

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
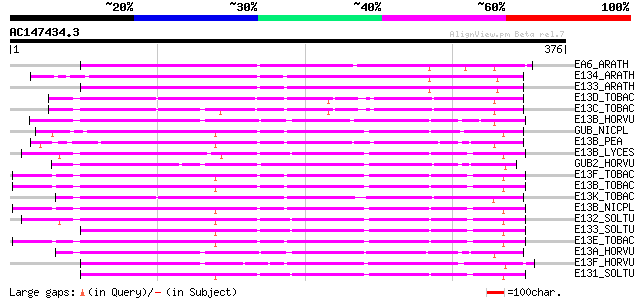
Query= AC147434.3 - phase: 0
(376 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
EA6_ARATH (Q06915) Probable glucan endo-1,3-beta-glucosidase A6 ... 187 3e-47
E134_ARATH (Q94CD8) Putative glucan endo-1,3-beta-glucosidase 4 ... 176 9e-44
E133_ARATH (Q9ZU91) Putative glucan endo-1,3-beta-glucosidase 3 ... 168 2e-41
E13D_TOBAC (P23433) Glucan endo-1,3-beta-glucosidase precursor (... 153 6e-37
E13C_TOBAC (P23432) Glucan endo-1,3-beta-glucosidase precursor (... 151 3e-36
E13B_HORVU (P15737) Glucan endo-1,3-beta-glucosidase GII precurs... 150 5e-36
GUB_NICPL (P07979) Lichenase precursor (EC 3.2.1.73) (Endo-beta-... 148 2e-35
E13B_PEA (Q03467) Glucan endo-1,3-beta-glucosidase precursor (EC... 146 7e-35
E13B_LYCES (Q01413) Glucan endo-1,3-beta-glucosidase B precursor... 144 3e-34
GUB2_HORVU (P12257) Lichenase II precursor (EC 3.2.1.73) (Endo-b... 144 5e-34
E13F_TOBAC (P27666) Glucan endo-1,3-beta-glucosidase, basic vacu... 143 6e-34
E13B_TOBAC (P15797) Glucan endo-1,3-beta-glucosidase, basic vacu... 143 8e-34
E13K_TOBAC (P52398) Glucan endo-1,3-beta-glucosidase, acidic iso... 142 1e-33
E13B_NICPL (P23431) Glucan endo-1,3-beta-glucosidase, basic vacu... 142 1e-33
E132_SOLTU (P52401) Glucan endo-1,3-beta-glucosidase, basic isof... 142 1e-33
E133_SOLTU (P52402) Glucan endo-1,3-beta-glucosidase, basic isof... 142 2e-33
E13E_TOBAC (P23546) Glucan endo-1,3-beta-glucosidase, basic vacu... 141 2e-33
E13A_HORVU (P34742) Glucan endo-1,3-beta-glucosidase GI (EC 3.2.... 141 2e-33
E13F_HORVU (Q02439) Putative glucan endo-1,3-beta-glucosidase GV... 141 3e-33
E131_SOLTU (P52400) Glucan endo-1,3-beta-glucosidase, basic isof... 141 3e-33
>EA6_ARATH (Q06915) Probable glucan endo-1,3-beta-glucosidase A6
precursor (EC 3.2.1.39) ((1->3)-beta-glucan
endohydrolase) ((1->3)-beta-glucanase)
(Beta-1,3-endoglucanase) (Anther-specific protein A6)
Length = 478
Score = 187 bits (476), Expect = 3e-47
Identities = 113/315 (35%), Positives = 185/315 (57%), Gaps = 13/315 (4%)
Query: 49 PSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAW 108
PSP + ++++K H++L + DP + L TN+ + +T+PN+ +T +++N++IA W
Sbjct: 54 PSPYQSINFIKSIKAGHVKLYDADPESLTLLSQTNLYVTITVPNHQITALSSNQTIADEW 113
Query: 109 IYTHVLPFYPRAKITTISVGNAFTDVYPESIN-NLLPAISNVHISLRDLGIRKISVSTSF 167
+ T++LP+YP+ +I + VGN +++ NL+PA+ + SLR GI I V T
Sbjct: 114 VRTNILPYYPQTQIRFVLVGNEILSYNSGNVSVNLVPAMRKIVNSLRLHGIHNIKVGTPL 173
Query: 168 SFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLG 226
+ + ++ S FPPS+ F+E ++ PLL+FL+ TNS F +N++PY + P L
Sbjct: 174 A-MDSLRSSFPPSNGTFREEITGPVMLPLLKFLNGTNSYFFLNVHPYFRWSRNPMNTSLD 232
Query: 227 IALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSG--S 284
ALFQ H + D TG+ Y+NL D M+D+V+ AM GY + + ++ETGWP+ G
Sbjct: 233 FALFQGH--STYTDPQTGLVYRNLLDQMLDSVLFAMTKLGYPHMRLAISETGWPNFGDID 290
Query: 285 ELDANSAYAEIYLKGLVKHLKSG--AGTPLLKDGVKGVYLYELFD---KEGAGNGRNWGI 339
E AN A Y + L+K + + GTP +++ LF+ K G+G R+WGI
Sbjct: 291 ETGANILNAATYNRNLIKKMSASPPIGTPSRPGLPIPTFVFSLFNENQKSGSGTQRHWGI 350
Query: 340 LYPNGSTKYDKIDFS 354
L+P+GS YD +DF+
Sbjct: 351 LHPDGSPIYD-VDFT 364
>E134_ARATH (Q94CD8) Putative glucan endo-1,3-beta-glucosidase 4
precursor (EC 3.2.1.39) ((1->3)-beta-glucan
endohydrolase) ((1->3)-beta-glucanase)
(Beta-1,3-endoglucanase) (Beta-1,3-glucanase)
Length = 505
Score = 176 bits (446), Expect = 9e-44
Identities = 121/343 (35%), Positives = 185/343 (53%), Gaps = 18/343 (5%)
Query: 15 SLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDPS 74
+L LLLS+L + + IGV T + PPPS I T +++ ++TH+RL + +
Sbjct: 10 ALLLLLSILACSNAAF--IGVNIG--TDLTNMPPPSD--IVTLLKSQQITHVRLYDANSH 63
Query: 75 IIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTDV 134
++++ T+I + + + N + I S A AW+ +V + P IT I+VG+
Sbjct: 64 MLKAFANTSIEVMVGVTNEEILKIGRFPSAAAAWVNKNVAAYIPSTNITAIAVGSEVLTT 123
Query: 135 YPESINNLLPAISNVHISLRDLGIR-KISVSTSFSFVTAVTSPFPPSSAAFQEIPGVN-L 192
