

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
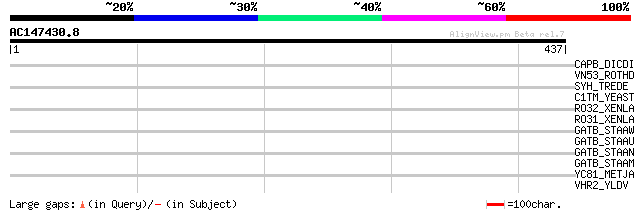
Query= AC147430.8 + phase: 1 /pseudo
(437 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CAPB_DICDI (P13021) F-actin capping protein beta subunit (CAP32) 34 0.65
VN53_ROTHD (P35423) Nonstructural RNA-binding protein 53 (NS53) ... 32 2.5
SYH_TREDE (P60920) Histidyl-tRNA synthetase (EC 6.1.1.21) (Histi... 32 2.5
C1TM_YEAST (P09440) C-1-tetrahydrofolate synthase, mitochondrial... 32 2.5
RO32_XENLA (P51992) Heterogeneous nuclear ribonucleoprotein A3 h... 32 4.2
RO31_XENLA (P51968) Heterogeneous nuclear ribonucleoprotein A3 h... 32 4.2
GATB_STAAW (P64202) Aspartyl/glutamyl-tRNA(Asn/Gln) amidotransfe... 31 7.2
GATB_STAAU (Q9RF06) Aspartyl/glutamyl-tRNA(Asn/Gln) amidotransfe... 31 7.2
GATB_STAAN (P99169) Aspartyl/glutamyl-tRNA(Asn/Gln) amidotransfe... 31 7.2
GATB_STAAM (P64201) Aspartyl/glutamyl-tRNA(Asn/Gln) amidotransfe... 31 7.2
YC81_METJA (Q58677) Hypothetical protein MJ1281 30 9.4
VHR2_YLDV (Q9DHP6) Probable host range protein 2 30 9.4
>CAPB_DICDI (P13021) F-actin capping protein beta subunit (CAP32)
Length = 272
Score = 34.3 bits (77), Expect = 0.65
Identities = 25/96 (26%), Positives = 43/96 (44%), Gaps = 8/96 (8%)
Query: 101 RGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYKAAGEMLMRKRK 160
RGT V + K + ++++ V+ + DN DN AG + + K
Sbjct: 148 RGTWDSIHVVEVKLGKKDKAVYKLTSTVMLSIETDN------DNTGKVNLAGSLTRQDEK 201
Query: 161 KLYWTPCAAHCLDL--MLEDLEKKIPIHGETIPKGK 194
+ + HC+++ M+ED+E K+ ETI GK
Sbjct: 202 EYTFNEVDTHCVNIGKMVEDMESKLRQTLETIYFGK 237
>VN53_ROTHD (P35423) Nonstructural RNA-binding protein 53 (NS53)
(NCVP2)
Length = 486
Score = 32.3 bits (72), Expect = 2.5
Identities = 28/104 (26%), Positives = 48/104 (45%), Gaps = 19/104 (18%)
Query: 49 IRVKLLNQEVKSTND---ALEEHKREWKKTSCTIMTDE--WTDRRRRTILNFLVNSPRGT 103
I KL+ ++N A E H +W C+I + W D R + + N + N
Sbjct: 297 IASKLIKPNYVASNHNSLATEVHNCKW----CSINNNSIVWNDFRIKNVYNDIFN----- 347
Query: 104 VFLKSVDASNI----CKTSEKIFEMIDAVVEEVGEDNVVQVVTD 143
F++++ SN+ C + EKI+E I V+ E+ +VT+
Sbjct: 348 -FIRALVKSNLYVGHCSSEEKIYESIKEVLNVCKENEWNMLVTE 390
>SYH_TREDE (P60920) Histidyl-tRNA synthetase (EC 6.1.1.21)
(Histidine--tRNA ligase) (HisRS)
Length = 448
Score = 32.3 bits (72), Expect = 2.5
Identities = 14/59 (23%), Positives = 36/59 (60%), Gaps = 1/59 (1%)
Query: 92 ILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAVVEEVGEDNVVQVVTDNAANYKA 150
+ NF ++ +F + + + N+ + SE++ ++D + + +GED V++++TD ++ A
Sbjct: 156 VSNFKIHVSHRGIFNRFLKSLNLSEDSEEVLRIVDKLAK-IGEDEVLKLLTDISSEESA 213
>C1TM_YEAST (P09440) C-1-tetrahydrofolate synthase, mitochondrial
precursor (C1-THF synthase) [Includes:
Methylenetetrahydrofolate dehydrogenase (EC 1.5.1.5);
Methenyltetrahydrofolate cyclohydrolase (EC 3.5.4.9);
Formyltetrahydrofolate synthetase (EC 6.3.
