

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147404.5 + phase: 0
(498 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
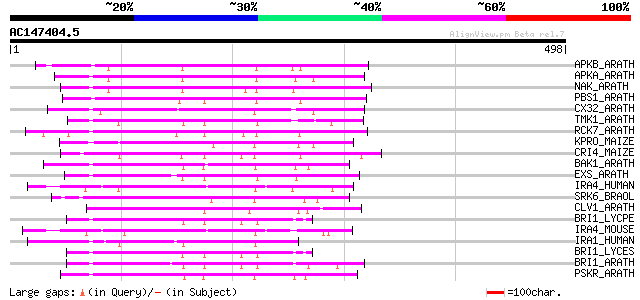
Score E
Sequences producing significant alignments: (bits) Value
APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor ... 161 4e-39
APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor ... 155 2e-37
NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK ... 151 3e-36
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 141 3e-33
CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx3... 138 4e-32
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 136 1e-31
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 134 5e-31
KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1 precu... 122 2e-27
CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4 pr... 121 4e-27
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 119 1e-26
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 119 2e-26
IRA4_HUMAN (Q9NWZ3) Interleukin-1 receptor-associated kinase-4 (... 113 1e-24
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 113 1e-24
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 107 7e-23
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 107 7e-23
IRA4_MOUSE (Q8R4K2) Interleukin-1 receptor-associated kinase-4 (... 107 9e-23
IRA1_HUMAN (P51617) Interleukin-1 receptor-associated kinase 1 (... 107 9e-23
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 106 1e-22
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 106 1e-22
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 97 7e-20
>APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor (EC
2.7.1.-)
Length = 412
Score = 161 bits (407), Expect = 4e-39
Identities = 112/330 (33%), Positives = 173/330 (51%), Gaps = 38/330 (11%)
Query: 24 SKKTDPGDDGGDGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLE 83
S +T+P +G + Q P K + EL+ AT F + ++ E G + V+KG ++
Sbjct: 37 SIRTNPRTEG----EILQSPNLKSFTFAELKAATRNFRPDSVLGEGGFGS---VFKGWID 89
Query: 84 NNRL----------VAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERL 133
L +AVK+ ++ W Q++LAE +G+ H +V LIG C E + RL
Sbjct: 90 EQTLTASKPGTGVVIAVKKLNQDGWQGHQEWLAEVNYLGQFSHPNLVKLIGYCLEDEHRL 149
Query: 134 LVAEYMPNDTLSKHLFHWDK--QPLPWEMRVRVAYHVAQALDHC-SMENRKIYHDLNAYR 190
LV E+MP +L HLF QPL W +R++VA A+ L + E IY D
Sbjct: 150 LVYEFMPRGSLENHLFRRGSYFQPLSWTLRLKVALGAAKGLAFLHNAETSVIYRDFKTSN 209
Query: 191 ILFDEDGDPRLSSFGLMKNSRDG-KSY-STNL----AYTPPEFLRTGRIIAESVIYSYGT 244
IL D + + +LS FGL K+ G KS+ ST + Y PE+L TG + +S +YSYG
Sbjct: 210 ILLDSEYNAKLSDFGLAKDGPTGDKSHVSTRIMGTYGYAAPEYLATGHLTTKSDVYSYGV 269
Query: 245 VLLDLLSG-----KHIPPSH------ALDLIRGKNALL-LMDSSLEGQYANDDATKLVEL 292
VLL++LSG K+ PP A L+ K L ++D+ L+ QY+ ++A K+ L
Sbjct: 270 VLLEVLSGRRAVDKNRPPGEQKLVEWARPLLANKRKLFRVIDNRLQDQYSMEEACKVATL 329
Query: 293 ASKCLQFEARERPDIKFLLTAVTPLQKQKE 322
