQAANRVDEPLIS 185
Query: 184 -PDQIQVTKEQIP-GILKGE----LKEFGEKMHDAEMKSYGEI-INTFEELEKAYVKDYK 236
P +P G++ G+ + +M +S + +NTF L+ V
Sbjct: 186 HPGLASYRVRDLPDGVVSGDFNYVISLLVHRMGQCLPRSAAAVALNTFPGLDPPDVTAAL 245
Query: 237 KEKNGKVWFVGPVSLC-NKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCN 295
E GP L +D D A A H CL WL + V Y G++
Sbjct: 246 AEILPNCVPFGPYHLLLAEDDADTA-----APADPHGCLAWLGRQPARGVAYVSFGTVAC 300
Query: 296 LIPSQLMELALALEATNRPFIWVIRE 321
P +L ELA LEA+ PF+W +RE
Sbjct: 301 PRPDELRELAAGLEASAAPFLWSLRE 326
>UFO1_MAIZE (P16166) Flavonol 3-O-glucosyltransferase (EC 2.4.1.91)
(UDP-glucose flavonoid 3-O-glucosyltransferase)
(Bronze-1) (Bz-McC allele)
Length = 471
Score = 47.4 bits (111), Expect = 5e-05
Identities = 72/327 (22%), Positives = 120/327 (36%), Gaps = 28/327 (8%)
Query: 11 PHFVLFPLIAQGHIIPMIDIAKLLAQR----GVIVTIFTTPKNASRFTSVLSRAVSSGLQ 66
PH + H ++ IA+ LA G ++ +T + ++ S + GL
Sbjct: 12 PHVAVVAFPFSSHAAVLLSIARALAAAAAPSGATLSFLSTASSLAQLRKASSASAGHGLP 71
Query: 67 IKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCII 126
+ + P G P E + + + M A + A + +C++
Sbjct: 72 GNLRFVEVPD---GAPAAEETVPVPRQMQLFMEAAEAGGVKAWLEAARAAAGGARVTCVV 128
Query: 127 SDFFIPWTIQIAENHKIPRIS-FHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGI-- 183
D F+ A + P + + SC L I+T + E V ++ V G+
Sbjct: 129 GDAFVWPAADAAASAGAPWVPVWTAASCALL---AHIRTDALREDVGDQAAN-RVDGLLI 184
Query: 184 --PDQIQVTKEQIP-GILKGE----LKEFGEKMHDAEMKSYGEI-INTFEELEKAYVKDY 235
P +P G++ G+ + +M +S + +NTF L+ V
Sbjct: 185 SHPGLASYRVRDLPDGVVSGDFNYVINLLVHRMGQCLPRSAAAVALNTFPGLDPPDVTAA 244
Query: 236 KKEKNGKVWFVGPVSLC-NKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLC 294
E GP L +D D A A H CL WL + V Y G++
Sbjct: 245 LAEILPNCVPFGPYHLLLAEDDADTA-----APADPHGCLAWLGRQPARGVAYVSFGTVA 299
Query: 295 NLIPSQLMELALALEATNRPFIWVIRE 321
P +L ELA LE + PF+W +RE
Sbjct: 300 CPRPDELRELAAGLEDSGAPFLWSLRE 326
>VGR2_HUMAN (P35968) Vascular endothelial growth factor receptor 2
precursor (EC 2.7.1.112) (VEGFR-2) (Kinase insert domain
receptor) (Protein-tyrosine kinase receptor Flk-1)
Length = 1356
Score = 32.0 bits (71), Expect = 2.4
Identities = 14/32 (43%), Positives = 23/32 (71%)
Query: 265 IASISEHHCLKWLDLHQPKSVVYACLGSLCNL 296
IAS+S+ H + ++ ++ K+VV CLGS+ NL
Sbjct: 126 IASVSDQHGVVYITENKNKTVVIPCLGSISNL 157
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.139 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,382,895
Number of Sequences: 164201
Number of extensions: 1746401
Number of successful extensions: 4610
Number of sequences better than 10.0: 35
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 4568
Number of HSP's gapped (non-prelim): 39
length of query: 346
length of database: 59,974,054
effective HSP length: 111
effective length of query: 235
effective length of database: 41,747,743
effective search space: 9810719605
effective search space used: 9810719605
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 66 (30.0 bits)
Medicago: description of AC147435.1