P L A++N+H +L + K+ VS+ S + + PFPPS++ F P N
Sbjct: 124 IPHVAPILASALNNIHKALVASNLNFKVKVSSPMS-MDIMPKPFPPSTSTFS--PSWNTT 180
Query: 193 IGPLLQFLSDTNSSFLINLYPYNLYRLRPEI-PLGIALFQE-HPFNFRDDFTTGVRYKNL 250
+ LLQFL +T S F++N YPY Y I PL ALF++ P D T + Y ++
Sbjct: 181 VYQLLQFLKNTGSFFMLNAYPYYGYTTANGIFPLDYALFKQLSPVKQIVDPNTLLHYNSM 240
Query: 251 FDIMVDAVVSAMALEGYETIPVIVTETGWPSSG--SELDANSAYAEIYLKGLVKHLKSGA 308
FD MVDA +M + IPV+VTETGWPSSG E A A AE + L+K + + +
Sbjct: 241 FDAMVDAAYYSMEALNFSKIPVVVTETGWPSSGGSDEAAATVANAETFNTNLIKRVLNNS 300
Query: 309 GTPLLKDGVKGVYLYELFDKE---GAGNGRNWGILYPNGSTKY 348
G P D Y+YEL++++ G + RNWGIL+PNG++ Y
Sbjct: 301 GPPSQPDIPINTYIYELYNEDKRSGPVSERNWGILFPNGTSVY 343
>E133_ARATH (Q9ZU91) Putative glucan endo-1,3-beta-glucosidase 3
precursor (EC 3.2.1.39) ((1->3)-beta-glucan
endohydrolase) ((1->3)-beta-glucanase)
(Beta-1,3-endoglucanase) (Beta-1,3-glucanase)
Length = 501
Score = 168 bits (425), Expect = 2e-41
Identities = 104/308 (33%), Positives = 166/308 (53%), Gaps = 10/308 (3%)
Query: 49 PSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAW 108
PSP ++ +++ + +RL + D S++ + +T + + +++PN + I+ + + A W
Sbjct: 35 PSPTQVVALLKSQNINRVRLYDADRSMLLAFAHTGVQVIISVPNDQLLGISQSNATAANW 94
Query: 109 IYTHVLPFYPRAKITTISVGNAFTDVYPESINNLLPAISNVHISLRDLGI-RKISVSTSF 167
+ +V +YP ITTI+VG+ + + L+ A+ + +L + R+I VST
Sbjct: 95 VTRNVAAYYPATNITTIAVGSEVLTSLTNAASVLVSALKYIQAALVTANLDRQIKVSTPH 154
Query: 168 SFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLY-RLRPEIPLG 226
S T + FPPS A F + +I PLL+FL T S L+N+YPY Y + IPL
Sbjct: 155 S-STIILDSFPPSQAFFNK-TWDPVIVPLLKFLQSTGSPLLLNVYPYFDYVQSNGVIPLD 212
Query: 227 IALFQEHPFNFRD-DFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSG-- 283
ALFQ N D T + Y N+FD +VDA AM+ + IP++VTE+GWPS G
Sbjct: 213 YALFQPLQANKEAVDANTLLHYTNVFDAIVDAAYFAMSYLNFTNIPIVVTESGWPSKGGP 272
Query: 284 SELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKGVYLYELFDKE---GAGNGRNWGIL 340
SE DA A Y L++H+ + GTP Y+YEL++++ G + +NWG+
Sbjct: 273 SEHDATVENANTYNSNLIQHVINKTGTPKHPGTAVTTYIYELYNEDTRPGPVSEKNWGLF 332
Query: 341 YPNGSTKY 348
Y NG+ Y
Sbjct: 333 YTNGTPVY 340
>E13D_TOBAC (P23433) Glucan endo-1,3-beta-glucosidase precursor (EC
3.2.1.39) ((1->3)-beta-glucan endohydrolase)
((1->3)-beta-glucanase) (Beta-1,3-endoglucanase)
Length = 351
Score = 153 bits (387), Expect = 6e-37
Identities = 104/329 (31%), Positives = 173/329 (51%), Gaps = 23/329 (6%)
Query: 27 TTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISL 86
T + IGV Y PS + + + +R+ PD +I ++L +NI +
Sbjct: 30 TGAQSNIGVCYGEIANN----LPSEQDVINLYKANGIRKMRIYYPDTNIFKALNGSNIEI 85
Query: 87 FLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTDVYPESINN-LLPA 145
L +PN + +A N SIA W+ ++ +P K IS+GN + + LL A
Sbjct: 86 ILEVPNQDLEALA-NSSIANGWVQDNIRSHFPYVKFKYISIGNEVSPTNNGQYSQFLLHA 144
Query: 146 ISNVHISLRDLGIR-KISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTN 204
+ NV+ +L G++ KI VST+ ++ + + +PP + F+E + I P+++FL+ N
Sbjct: 145 MKNVYNALAAAGLQDKIKVSTA-TYSGLLANTYPPKDSIFRE-ELKSFINPIIEFLARNN 202
Query: 205 SSFLINLYPY--NLYRLRPEIPLGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAM 262
L N+YPY ++Y ++PL ALF + N +TG Y+NLFD ++D++ A+
Sbjct: 203 LPLLANIYPYFGHIYNT-VDVPLSYALFNQQETN-----STG--YQNLFDALLDSIYFAV 254
Query: 263 ALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKGVYL 322
G + +IV+E+GWPS G+ A A+ Y + LV H+K GAGTP + YL
Sbjct: 255 EKAGGPNVEIIVSESGWPSEGNSA-ATIENAQTYYRNLVNHVKGGAGTPKKPGRIIETYL 313
Query: 323 YELFD---KEGAGNGRNWGILYPNGSTKY 348
+ +FD K+G +++G+ YPN + KY
Sbjct: 314 FAMFDENEKQGEITEKHFGLFYPNRAAKY 342
>E13C_TOBAC (P23432) Glucan endo-1,3-beta-glucosidase precursor (EC
3.2.1.39) ((1->3)-beta-glucan endohydrolase)
((1->3)-beta-glucanase) (Beta-1,3-endoglucanase)
Length = 351
Score = 151 bits (381), Expect = 3e-36
Identities = 105/332 (31%), Positives = 173/332 (51%), Gaps = 29/332 (8%)
Query: 27 TTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISL 86
T + IGV Y PS + + + +R+ D +I +SL +NI +
