

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147406.4 - phase: 0
(310 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
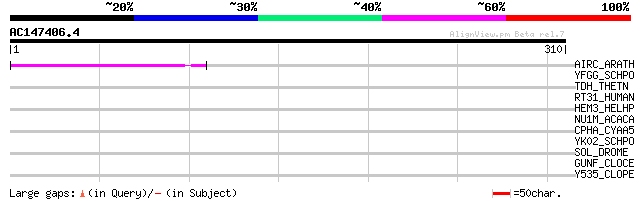
Score E
Sequences producing significant alignments: (bits) Value
AIRC_ARATH (Q94BT2) Auxin-induced in root cultures protein 12 pr... 44 7e-04
YFGG_SCHPO (O13854) Hypothetical serine/threonine-rich protein C... 31 3.4
TDH_THETN (Q8R7K0) L-threonine 3-dehydrogenase (EC 1.1.1.103) 31 3.4
RT31_HUMAN (Q92665) 28S ribosomal protein S31, mitochondrial pre... 31 3.4
HEM3_HELHP (Q7VFE9) Porphobilinogen deaminase (EC 2.5.1.61) (PBG... 31 3.4
NU1M_ACACA (Q37381) NADH-ubiquinone oxidoreductase chain 1 (EC 1... 30 5.8
CPHA_CYAA5 (Q9KGY4) Cyanophycin synthetase (EC 6.-.-.-) 30 5.8
YK02_SCHPO (Q9HDY9) Hypothetical serine/threonine-rich protein P... 30 7.6
SOL_DROME (P27398) Small optic lobes protein (EC 3.4.22.-) 30 7.6
GUNF_CLOCE (P37698) Endoglucanase F precursor (EC 3.2.1.4) (Endo... 30 7.6
Y535_CLOPE (Q8XN03) Hypothetical UPF0059 protein CPE0535 30 10.0
>AIRC_ARATH (Q94BT2) Auxin-induced in root cultures protein 12
precursor
Length = 252
Score = 43.5 bits (101), Expect = 7e-04
Identities = 31/111 (27%), Positives = 50/111 (44%), Gaps = 4/111 (3%)
Query: 1 MIGAQALVAIPQASGSPKAYTSNIVDTSTRLQEGTISYPVSGLSA-TYQNNKVTIFATLT 59
M G+QA +A G+ + + + + L EG +++ L A + ++ IF T+
Sbjct: 91 MAGSQAFLAYRSGGGAAPVVKTYNISSYSSLVEGKLAFDFWNLRAESLSGGRIAIFTTVK 150
Query: 60 LPNGTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASG 110
+P G S+ VWQ G ++ P H + S VL S T AA G
Sbjct: 151 VPAGADSVNQVWQIGGNVTNGRPGVHPFGPDNLGSHRVL---SFTEDAAPG 198
>YFGG_SCHPO (O13854) Hypothetical serine/threonine-rich protein
C19G12.16c in chromosome I precursor
Length = 670
Score = 31.2 bits (69), Expect = 3.4
Identities = 27/108 (25%), Positives = 45/108 (41%), Gaps = 2/108 (1%)
Query: 6 ALVAIPQASGSPKAYTSNIVDTSTRLQEGTISYPVSGLSATYQNNKVTIFATLTLPNGTT 65
++++ S S TS I TST + + S +S + +Y N T +T T + T
Sbjct: 232 SVISSASLSSSSVLPTSIITSTSTPVTVSSSS--LSSFTPSYSTNLTTTGSTTTTGSATV 289
Query: 66 SLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGS 113
S + + + S P S +S +S S TS +G G+
Sbjct: 290 SSSPFYSNSSVIPTSVPSSVSSFTSSSSSYTTTLTASNTSVTYTGTGT 337
>TDH_THETN (Q8R7K0) L-threonine 3-dehydrogenase (EC 1.1.1.103)
Length = 347
Score = 31.2 bits (69), Expect = 3.4
Identities = 29/107 (27%), Positives = 42/107 (39%), Gaps = 10/107 (9%)
Query: 50 NKVTIFATLTLPNGTTSLVHVWQDGVLSSDSTPQEHSHE--------SSHQNSKEVLDLV 101
++V I T GT ++VW + S P+ HE + S +V DLV
Sbjct: 30 DEVLIKVKATSICGTDVHIYVWNEWAKSRIKPPKTMGHEFVGEVVEIGENVTSVKVGDLV 89
Query: 102 SGTSQAASGIGSRQRRRNTHGVLNAISWGILMPTGAVIARYLKVFKS 148
S + G R N H N + G+ T A Y+KV +S
Sbjct: 90 SAETHIVCGKCRACRTGNAHICENTLILGV--DTDGAFAEYIKVPES 134
>RT31_HUMAN (Q92665) 28S ribosomal protein S31, mitochondrial
precursor (S31mt) (MRP-S31) (Imogen 38)
Length = 395
Score = 31.2 bits (69), Expect = 3.4
Identities = 26/104 (25%), Positives = 45/104 (43%), Gaps = 5/104 (4%)
Query: 11 PQASGSPKAYTSNIVDTSTRLQEGTISYPVSGLSATYQNNKVTIFATLTLPNGTTSLVHV 70
P +SGSP+ + I+ + R GT+ Y S L A +NN F T ++ V
Sbjct: 18 PLSSGSPETSAAAIMLLTVR--HGTVRYRSSALLARTKNNIQRYFGTNSVICSKKDKQSV 75
Query: 71 WQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGSR 114
+ + S Q+ E++ K++L ++ G S + R
Sbjct: 76 RTEEISKETSESQDSEKENT---KKDLLGIIKGMKVELSTVNVR 116
>HEM3_HELHP (Q7VFE9) Porphobilinogen deaminase (EC 2.5.1.61) (PBG)
(Hydroxymethylbilane synthase) (HMBS)
(Pre-uroporphyrinogen synthase)
Length = 321
Score = 31.2 bits (69), Expect = 3.4