A +CL FE + RP++ +++ + +Q E
Sbjct: 330 ALRCLTFEIKLRPNMNEVVSHLEHIQTLNE 359
>APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor (EC
2.7.1.-)
Length = 410
Score = 155 bits (393), Expect = 2e-37
Identities = 104/309 (33%), Positives = 163/309 (52%), Gaps = 34/309 (11%)
Query: 41 QVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRL----------VAV 90
Q P K + EL+ AT F + ++ E G V+KG ++ L +AV
Sbjct: 49 QSPNLKSFSFAELKSATRNFRPDSVLGEGGF---GCVFKGWIDEKSLTASRPGTGLVIAV 105
Query: 91 KRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFH 150
K+ ++ W Q++LAE +G+ H+ +V LIG C E + RLLV E+MP +L HLF
Sbjct: 106 KKLNQDGWQGHQEWLAEVNYLGQFSHRHLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFR 165
Query: 151 WDK--QPLPWEMRVRVAYHVAQALDHC-SMENRKIYHDLNAYRILFDEDGDPRLSSFGLM 207
QPL W++R++VA A+ L S E R IY D IL D + + +LS FGL
Sbjct: 166 RGLYFQPLSWKLRLKVALGAAKGLAFLHSSETRVIYRDFKTSNILLDSEYNAKLSDFGLA 225
Query: 208 KNSRDG-KSY-STNL----AYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHI----PP 257
K+ G KS+ ST + Y PE+L TG + +S +YS+G VLL+LLSG+ P
Sbjct: 226 KDGPIGDKSHVSTRVMGTHGYAAPEYLATGHLTTKSDVYSFGVVLLELLSGRRAVDKNRP 285
Query: 258 SHALDLIRGKNALL--------LMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKF 309
S +L+ L ++D+ L+ QY+ ++A K+ L+ +CL E + RP++
Sbjct: 286 SGERNLVEWAKPYLVNKRKIFRVIDNRLQDQYSMEEACKVATLSLRCLTTEIKLRPNMSE 345
Query: 310 LLTAVTPLQ 318
+++ + +Q
Sbjct: 346 VVSHLEHIQ 354
>NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK (EC
2.7.1.37)
Length = 389
Score = 151 bits (382), Expect = 3e-36
Identities = 104/310 (33%), Positives = 159/310 (50%), Gaps = 34/310 (10%)
Query: 46 KEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRL----------VAVKRFSK 95
K + L+EL+ AT F + +V E G V+KG ++ + L +AVKR ++
Sbjct: 54 KNFSLSELKSATRNFRPDSVVGEGGF---GCVFKGWIDESSLAPSKPGTGIVIAVKRLNQ 110
Query: 96 QSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDK-- 153
+ + +++LAE +G++ H +V LIG C E + RLLV E+M +L HLF
Sbjct: 111 EGFQGHREWLAEINYLGQLDHPNLVKLIGYCLEEEHRLLVYEFMTRGSLENHLFRRGTFY 170
Query: 154 QPLPWEMRVRVAYHVAQALDHC-SMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKNS-- 210
QPL W RVR+A A+ L + + + IY D A IL D + + +LS FGL ++
Sbjct: 171 QPLSWNTRVRMALGAARGLAFLHNAQPQVIYRDFKASNILLDSNYNAKLSDFGLARDGPM 230
Query: 211 RDGKSYSTNL----AYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGK-----------HI 255
D ST + Y PE+L TG + +S +YS+G VLL+LLSG+ H
Sbjct: 231 GDNSHVSTRVMGTQGYAAPEYLATGHLSVKSDVYSFGVVLLELLSGRRAIDKNQPVGEHN 290
Query: 256 PPSHALDLIRGKNALL-LMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTAV 314
A + K LL +MD L+GQY+ A K+ LA C+ +A+ RP + ++ +
Sbjct: 291 LVDWARPYLTNKRRLLRVMDPRLQGQYSLTRALKIAVLALDCISIDAKSRPTMNEIVKTM 350
Query: 315 TPLQKQKEVA 324
L QKE +
Sbjct: 351 EELHIQKEAS 360
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 141 bits (356), Expect = 3e-33
Identities = 100/296 (33%), Positives = 153/296 (50%), Gaps = 26/296 (8%)