Sbjct: 30 TGAQSNIGVCYGKIANN----LPSEQDVINLYKANGIRKMRIYNSDTNIFKSLNGSNIEI 85
Query: 87 FLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTDVYPESINN----L 142
L +PN + +A N SIA W+ ++ +P K IS+GN +V P + L
Sbjct: 86 ILDVPNQDLEALA-NSSIANGWVQDNIRSHFPYVKFKYISIGN---EVSPSNNGQYSQFL 141
Query: 143 LPAISNVHISLRDLGIR-KISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLS 201
L A+ NV+ +L G++ KI V+T+ ++ + + +PP + F+E + I P+++FL+
Sbjct: 142 LHAMENVYNALAAAGLQDKIKVTTA-TYSGLLANTYPPKDSIFRE-EFKSFINPIIEFLA 199
Query: 202 DTNSSFLINLYPY--NLYRLRPEIPLGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVV 259
N L N+YPY ++Y ++PL ALF + N +TG Y+NLFD ++D++
Sbjct: 200 RNNLPLLANIYPYFGHIYNT-VDVPLSYALFNQQGTN-----STG--YQNLFDALLDSIY 251
Query: 260 SAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKG 319
A+ G + +IV+E+GWPS G+ A A+ Y + LV H+K GAGTP +
Sbjct: 252 FAVEKAGGPNVEIIVSESGWPSEGNSA-ATIENAQTYYRNLVNHVKGGAGTPKKPGRIVE 310
Query: 320 VYLYELFD---KEGAGNGRNWGILYPNGSTKY 348
YL+ +FD K G +++G+ YPN + KY
Sbjct: 311 TYLFAMFDENEKNGEVTEKHFGLFYPNRTAKY 342
>E13B_HORVU (P15737) Glucan endo-1,3-beta-glucosidase GII precursor
(EC 3.2.1.39) ((1->3)-beta-glucan endohydrolase GII)
((1->3)-beta-glucanase isoenzyme GII)
(Beta-1,3-endoglucanase GII)
Length = 334
Score = 150 bits (379), Expect = 5e-36
Identities = 106/340 (31%), Positives = 171/340 (50%), Gaps = 22/340 (6%)
Query: 14 FSLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDP 73
F+ L + TS+ +IGV Y PS + ++ + +R+ D
Sbjct: 10 FAAALFIGAFAAVPTSVQSIGVCYGVIGNN----LPSRSDVVQLYRSKGINGMRIYFADG 65
Query: 74 SIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTD 133
+ +L + I L L I N + IA + S A +W+ +V P+YP I I+ GN +
Sbjct: 66 QALSALRNSGIGLILDIGNDQLANIAASTSNAASWVQNNVRPYYPAVNIKYIAAGN---E 122
Query: 134 VYPESINNLLPAISNVHISLRDLGIRKISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLI 193
V + ++LPA+ N++ +L G+ I VSTS F V + FPPS+ F+ +
Sbjct: 123 VQGGATQSILPAMRNLNAALSAAGLGAIKVSTSIRF-DEVANSFPPSAGVFKNA----YM 177
Query: 194 GPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLGIALFQEHPFNFRDDFTTGVRYKNLFD 252
+ + L+ T + L N+YPY YR P I L A FQ P D G+ Y +LFD
Sbjct: 178 TDVARLLASTGAPLLANVYPYFAYRDNPGSISLNYATFQ--PGTTVRDQNNGLTYTSLFD 235
Query: 253 IMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLKGLVKHLKSGAGTPL 312
MVDAV +A+ G + V+V+E+GWPS+G A++ A Y +GL+ H+ G GTP
Sbjct: 236 AMVDAVYAALEKAGAPAVKVVVSESGWPSAGG-FAASAGNARTYNQGLINHV--GGGTPK 292
Query: 313 LKDGVKGVYLYELFD---KEGAGNGRNWGILYPNGSTKYD 349
++ ++ Y++ +F+ K G R++G+ P+ S Y+
Sbjct: 293 KREALE-TYIFAMFNENQKTGDATERSFGLFNPDKSPAYN 331
>GUB_NICPL (P07979) Lichenase precursor (EC 3.2.1.73)
(Endo-beta-1,3-1,4 glucanase)
Length = 370
Score = 148 bits (374), Expect = 2e-35
Identities = 108/339 (31%), Positives = 181/339 (52%), Gaps = 20/339 (5%)
Query: 18 LLLSLLPTTT--TSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDPSI 75
+LL LL ++T ++GV Y ++ PP S ++ ++ + +RL +P+ +
Sbjct: 16 ILLGLLVSSTEIVGAQSVGVCYGMLG--NNLPPAS--QVVQLYKSKNIRRMRLYDPNQAA 71
Query: 76 IRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTDVY 135
+++L +NI + L +PN + IA N S A W+ +V F+P K I+VGN + V
Sbjct: 72 LQALRGSNIEVMLGVPNSDLQNIAANPSNANNWVQRNVRNFWPAVKFRYIAVGNEVSPVT 131
Query: 136 PES--INNLLPAISNVHISLRDLGIR-KISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNL 192
S LLPA+ N+ ++ G++ I VS+S +T + + FPPS +F+ +
Sbjct: 132 GTSSLTRYLLPAMRNIRNAISSAGLQNNIKVSSSVD-MTLIGNSFPPSQGSFRN-DVRSF 189
Query: 193 IGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLGIALFQEHPFNFRDDFTTGVRYKNLF 251
I P++ F+ NS L+N+YPY Y P +I L ALF +D + Y+NLF
Sbjct: 190 IDPIIGFVRRINSPLLVNIYPYFSYAGNPRDISLPYALFTAPNVVVQDG---SLGYRNLF 246
Query: 252 DIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLKGLVKHLKSGAGTP 311
D M DAV +A++ G +I ++V+E+GWPS+G+ A + A Y K L++H+K G+P
Sbjct: 247 DAMSDAVYAALSRAGGGSIEIVVSESGWPSAGA-FAATTNNAATYYKNLIQHVK--RGSP 303
Query: 312 LLKDGVKGVYLYELFDKEGAGN--GRNWGILYPNGSTKY 348
+ V YL+ +FD+ +++G+ PN KY
Sbjct: 304 RRPNKVIETYLFAMFDENNKNPELEKHFGLFSPNKQPKY 342