Length = 975
Score = 32.3 bits (72), Expect = 2.5
Identities = 20/71 (28%), Positives = 35/71 (49%), Gaps = 6/71 (8%)
Query: 151 AGEMLMRKRKKLYWTPCAAHCLDLMLEDLEKKIP-----IHGETIPKGKKITTFIYARTS 205
AGE L +K K Y+ PC + +LE+ K+ + G + G I + + +
Sbjct: 161 AGE-LAKKGGKPYFIPCTPYGCMKLLEEAHVKLDGKNAVVLGRSSIVGNPIASLLKNANA 219
Query: 206 LISILHSHTKN 216
+++ HSHT+N
Sbjct: 220 TVTVCHSHTRN 230
>RO32_XENLA (P51992) Heterogeneous nuclear ribonucleoprotein A3
homolog 2 (hnRNP A3(B))
Length = 385
Score = 31.6 bits (70), Expect = 4.2
Identities = 14/32 (43%), Positives = 20/32 (61%), Gaps = 1/32 (3%)
Query: 59 KSTNDALEEHKREW-KKTSCTIMTDEWTDRRR 89
++T+D+L EH +W K T C +M D T R R
Sbjct: 37 ETTDDSLREHFEQWGKLTDCVVMRDPQTKRSR 68
>RO31_XENLA (P51968) Heterogeneous nuclear ribonucleoprotein A3
homolog 1 (hnRNP A3(A))
Length = 373
Score = 31.6 bits (70), Expect = 4.2
Identities = 14/32 (43%), Positives = 20/32 (61%), Gaps = 1/32 (3%)
Query: 59 KSTNDALEEHKREW-KKTSCTIMTDEWTDRRR 89
++T+D+L EH +W K T C +M D T R R
Sbjct: 37 ETTDDSLREHFEQWGKLTDCVVMRDPQTKRSR 68
>GATB_STAAW (P64202) Aspartyl/glutamyl-tRNA(Asn/Gln)
amidotransferase subunit B (EC 6.3.5.-) (Asp/Glu-ADT
subunit B)
Length = 475
Score = 30.8 bits (68), Expect = 7.2
Identities = 27/115 (23%), Positives = 54/115 (46%), Gaps = 6/115 (5%)
Query: 47 YDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIM--TDEWTDRRRRTILNFLVNSPRGTV 104
YD V L +E+ ++ EH + K TS +M +E+ ++ + +L+ +
Sbjct: 312 YDAHVLTLTKEMSDFFESTIEHGADVKLTSNWLMGGVNEYLNKNQVELLDTKLTPENLAG 371
Query: 105 FLKSV-DASNICKTSEKIFEMIDAV---VEEVGEDNVVQVVTDNAANYKAAGEML 155
+K + D + K ++K+F + A +++ EDN + ++D A K E L
Sbjct: 372 MIKLIEDGTMSSKIAKKVFPELAAKGGNAKQIMEDNGLVQISDEATLLKFVNEAL 426
>GATB_STAAU (Q9RF06) Aspartyl/glutamyl-tRNA(Asn/Gln)
amidotransferase subunit B (EC 6.3.5.-) (Asp/Glu-ADT
subunit B)
Length = 475
Score = 30.8 bits (68), Expect = 7.2
Identities = 27/115 (23%), Positives = 54/115 (46%), Gaps = 6/115 (5%)
Query: 47 YDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIM--TDEWTDRRRRTILNFLVNSPRGTV 104
YD V L +E+ ++ EH + K TS +M +E+ ++ + +L+ +
Sbjct: 312 YDAHVLTLTKEMSDFFESTIEHGADVKLTSNWLMGGVNEYLNKNQVELLDTKLTPENLAG 371
Query: 105 FLKSV-DASNICKTSEKIFEMIDAV---VEEVGEDNVVQVVTDNAANYKAAGEML 155
+K + D + K ++K+F + A +++ EDN + ++D A K E L
Sbjct: 372 MIKLIEDGTMSSKIAKKVFPELAAKGGNAKQIMEDNGLVQISDEATLLKFVNEAL 426
>GATB_STAAN (P99169) Aspartyl/glutamyl-tRNA(Asn/Gln)
amidotransferase subunit B (EC 6.3.5.-) (Asp/Glu-ADT
subunit B)
Length = 475
Score = 30.8 bits (68), Expect = 7.2