Query: 48 YGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENN-RLVAVKRFSKQSWPDAQQFLA 106
+ EL AT F + + E G VYKG+L++ ++VAVK+ + ++FL
Sbjct: 74 FAFRELAAATMNFHPDTFLGEGGFGR---VYKGRLDSTGQVVAVKQLDRNGLQGNREFLV 130
Query: 107 EAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHW--DKQPLPWEMRVRV 164
E + + H +VNLIG CA+GD+RLLV E+MP +L HL DK+ L W MR+++
Sbjct: 131 EVLMLSLLHHPNLVNLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPPDKEALDWNMRMKI 190
Query: 165 AYHVAQALD--HCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKNSRDG-KSY-STNL 220
A A+ L+ H IY D + IL DE P+LS FGL K G KS+ ST +
Sbjct: 191 AAGAAKGLEFLHDKANPPVIYRDFKSSNILLDEGFHPKLSDFGLAKLGPTGDKSHVSTRV 250
Query: 221 ----AYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSH-----------ALDLIR 265
Y PE+ TG++ +S +YS+G V L+L++G+ S A L
Sbjct: 251 MGTYGYCAPEYAMTGQLTVKSDVYSFGVVFLELITGRKAIDSEMPHGEQNLVAWARPLFN 310
Query: 266 GKNALL-LMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTAVTPLQKQ 320
+ + L D L+G++ + + +AS C+Q +A RP I ++TA++ L Q
Sbjct: 311 DRRKFIKLADPRLKGRFPTRALYQALAVASMCIQEQAATRPLIADVVTALSYLANQ 366
>CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx32,
chloroplast precursor (EC 2.7.1.37)
Length = 419
Score = 138 bits (347), Expect = 4e-32
Identities = 88/316 (27%), Positives = 156/316 (48%), Gaps = 39/316 (12%)
Query: 35 DGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKG----------KLEN 84
D + + P K Y +L+ AT F + ++ + G VY+G ++ +
Sbjct: 61 DSGKLLESPNLKVYNFLDLKTATKNFKPDSMLGQGGF---GKVYRGWVDATTLAPSRVGS 117
Query: 85 NRLVAVKRFSKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTL 144
+VA+KR + +S ++ +E +G + H+ +V L+G C E E LLV E+MP +L
Sbjct: 118 GMIVAIKRLNSESVQGFAEWRSEVNFLGMLSHRNLVKLLGYCREDKELLLVYEFMPKGSL 177
Query: 145 SKHLFHWDKQPLPWEMRVRVAYHVAQALDHC-SMENRKIYHDLNAYRILFDEDGDPRLSS 203
HLF P PW++R+++ A+ L S++ IY D A IL D + D +LS
Sbjct: 178 ESHLFR-RNDPFPWDLRIKIVIGAARGLAFLHSLQREVIYRDFKASNILLDSNYDAKLSD 236
Query: 204 FGLMK-NSRDGKSYST-----NLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPP 257
FGL K D KS+ T Y PE++ TG + +S ++++G VLL++++G
Sbjct: 237 FGLAKLGPADEKSHVTTRIMGTYGYAAPEYMATGHLYVKSDVFAFGVVLLEIMTGL---T 293
Query: 258 SHALDLIRGKNALL---------------LMDSSLEGQYANDDATKLVELASKCLQFEAR 302
+H RG+ +L+ +MD ++GQY AT++ + C++ + +
Sbjct: 294 AHNTKRPRGQESLVDWLRPELSNKHRVKQIMDKGIKGQYTTKVATEMARITLSCIEPDPK 353
Query: 303 ERPDIKFLLTAVTPLQ 318
RP +K ++ + +Q
Sbjct: 354 NRPHMKEVVEVLEHIQ 369
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 136 bits (343), Expect = 1e-31
Identities = 94/297 (31%), Positives = 156/297 (51%), Gaps = 41/297 (13%)
Query: 53 LRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQ--SWPDAQQFLAEAAG 110
LR T+ FS++ I+ G VVYKG+L + +AVKR + +F +E A
Sbjct: 581 LRSVTNNFSSDNILGSGGF---GVVYKGELHDGTKIAVKRMENGVIAGKGFAEFKSEIAV 637
Query: 111 VGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDKQ---PLPWEMRVRVAYH 167