>E13B_PEA (Q03467) Glucan endo-1,3-beta-glucosidase precursor (EC
3.2.1.39) ((1->3)-beta-glucan endohydrolase)
((1->3)-beta-glucanase) (Beta-1,3-endoglucanase)
Length = 370
Score = 146 bits (369), Expect = 7e-35
Identities = 114/344 (33%), Positives = 180/344 (52%), Gaps = 26/344 (7%)
Query: 15 SLYLL----LSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEE 70
SL+LL ++L+PTT IG+ Y + PP+ + I N + +RL +
Sbjct: 15 SLFLLELFTINLIPTTDAQ---IGICYGM---MGNNLPPANEVIALYKAN-NIKRMRLYD 67
Query: 71 PDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNA 130
P+ + +L + I L L IPN + +ATN+ AR W+ +VL FYP KI I+VGN
Sbjct: 68 PNQPALNALRDSGIELILGIPNSDLQTLATNQDSARQWVQRNVLNFYPSVKIKYIAVGNE 127
Query: 131 FTDVYPES--INNLLPAISNVHISLRDLGIR-KISVSTSFSFVTAVTSPFPPSSAAFQEI 187
+ V S +LPA NV+ ++R G+ +I V+T+ +T + + FPPS +F+
Sbjct: 128 VSPVGGSSWLAQYVLPATQNVYQAIRAQGLHDQIKVTTAID-MTLIGNSFPPSKGSFRS- 185
Query: 188 PGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLGIALFQEHPFNFRDDFTTGVR 246
+ + P + +L + L+N+YPY + P +I L ALF P D G
Sbjct: 186 DVRSYLDPFIGYLVYAGAPLLVNVYPYFSHIGNPRDISLPYALFTS-PGVMVQDGPNG-- 242
Query: 247 YKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLKGLVKHLKS 306
Y+NLFD M+D+V +A+ G + V+V+E+GWPS G + A IYL L++H+
Sbjct: 243 YQNLFDAMLDSVHAALDNTGIGWVNVVVSESGWPSDGGSATSYD-NARIYLDNLIRHV-- 299
Query: 307 GAGTPLLKDGVKGVYLYELFD--KEGAGNGRNWGILYPNGSTKY 348
G GTP + YL+ +FD ++ +++G+ YPN KY
Sbjct: 300 GKGTP-RRPWATEAYLFAMFDENQKSPELEKHFGVFYPNKQKKY 342
>E13B_LYCES (Q01413) Glucan endo-1,3-beta-glucosidase B precursor
(EC 3.2.1.39) ((1->3)-beta-glucan endohydrolase B)
((1->3)-beta-glucanase B) (Basic beta-1,3-glucanase)
(Beta-1,3-endoglucanase B)
Length = 360
Score = 144 bits (364), Expect = 3e-34
Identities = 106/351 (30%), Positives = 180/351 (51%), Gaps = 25/351 (7%)
Query: 9 ISLTTFSLYLLLSLLPTTTTSLPT--IGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHL 66
++ + ++ +LL LL T + IGV Y PS + ++ + L
Sbjct: 1 MATSQIAIIVLLGLLVATNIHITEAQIGVCYGMMGNN----LPSHSEVIQLYKSRNIRRL 56
Query: 67 RLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTIS 126
RL +P+ + +L +NI + L +PN V I++ AR W+ +V F+P KI I+
Sbjct: 57 RLYDPNHGALNALRGSNIEVILGLPNVDVKHISSGMEHARWWVQKNVRDFWPHVKIKYIA 116
Query: 127 VGNAFTDVYPESINNL----LPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSPFPPSS 181
VGN + V +NL +PA+ N++ ++ + G+ I VSTS +T + + +PPS
Sbjct: 117 VGNEISPV--TGTSNLAPFQVPALVNIYKAIGEAGLGNDIKVSTSVD-MTLIGNSYPPSQ 173
Query: 182 AAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLGIALFQEHPFNFRDD 240
+F+ P++ FL DT + L+N+YPY Y P +I L ALF +D
Sbjct: 174 GSFRNDVRW-FTDPIVGFLRDTRAPLLVNIYPYFSYSGNPGQISLPYALFTAPNVVVQDG 232
Query: 241 FTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLKGL 300
+Y+NLFD M+D+V +AM G ++ ++V+E+GWPS+G+ A A+ YL+ L
Sbjct: 233 ---SRQYRNLFDAMLDSVYAAMDRTGGGSVGIVVSESGWPSAGA-FGATHENAQTYLRNL 288
Query: 301 VKHLKSGAGTPLLKDGVKGVYLYELFDKEGAGN--GRNWGILYPNGSTKYD 349
++H K G+ K G Y++ +FD+ +++G+ PN KY+
Sbjct: 289 IQHAKEGSPR---KPGPIETYIFAMFDENNKNPELEKHFGMFSPNKQPKYN 336
>GUB2_HORVU (P12257) Lichenase II precursor (EC 3.2.1.73)
(Endo-beta-1,3-1,4 glucanase II)
((1->3,1->4)-beta-glucanase isoenzyme EII) (Fragment)
Length = 312
Score = 144 bits (362), Expect = 5e-34
Identities = 97/318 (30%), Positives = 165/318 (51%), Gaps = 20/318 (6%)
Query: 29 SLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISLFL 88
S+ +IGV Y + P+ + + + + +RL P+ + ++++ T I++ +
Sbjct: 3 SVESIGVCYGMSANN----LPAASTVVSMFKFNGIKSMRLYAPNQAALQAVGGTGINVVV 58
Query: 89 TIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTDVYPESINNLLPAISN 148
PN +++ +A + + A +W+ +++ YP+ + VGN +V + NL+PA+ N
Sbjct: 59 GAPNDVLSNLAASPAAAASWVKSNIQA-YPKVSFRYVCVGN---EVAGGATRNLVPAMKN 114
Query: 149 VHISLRDLGIRKISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFL 208
VH +L G+ I V+TS S A+ F P SA +GP++QFL+ TN+ +
Sbjct: 115 VHGALVAAGLGHIKVTTSVS--QAILGVFSPPSAGSFTGEAAAFMGPVVQFLARTNAPLM 172
Query: 209 INLYPYNLYRLRPE-IPLGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAMALEGY 267