Identities = 18/56 (32%), Positives = 29/56 (51%), Gaps = 1/56 (1%)
Query: 30 RLQEGTISYPVSGLSATYQNNKVTIFATLTLPNGTTSLVHVWQDGVLSSDSTPQEH 85
R EG P+ G+ A + NNK+T+ A + LP+G+ L +D + D E+
Sbjct: 240 RTLEGGCQVPI-GVYAQFSNNKLTLQAIVGLPDGSEVLQDSVEDSINIDDINASEN 294
>NU1M_ACACA (Q37381) NADH-ubiquinone oxidoreductase chain 1 (EC
1.6.5.3)
Length = 369
Score = 30.4 bits (67), Expect = 5.8
Identities = 22/72 (30%), Positives = 36/72 (49%), Gaps = 4/72 (5%)
Query: 185 ITYDTHRALAIVLVTL---ATLQVFALFLRPNKDHKLRFYWNIYHHV-VGYVTISISIVN 240
IT+ H + I+ V + A + V A R D +R W ++ + +G V ISIS+
Sbjct: 288 ITWIYHSLVYIIKVYIYLFAFIWVRATLPRYRYDQLMRLGWKVFVPLTIGLVFISISLTL 347
Query: 241 VFKGFEALGDFV 252
F GF + G ++
Sbjct: 348 FFDGFPSEGGYI 359
>CPHA_CYAA5 (Q9KGY4) Cyanophycin synthetase (EC 6.-.-.-)
Length = 872
Score = 30.4 bits (67), Expect = 5.8
Identities = 13/28 (46%), Positives = 19/28 (67%), Gaps = 4/28 (14%)
Query: 244 GFEALGDFVGDRYKNWKHAYIGIIGALG 271
G+EA+G+FV KNWK +G++G G
Sbjct: 752 GYEAVGEFV----KNWKGQRLGVVGGPG 775
>YK02_SCHPO (Q9HDY9) Hypothetical serine/threonine-rich protein
P11E10.02c in chromosome I precursor
Length = 1082
Score = 30.0 bits (66), Expect = 7.6
Identities = 29/122 (23%), Positives = 53/122 (42%), Gaps = 6/122 (4%)
Query: 9 AIPQASGSPKAYTSN-IVDTSTRLQEG-----TISYPVSGLSATYQNNKVTIFATLTLPN 62
A P SP + +S+ +V+TS+ L + T + S ++ T N+ + T LP+
Sbjct: 263 ATPSTLSSPASSSSSYLVETSSTLTDSVFTTVTATSDSSVITYTLINSVTSSSETTNLPS 322
Query: 63 GTTSLVHVWQDGVLSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASGIGSRQRRRNTHG 122
++SLV + + SS + S + S D ++ S +S + S T
Sbjct: 323 SSSSLVTIGESSFPSSLLSLLTQSFSTVRSTSSSSTDQLTSASPISSSVISPSVSSPTSS 382
Query: 123 VL 124
+L
Sbjct: 383 IL 384
>SOL_DROME (P27398) Small optic lobes protein (EC 3.4.22.-)
Length = 1594
Score = 30.0 bits (66), Expect = 7.6
Identities = 17/58 (29%), Positives = 30/58 (51%), Gaps = 6/58 (10%)
Query: 59 TLP-NGTTSLVHVWQDGV-----LSSDSTPQEHSHESSHQNSKEVLDLVSGTSQAASG 110
T+P NG V+ W + + +SS S H H +S+ NS ++++ S ++SG
Sbjct: 402 TIPRNGVFIAVNEWSEPMASSSSVSSSSNHHHHHHSNSNSNSSGNSNIINNNSSSSSG 459
>GUNF_CLOCE (P37698) Endoglucanase F precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase F) (Cellulase F) (EGCCF)
Length = 722
Score = 30.0 bits (66), Expect = 7.6
Identities = 21/76 (27%), Positives = 34/76 (44%), Gaps = 10/76 (13%)
Query: 88 ESSHQNSKEVLDLVSGTSQAASGIGSRQRRRNTHGVLN-----AISWGILMPTGAVIARY 142
E+S QN++++LD + + GI + ++R + H L+ W MP G VI
Sbjct: 544 ETSRQNAQKLLDAMWNNYSDSKGISTVEQRGDYHRFLDQEVFVPAGWTGKMPNGDVIKSG 603
Query: 143 LKVFK-----SADPAW 153
+K DP W
Sbjct: 604 VKFIDIRSKYKQDPEW 619
>Y535_CLOPE (Q8XN03) Hypothetical UPF0059 protein CPE0535
Length = 213
Score = 29.6 bits (65), Expect = 10.0
Identities = 16/51 (31%), Positives = 28/51 (54%), Gaps = 5/51 (9%)
Query: 228 VVGYVTISISIVNVFKGFEALGDFVGDRYKNWKHAYIGIIGALGGIAVLLE 278
++G +T + + V +G+ +GD +KN+ G+I L GI +LLE
Sbjct: 158 IIGIITFVLCFLGVI-----IGEKLGDIFKNYAEIVGGVILILIGINILLE 203
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 36,812,482
Number of Sequences: 164201
Number of extensions: 1536724
Number of successful extensions: 3901
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 3897
Number of HSP's gapped (non-prelim): 12
length of query: 310
length of database: 59,974,054
effective HSP length: 110
effective length of query: 200
effective length of database: 41,911,944
effective search space: 8382388800
effective search space used: 8382388800
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC147406.4