Identities = 27/115 (23%), Positives = 54/115 (46%), Gaps = 6/115 (5%)
Query: 47 YDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIM--TDEWTDRRRRTILNFLVNSPRGTV 104
YD V L +E+ ++ EH + K TS +M +E+ ++ + +L+ +
Sbjct: 312 YDAHVLTLTKEMSDFFESTIEHGADVKLTSNWLMGGVNEYLNKNQVELLDTKLTPENLAG 371
Query: 105 FLKSV-DASNICKTSEKIFEMIDAV---VEEVGEDNVVQVVTDNAANYKAAGEML 155
+K + D + K ++K+F + A +++ EDN + ++D A K E L
Sbjct: 372 MIKLIEDGTMSSKIAKKVFPELAAKGGNAKQIMEDNGLVQISDEATLLKFVNEAL 426
>GATB_STAAM (P64201) Aspartyl/glutamyl-tRNA(Asn/Gln)
amidotransferase subunit B (EC 6.3.5.-) (Asp/Glu-ADT
subunit B)
Length = 475
Score = 30.8 bits (68), Expect = 7.2
Identities = 27/115 (23%), Positives = 54/115 (46%), Gaps = 6/115 (5%)
Query: 47 YDIRVKLLNQEVKSTNDALEEHKREWKKTSCTIM--TDEWTDRRRRTILNFLVNSPRGTV 104
YD V L +E+ ++ EH + K TS +M +E+ ++ + +L+ +
Sbjct: 312 YDAHVLTLTKEMSDFFESTIEHGADVKLTSNWLMGGVNEYLNKNQVELLDTKLTPENLAG 371
Query: 105 FLKSV-DASNICKTSEKIFEMIDAV---VEEVGEDNVVQVVTDNAANYKAAGEML 155
+K + D + K ++K+F + A +++ EDN + ++D A K E L
Sbjct: 372 MIKLIEDGTMSSKIAKKVFPELAAKGGNAKQIMEDNGLVQISDEATLLKFVNEAL 426
>YC81_METJA (Q58677) Hypothetical protein MJ1281
Length = 1048
Score = 30.4 bits (67), Expect = 9.4
Identities = 16/56 (28%), Positives = 29/56 (51%)
Query: 367 DIEVRKGLMDCITRMVEDPEEQAKIEVQMHDFKKQIGYFGTKIAKLTLDKSTPADW 422
+I++ K D + V+ E A E +++K IG+ G K+ +T++K DW
Sbjct: 759 EIDLAKIKKDVVVANVDVDELMAGAEDNKNNYKIVIGWKGNKLYLITMEKDKFEDW 814
>VHR2_YLDV (Q9DHP6) Probable host range protein 2
Length = 178
Score = 30.4 bits (67), Expect = 9.4
Identities = 19/81 (23%), Positives = 38/81 (46%), Gaps = 10/81 (12%)
Query: 69 KREWKKTSCTIMTDEWTDRRRRTILNFLVNSPRGTVFLKSVDASNICKTSEKIFEMIDAV 128
K E K ++ +W++ + + +VN+ SVD K SE +++++ +
Sbjct: 38 KEEKKFNIIFVLKPDWSEVDKVKPIRMIVNN-------NSVDVE---KVSESLYQVVYSA 87
Query: 129 VEEVGEDNVVQVVTDNAANYK 149
+ D+ V+V +DN YK
Sbjct: 88 SFSINSDSYVKVFSDNPDKYK 108
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.138 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 50,650,648
Number of Sequences: 164201
Number of extensions: 2090964
Number of successful extensions: 6566
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 6563
Number of HSP's gapped (non-prelim): 14
length of query: 437
length of database: 59,974,054
effective HSP length: 113
effective length of query: 324
effective length of database: 41,419,341
effective search space: 13419866484
effective search space used: 13419866484
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 67 (30.4 bits)
Medicago: description of AC147430.8