N+YPY + P + +G ALF RD Y+NLFD VDA +AM G
Sbjct: 173 ANIYPYLAWAYNPSAMDMGYALFNASGTVVRDG---AYGYQNLFDTTVDAFYTAMGKHGG 229
Query: 268 ETIPVIVTETGWPSSGSELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKGVYLYELFD 327
++ ++V+E+GWPS G A A A Y + L+ H+ G GTP G Y++ +F+
Sbjct: 230 SSVKLVVSESGWPSGGGTA-ATPANARFYNQHLINHV--GRGTP-RHPGAIETYIFAMFN 285
Query: 328 KEGAGNG--RNWGILYPN 343
+ +G +NWG+ YPN
Sbjct: 286 ENQKDSGVEQNWGLFYPN 303
>E13F_TOBAC (P27666) Glucan endo-1,3-beta-glucosidase, basic
vacuolar isoform GLB precursor (EC 3.2.1.39)
((1->3)-beta-glucan endohydrolase)
((1->3)-beta-glucanase) (Beta-1,3-endoglucanase, basic)
(Glucanase GLB)
Length = 370
Score = 143 bits (361), Expect = 6e-34
Identities = 105/353 (29%), Positives = 180/353 (50%), Gaps = 21/353 (5%)
Query: 3 HHSTFMISLTTFSLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLK 62
H++ M ++T L L+ S + +IGV Y P+ + ++
Sbjct: 7 HNTPQMAAITLLGLLLVASTIEIAGAQ--SIGVCYGMLGNN----LPNHWEVIQLYKSRN 60
Query: 63 LTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKI 122
+ LRL +P+ +++L +NI + L +PN V IA+ AR W+ +V F+P KI
Sbjct: 61 IGRLRLYDPNHGALQALKGSNIEVMLGLPNSDVKHIASGMEHARWWVQKNVKDFWPDVKI 120
Query: 123 TTISVGNAFTDVYPES--INNLLPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSPFPP 179
I+VGN + V S + L PA+ N++ ++ + G+ I VSTS +T + + +PP
Sbjct: 121 KYIAVGNEISPVTGTSYLTSFLTPAMVNIYKAIGEAGLGNNIKVSTSVD-MTLIGNSYPP 179
Query: 180 SSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLGIALFQEHPFNFR 238
S +F+ P++ FL DT + L+N+YPY Y P +I L +LF +
Sbjct: 180 SQGSFRN-DARWFTDPIVGFLRDTRAPLLVNIYPYFSYSGNPGQISLPYSLFTAPNVVVQ 238
Query: 239 DDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLK 298
D +Y+NLFD M+D+V +A+ G ++ ++V+E+GWPS+G+ A A YL+
Sbjct: 239 DG---SRQYRNLFDAMLDSVYAALERSGGASVGIVVSESGWPSAGA-FGATYDNAATYLR 294
Query: 299 GLVKHLKSGAGTPLLKDGVKGVYLYELFDKEGAGN--GRNWGILYPNGSTKYD 349
L++H K G+ K G Y++ +FD+ +++G+ PN KY+
Sbjct: 295 NLIQHAKEGSPR---KPGPIETYIFAMFDENNKNPELEKHFGLFSPNKQPKYN 344
>E13B_TOBAC (P15797) Glucan endo-1,3-beta-glucosidase, basic
vacuolar isoform precursor (EC 3.2.1.39)
((1->3)-beta-glucan endohydrolase)
((1->3)-beta-glucanase) (Beta-1,3-endoglucanase, basic)
Length = 371
Score = 143 bits (360), Expect = 8e-34
Identities = 105/353 (29%), Positives = 180/353 (50%), Gaps = 21/353 (5%)
Query: 3 HHSTFMISLTTFSLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLK 62
H++ M ++T L L+ S + +IGV Y P+ + ++
Sbjct: 8 HNTPQMAAITLLGLLLVASSIDIAGAQ--SIGVCYGMLGNN----LPNHWEVIQLYKSRN 61
Query: 63 LTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKI 122
+ LRL +P+ +++L +NI + L +PN V IA+ AR W+ +V F+P KI
Sbjct: 62 IGRLRLYDPNHGALQALKGSNIEVMLGLPNSDVKHIASGMEHARWWVQKNVKDFWPDVKI 121
Query: 123 TTISVGNAFTDVYPES--INNLLPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSPFPP 179
I+VGN + V S + L PA+ N++ ++ + G+ I VSTS +T + + +PP
Sbjct: 122 KYIAVGNEISPVTGTSYLTSFLTPAMVNIYKAIGEAGLGNNIKVSTSVD-MTLIGNSYPP 180
Query: 180 SSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLGIALFQEHPFNFR 238
S +F+ P++ FL DT + L+N+YPY Y P +I L +LF +
Sbjct: 181 SQGSFRN-DARWFTDPIVGFLRDTRAPLLVNIYPYFSYSGNPGQISLPYSLFTAPNVVVQ 239
Query: 239 DDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLK 298
D +Y+NLFD M+D+V +A+ G ++ ++V+E+GWPS+G+ A A YL+
Sbjct: 240 DG---SRQYRNLFDAMLDSVYAALERSGGASVGIVVSESGWPSAGA-FGATYDNAATYLR 295
Query: 299 GLVKHLKSGAGTPLLKDGVKGVYLYELFDKEGAGN--GRNWGILYPNGSTKYD 349
L++H K G+ K G Y++ +FD+ +++G+ PN KY+
Sbjct: 296 NLIQHAKEGSPR---KPGPIETYIFAMFDENNKNPELEKHFGLFSPNKQPKYN 345
>E13K_TOBAC (P52398) Glucan endo-1,3-beta-glucosidase, acidic
isoform GL161 precursor (EC 3.2.1.39)
((1->3)-beta-glucan endohydrolase)
((1->3)-beta-glucanase) (Beta-1,3-endoglucanase)
Length = 331
Score = 142 bits (359), Expect = 1e-33
Identities = 95/322 (29%), Positives = 161/322 (49%), Gaps = 19/322 (5%)
Query: 32 TIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISLFLTIP 91
+IGV Y PS + + LR+ PD +I ++L +NI + L +P
Sbjct: 11 SIGVCYGKAANN----LPSDQDVINLYNANGIRKLRIYYPDKNIFKALNGSNIEIILGVP 66