+ KVRH+ +V L+G C +G+E+LLV EYMP TLS+HLF W ++ PL W+ R+ +A
Sbjct: 638 LTKVRHRHLVTLLGYCLDGNEKLLVYEYMPQGTLSRHLFEWSEEGLKPLLWKQRLTLALD 697
Query: 168 VAQALD--HCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKNSRDGK-SYSTNLA--- 221
VA+ ++ H I+ DL IL +D +++ FGL++ + +GK S T +A
Sbjct: 698 VARGVEYLHGLAHQSFIHRDLKPSNILLGDDMRAKVADFGLVRLAPEGKGSIETRIAGTF 757
Query: 222 -YTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALDLIRGKNALLLMDSSLEGQ 280
Y PE+ TGR+ + +YS+G +L++L++G+ +LD + + ++ L+ S +
Sbjct: 758 GYLAPEYAVTGRVTTKVDVYSFGVILMELITGR-----KSLDESQPEESIHLV-SWFKRM 811
Query: 281 YANDDAT--------------------KLVELASKCLQFEARERPDIKFLLTAVTPL 317
Y N +A+ + ELA C E +RPD+ + ++ L
Sbjct: 812 YINKEASFKKAIDTTIDLDEETLASVHTVAELAGHCCAREPYQRPDMGHAVNILSSL 868
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 134 bits (337), Expect = 5e-31
Identities = 106/348 (30%), Positives = 164/348 (46%), Gaps = 44/348 (12%)
Query: 15 EDPPTALPDSKKTDP----GDDGGDGDQESQVPVFKEYGLN--------------ELRRA 56
+D DS K P ++GG G ++ K LN EL A
Sbjct: 40 DDQTQPSSDSTKVSPYRDVNNEGGVGKEDQLSLDVKGLNLNDQVTGKKAQTFTFQELAEA 99
Query: 57 THEFSTEYIVSESGEKAPNVVYKGKLEN-NRLVAVKRFSKQSWPDAQQFLAEAAGVGKVR 115
T F ++ + E G V+KG +E +++VA+K+ + ++F+ E +
Sbjct: 100 TGNFRSDCFLGEGGF---GKVFKGTIEKLDQVVAIKQLDRNGVQGIREFVVEVLTLSLAD 156
Query: 116 HKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHL--FHWDKQPLPWEMRVRVAYHVAQALD 173
H +V LIG CAEGD+RLLV EYMP +L HL K+PL W R+++A A+ L+
Sbjct: 157 HPNLVKLIGFCAEGDQRLLVYEYMPQGSLEDHLHVLPSGKKPLDWNTRMKIAAGAARGLE 216
Query: 174 --HCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMK--NSRDGKSYSTNL----AYTPP 225
H M IY DL IL ED P+LS FGL K S D ST + Y P
Sbjct: 217 YLHDRMTPPVIYRDLKCSNILLGEDYQPKLSDFGLAKVGPSGDKTHVSTRVMGTYGYCAP 276
Query: 226 EFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPS-----------HALDLIRG-KNALLLM 273
++ TG++ +S IYS+G VLL+L++G+ + A L + +N ++
Sbjct: 277 DYAMTGQLTFKSDIYSFGVVLLELITGRKAIDNTKTRKDQNLVGWARPLFKDRRNFPKMV 336
Query: 274 DSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTAVTPLQKQK 321
D L+GQY + + +++ C+Q + RP + ++ A+ L K
Sbjct: 337 DPLLQGQYPVRGLYQALAISAMCVQEQPTMRPVVSDVVLALNFLASSK 384
>KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1
precursor (EC 2.7.1.37)
Length = 817
Score = 122 bits (306), Expect = 2e-27
Identities = 95/292 (32%), Positives = 143/292 (48%), Gaps = 34/292 (11%)
Query: 45 FKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQF 104
F+ Y EL +AT +F E ESG VYKG LE++R VAVK+ + F
Sbjct: 521 FRRYSYRELVKATRKFKVELGRGESG-----TVYKGVLEDDRHVAVKKLENVR-QGKEVF 574
Query: 105 LAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLF-HWDKQPLPWEMRVR 163
AE + +G++ H +V + G C+EG RLLV+EY+ N +L+ LF L WE R
Sbjct: 575 QAELSVIGRINHMNLVRIWGFCSEGSHRLLVSEYVENGSLANILFSEGGNILLDWEGRFN 634
Query: 164 VAYHVAQALDHCSMENRK--IYHDLNAYRILFDEDGDPRLSSFGLMKNSRDGKSYST--- 218
+A VA+ L + E + I+ D+ IL D+ +P+++ FGL+K G S