Query: 92 NYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTDVYPESINN-LLPAISNVH 150
N + +A N SIA W+ ++ +P K IS+GN + + + LL A+ NV+
Sbjct: 67 NQDLEALA-NSSIANGWVQDNIRSHFPYVKFKYISIGNKVSPTNNDQYSEFLLQAMKNVY 125
Query: 151 ISLRDLGIRKISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLIN 210
+L G++ + ++ ++ + + +PP + F+E + I P++QFL+ N L N
Sbjct: 126 NALAAAGLQDMIKVSTVTYSGVLANTYPPERSIFRE-EFKSFINPIIQFLARNNLPLLAN 184
Query: 211 LYPYNLYRLRP-EIPLGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAMALEGYET 269
+YPY ++ ++ L ALF + T Y+NLFD ++D++ A+ G
Sbjct: 185 VYPYFVHVSNTADVSLSYALFTQQG-------TNSAGYQNLFDAILDSMYFAVEKAGGPN 237
Query: 270 IPVIVTETGWPSSGSELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKGVYLYELF--- 326
+ +IV+E+GWPS GS A A+ Y + L+ H+KSGAGTP YL+ +F
Sbjct: 238 VEIIVSESGWPSEGSSA-ATIENAQTYYRNLINHVKSGAGTPKKPGKTIETYLFAMFDEN 296
Query: 327 DKEGAGNGRNWGILYPNGSTKY 348
DK G +++G+ P+ KY
Sbjct: 297 DKIGEITEKHFGLFSPDQRAKY 318
>E13B_NICPL (P23431) Glucan endo-1,3-beta-glucosidase, basic
vacuolar isoform precursor (EC 3.2.1.39)
((1->3)-beta-glucan endohydrolase)
((1->3)-beta-glucanase) (Beta-1,3-endoglucanase, basic)
Length = 365
Score = 142 bits (359), Expect = 1e-33
Identities = 105/353 (29%), Positives = 180/353 (50%), Gaps = 21/353 (5%)
Query: 3 HHSTFMISLTTFSLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLK 62
H++ M ++T L L+ S + +IGV Y P+ + ++
Sbjct: 7 HNTPQMAAITLLGLLLVASSIEIAGAE--SIGVCYGMLGNN----LPNHWEVIQLYKSRN 60
Query: 63 LTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKI 122
+ LRL +P+ +++L +NI + L +PN V IA+ AR W+ +V F+P KI
Sbjct: 61 IGRLRLYDPNHGALQALKGSNIEVMLGLPNSDVKHIASGMEHARWWVQKNVKDFWPDVKI 120
Query: 123 TTISVGNAFTDVYPES--INNLLPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSPFPP 179
I+VGN + V S + L PA+ N++ ++ + G+ I VSTS +T + + +PP
Sbjct: 121 KYIAVGNEISPVTGTSYLTSFLTPAMVNIYKAIGEAGLGNNIKVSTSVD-MTLIGNSYPP 179
Query: 180 SSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLGIALFQEHPFNFR 238
S +F+ + P++ FL DT + L+N+YPY Y P +I L +LF +
Sbjct: 180 SQGSFRN-DARWFVDPIVGFLRDTRAPLLVNIYPYFSYSGNPGQISLPYSLFTAPNVVVQ 238
Query: 239 DDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLK 298
D +Y+NLFD M+D+V +A+ G ++ ++V+E+GWPS+G+ A A YLK
Sbjct: 239 DG---SRQYRNLFDAMLDSVYAALERSGGASVGIVVSESGWPSAGA-FGATYDNAATYLK 294
Query: 299 GLVKHLKSGAGTPLLKDGVKGVYLYELFDKEGAGN--GRNWGILYPNGSTKYD 349
L++H K G+ K Y++ +FD+ +++G+ PN KY+
Sbjct: 295 NLIQHAKEGSPR---KPRPIETYIFAMFDENNKNPELEKHFGLFSPNKQPKYN 344
>E132_SOLTU (P52401) Glucan endo-1,3-beta-glucosidase, basic isoform
2 precursor (EC 3.2.1.39) ((1->3)-beta-glucan
endohydrolase) ((1->3)-beta-glucanase)
(Beta-1,3-endoglucanase)
Length = 363
Score = 142 bits (358), Expect = 1e-33
Identities = 105/349 (30%), Positives = 177/349 (50%), Gaps = 21/349 (6%)
Query: 9 ISLTTFSLYLLLSLLPTTTTSLPT--IGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHL 66
++ + ++ +LL LL T + +GV Y PS + ++ + L
Sbjct: 1 MATSQIAVIVLLGLLVATNIHITEAQLGVCYGMMGNN----LPSHSEVIQLYKSRNIGRL 56
Query: 67 RLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTIS 126
RL +P+ + +L +NI + L +PN V IA+ AR W+ +V F+P KI I+
Sbjct: 57 RLYDPNQGALNALRGSNIEVILGLPNVDVKHIASGMEHARWWVQKNVKDFWPDVKIKYIA 116
Query: 127 VGNAFTDVYPES--INNLLPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSPFPPSSAA 183
VGN + V S + +PA+ N++ ++ + G+ I VSTS +T + + +PPS +
Sbjct: 117 VGNEISPVTGTSSLTSFQVPALVNIYKAVGEAGLGNDIKVSTSVD-MTLIGNSYPPSQGS 175
Query: 184 FQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLGIALFQEHPFNFRDDFT 242
F+ P++ FL DT + L+N+YPY Y P +I L ALF +D
Sbjct: 176 FRNDVRW-FTDPIVGFLRDTRAPLLVNIYPYFSYSGNPGQISLPYALFTAPNVVVQDG-- 232
Query: 243 TGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLKGLVK 302
+Y+NLFD M+D+V +AM G ++ ++V+E GWPS+G+ A A YL+ L++
Sbjct: 233 -SRQYRNLFDAMLDSVYAAMERTGGGSVGIVVSECGWPSAGA-FGATQDNAATYLRNLIQ 290
Query: 303 HLKSGAGTPLLKDGVKGVYLYELFDKEGAGN--GRNWGILYPNGSTKYD 349
H K G+ K G Y++ +FD+ +++G+ PN KY+
Sbjct: 291 HAKEGSPR---KPGPIETYIFAMFDENNKNPELEKHFGLFSPNKQPKYN 336
>E133_SOLTU (P52402) Glucan endo-1,3-beta-glucosidase, basic isoform
3 precursor (EC 3.2.1.39) ((1->3)-beta-glucan
endohydrolase) ((1->3)-beta-glucanase)
(Beta-1,3-endoglucanase) (Fragment)
Length = 328
Score = 142 bits (357), Expect = 2e-33
Identities = 97/307 (31%), Positives = 162/307 (52%), Gaps = 15/307 (4%)
Query: 49 PSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAW 108
PS + ++ + LRL +P+ + +L +NI + L +PN V IA+ AR W
Sbjct: 4 PSHSEVIQLYKSRNIGRLRLYDPNHGALNALRGSNIEVILGLPNVDVKHIASGMEHARWW 63
Query: 109 IYTHVLPFYPRAKITTISVGNAFTDVYPES--INNLLPAISNVHISLRDLGI-RKISVST 165
+ +V F+P KI I+VGN + V S + +PA+ N++ ++ + G+ I VST
Sbjct: 64 VQKNVKDFWPDVKIKYIAVGNEISPVTGTSSLTSFQVPALVNIYKAIGEAGLGNDIKVST 123
Query: 166 SFSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIP 224
S +T + + +PPS +F+ P++ FL DT + L+N+YPY Y P +I
Sbjct: 124 SVD-MTLIGNSYPPSQGSFRNDVRW-FTDPIVGFLRDTRAPLLVNIYPYFSYSGNPGQIS 181
Query: 225 LGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGS 284
L ALF +D +Y+NLFD M+D+V +AM G ++ ++V+E+GWPS+G+
Sbjct: 182 LPYALFTAPNVVVQDG---SRQYRNLFDAMLDSVYAAMERTGGGSVGIVVSESGWPSAGA 238
Query: 285 ELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKGVYLYELFDKEGAGN--GRNWGILYP 342
A A YL+ L++H K G+ K G Y++ +FD+ +++G+ P
Sbjct: 239 -FGATQDNAATYLRNLIQHAKEGSPR---KPGPIETYIFAMFDENNKNPELEKHFGLFSP 294
Query: 343 NGSTKYD 349
N KY+
Sbjct: 295 NKQPKYN 301
>E13E_TOBAC (P23546) Glucan endo-1,3-beta-glucosidase, basic
vacuolar isoform GGIB50 precursor (EC 3.2.1.39)
((1->3)-beta-glucan endohydrolase)
((1->3)-beta-glucanase) (Beta-1,3-endoglucanase, basic)
(Glucanase GLA)
Length = 370
Score = 141 bits (356), Expect = 2e-33
Identities = 104/353 (29%), Positives = 180/353 (50%), Gaps = 21/353 (5%)
Query: 3 HHSTFMISLTTFSLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLK 62
H++ M ++T L L+ S + +IGV Y P+ + ++
Sbjct: 7 HNTPQMAAITLLGLLLVASSIDIAGAQ--SIGVCYGMLGNN----LPNHWEVIQLYKSRN 60
Query: 63 LTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKI 122
+ LRL +P+ +++L +NI + L +PN V IA+ AR W+ +V F+P KI
Sbjct: 61 IGRLRLYDPNHGALQALKGSNIEVMLGLPNSDVKHIASGMEHARWWVQKNVKDFWPDVKI 120
Query: 123 TTISVGNAFTDVYPES--INNLLPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSPFPP 179
I+VGN + V S + L PA+ N++ ++ + G+ I VSTS +T + + +PP
Sbjct: 121 KYIAVGNEISPVTGTSYLTSFLTPAMVNIYKAIGEAGLGNNIKVSTSVD-MTLIGNSYPP 179
Query: 180 SSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLGIALFQEHPFNFR 238
S +F+ + ++ FL DT + L+N+YPY Y P +I L +LF +
Sbjct: 180 SQGSFRN-DARWFVDAIVGFLRDTRAPLLVNIYPYFSYSGNPGQISLPYSLFTAPNVVVQ 238
Query: 239 DDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLK 298
D +Y+NLFD M+D+V +A+ G ++ ++V+E+GWPS+G+ A A YL+
Sbjct: 239 DG---SRQYRNLFDAMLDSVYAALERSGGASVGIVVSESGWPSAGA-FGATYDNAATYLR 294
Query: 299 GLVKHLKSGAGTPLLKDGVKGVYLYELFDKEGAGN--GRNWGILYPNGSTKYD 349
L++H K G+ K G Y++ +FD+ +++G+ PN KY+
Sbjct: 295 NLIQHAKEGSPR---KPGPIETYIFAMFDENNKNPELEKHFGLFSPNKQPKYN 344
>E13A_HORVU (P34742) Glucan endo-1,3-beta-glucosidase GI (EC
3.2.1.39) ((1->3)-beta-glucan endohydrolase GI)
((1->3)-beta-glucanase isoenzyme GI)
(Beta-1,3-endoglucanase GI)
Length = 310
Score = 141 bits (356), Expect = 2e-33
Identities = 104/322 (32%), Positives = 165/322 (50%), Gaps = 22/322 (6%)
Query: 32 TIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISLFLTIP 91
TIGV Y P + + ++ LT +R+ D + +L + I L L +
Sbjct: 1 TIGVCYGVVANN----LPPANEVVQLYRSNGLTGMRIYFADAKALSALRGSGIGLILDVG 56
Query: 92 -NYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTDVYPESINNLLPAISNVH 150
N ++ +A N S A W+ +V P+YP I I+ GN +V+ N++PA+ N+
Sbjct: 57 GNDVLASLAANASNAANWVRDNVRPYYPAVNIKYIAAGN---EVWGGDTQNIVPAMRNLG 113
Query: 151 ISLRDLGIRKISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLIN 210
+L+ G+ I VSTS F AVT+ FPPS+ F + + + + L+ T + L N
Sbjct: 114 AALKAPGLGTIKVSTSIRF-DAVTNTFPPSNGVFAQA----YMTDVARLLASTGAPLLTN 168
Query: 211 LYPYNLYRLRP-EIPLGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAMALEGYET 269
+YPY Y+ P +I L A F+ RD TG+ + LFD MVDAVV+A+ G
Sbjct: 169 VYPYFAYKDNPRDIQLNYATFRPGTTTVRDP-NTGLTSQCLFDAMVDAVVAALERSGAPG 227
Query: 270 IPVIVTETGWPSSGSELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKGVYLYELFD-- 327
+ V+V+E+GWPS+ S A + A Y +GL+ H+ G GTP + G Y++ +F+
Sbjct: 228 VRVVVSESGWPSA-SGFAATADNARAYNQGLIDHV--GGGTP-KRPGALETYIFAMFNEN 283
Query: 328 -KEGAGNGRNWGILYPNGSTKY 348
K G +++G+ P+ S Y
Sbjct: 284 FKTGELTEKHFGLFNPDKSPAY 305
>E13F_HORVU (Q02439) Putative glucan endo-1,3-beta-glucosidase GVI
precursor (EC 3.2.1.39) ((1->3)-beta-glucan
endohydrolase GVI) ((1->3)-beta-glucanase isoenzyme GVI)
(Beta-1,3-endoglucanase GVI) (Fragment)
Length = 321
Score = 141 bits (355), Expect = 3e-33
Identities = 94/312 (30%), Positives = 165/312 (52%), Gaps = 17/312 (5%)
Query: 49 PSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAW 108
PSPD++ + +T +R+ PD +++ +L + + + L N + P+A++ S A +W
Sbjct: 19 PSPDKVVALYKANNITDVRIFHPDTNVLEALRNSGLGVVLGTLNSDLAPLASDASYAASW 78
Query: 109 IYTHVLPFYPRAKITTISVGNAFTDVYP-ESINNLLPAISNVHISLRDLGIRKISVSTSF 167
++++V PF I+ GN +V P ES +LPA+ N+ +L+ G+ + V+T+
Sbjct: 79 VHSYVQPFAGAVSFRYINAGN---EVIPGESAALVLPAMKNLEAALQAAGL-SVPVTTAM 134
Query: 168 SFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLG 226
+ TS +PPS F E + +GP++ L+ + + L+N+YPY Y P + L
Sbjct: 135 ATSVLGTS-YPPSQGTFSE-AALPTVGPIVSHLASSGTPLLVNVYPYFAYSADPSSVRLD 192
Query: 227 IALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAM-ALEGYETIPVIVTETGWPSSGSE 285
AL D GV Y N+FD ++DAV +A+ G E++ ++V+ETGWPS G
Sbjct: 193 YALLSSSAAVAVTD--NGVEYANMFDAILDAVYAAVEKAGGGESLELVVSETGWPSGGGG 250
Query: 286 LDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKGVYLYELFDKEGAGNG--RNWGILYPN 343
A+ A Y+ LV+H+ GTP Y++ +F++ G +N+G+ P+
Sbjct: 251 YGASVENAAAYINNLVRHV---GGTPRRPGKAVETYIFAMFNENQKPEGVEQNFGMFQPD 307
Query: 344 GSTKYDKIDFSS 355
S Y +DF++
Sbjct: 308 MSQVY-HVDFTA 318
>E131_SOLTU (P52400) Glucan endo-1,3-beta-glucosidase, basic isoform
1 precursor (EC 3.2.1.39) ((1->3)-beta-glucan
endohydrolase) ((1->3)-beta-glucanase)
(Beta-1,3-endoglucanase) (Fragment)
Length = 337
Score = 141 bits (355), Expect = 3e-33
Identities = 97/307 (31%), Positives = 162/307 (52%), Gaps = 15/307 (4%)
Query: 49 PSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAW 108
PS + ++ + LRL +P+ + +L +NI + L +PN V IA+ AR W
Sbjct: 13 PSHSEVIQLYKSRNIGRLRLYDPNHGALNALRGSNIEVILGLPNVDVKHIASGMEHARWW 72
Query: 109 IYTHVLPFYPRAKITTISVGNAFTDVYPES--INNLLPAISNVHISLRDLGI-RKISVST 165
+ +V F+P KI I+VGN + V S + +PA+ N++ ++ + G+ I VST
Sbjct: 73 VQKNVKDFWPDVKIKYIAVGNEISPVTGTSSLTSFQVPALVNIYKAVGEAGLGNDIKVST 132
Query: 166 SFSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIP 224
S +T + + +PPS +F+ P++ FL DT + L+N+YPY Y P +I
Sbjct: 133 SVD-MTLIGNSYPPSQGSFRNDVRW-FTDPIVGFLRDTRAPLLVNIYPYFSYSGNPGQIS 190
Query: 225 LGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGS 284
L ALF +D +Y+NLFD M+D+V +AM G ++ ++V+E+GWPS+G+
Sbjct: 191 LPYALFTAPNAVVQDG---SRQYRNLFDAMLDSVYAAMERTGGGSVGIVVSESGWPSAGA 247
Query: 285 ELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKGVYLYELFDKEGAGN--GRNWGILYP 342
A A YL+ L++H K G+ K G Y++ +FD+ +++G+ P
Sbjct: 248 -FGATQDNAATYLRNLIQHAKEGSPR---KPGPIETYIFAMFDENNKNPELEKHFGLFSP 303
Query: 343 NGSTKYD 349
N KY+
Sbjct: 304 NKQPKYN 310
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.139 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 45,268,031
Number of Sequences: 164201
Number of extensions: 2047679
Number of successful extensions: 5987
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 45
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 5740
Number of HSP's gapped (non-prelim): 86
length of query: 376
length of database: 59,974,054
effective HSP length: 112
effective length of query: 264
effective length of database: 41,583,542
effective search space: 10978055088
effective search space used: 10978055088
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC147434.3