Sbjct: 635 IALGVAKGLAYLHHECLEWVIHCDVKPENILLDQAFEPKITDFGLVKLLNRGGSTQNVSH 694
Query: 219 ---NLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPP---------SHALDLIRG 266
L Y PE++ + I A+ +YSYG VLL+LL+G + S L+R
Sbjct: 695 VRGTLGYIAPEWVSSLPITAKVDVYSYGVVLLELLTGTRVSELVGGTDEVHSMLRKLVRM 754
Query: 267 KNALL----------LMDSSLEGQYANDDATKLVELASKCLQFEARERPDIK 308
+A L +DS L A L++LA CL+ + +RP ++
Sbjct: 755 LSAKLEGEEQSWIDGYLDSKLNRPVNYVQARTLIKLAVSCLEEDRSKRPTME 806
>CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4
precursor (EC 2.7.1.-)
Length = 901
Score = 121 bits (304), Expect = 4e-27
Identities = 92/319 (28%), Positives = 161/319 (49%), Gaps = 34/319 (10%)
Query: 46 KEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQS--WPDAQQ 103
+E+ EL +AT FS + S+ G+ + + V+KG L + +VAVKR K S +++
Sbjct: 491 QEFSYEELEQATGGFSED---SQVGKGSFSCVFKGILRDGTVVAVKRAIKASDVKKSSKE 547
Query: 104 FLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWD---KQPLPWEM 160
F E + ++ H ++NL+G C +G ERLLV E+M + +L +HL D K+ L W
Sbjct: 548 FHNELDLLSRLNHAHLLNLLGYCEDGSERLLVYEFMAHGSLYQHLHGKDPNLKKRLNWAR 607
Query: 161 RVRVAYHVAQALD--HCSMENRKIYHDLNAYRILFDEDGDPRLSSFGL--MKNSRDGKSY 216
RV +A A+ ++ H I+ D+ + IL DED + R++ FGL + + G
Sbjct: 608 RVTIAVQAARGIEYLHGYACPPVIHRDIKSSNILIDEDHNARVADFGLSILGPADSGTPL 667
Query: 217 ST----NLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSH---------ALDL 263
S L Y PE+ R + +S +YS+G VLL++LSG+ A+ L
Sbjct: 668 SELPAGTLGYLDPEYYRLHYLTTKSDVYSFGVVLLEILSGRKAIDMQFEEGNIVEWAVPL 727
Query: 264 IRGKNALLLMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTAV--------- 314
I+ + ++D L + K+ +A KC++ ++RP + + TA+
Sbjct: 728 IKAGDIFAILDPVLSPPSDLEALKKIASVACKCVRMRGKDRPSMDKVTTALEHALALLMG 787
Query: 315 TPLQKQKEVASHVLMGLTK 333
+P +Q + + V++G ++
Sbjct: 788 SPCIEQPILPTEVVLGSSR 806
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 119 bits (299), Expect = 1e-26
Identities = 89/300 (29%), Positives = 144/300 (47%), Gaps = 30/300 (10%)
Query: 31 DDGGDGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAV 90
D + D E + K + L EL+ A+ FS + I+ G VYKG+L + LVAV
Sbjct: 260 DVPAEEDPEVHLGQLKRFSLRELQVASDNFSNKNILGRGGF---GKVYKGRLADGTLVAV 316
Query: 91 KRFSKQSWPDAQ-QFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLF 149
KR ++ + QF E + H+ ++ L G C ERLLV YM N +++ L
Sbjct: 317 KRLKEERTQGGELQFQTEVEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLR 376
Query: 150 HWDKQ--PLPWEMRVRVAYHVAQAL----DHCSMENRKIYHDLNAYRILFDEDGDPRLSS 203
+ PL W R R+A A+ L DHC + + I+ D+ A IL DE+ + +
Sbjct: 377 ERPESQPPLDWPKRQRIALGSARGLAYLHDHC--DPKIIHRDVKAANILLDEEFEAVVGD 434
Query: 204 FGLMKNSRDGKSYSTN-----LAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHI--- 255
FGL K ++ T + + PE+L TG+ ++ ++ YG +LL+L++G+
Sbjct: 435 FGLAKLMDYKDTHVTTAVRGTIGHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDL 494
Query: 256 ------PPSHALDLIRG----KNALLLMDSSLEGQYANDDATKLVELASKCLQFEARERP 305
LD ++G K L+D L+G Y +++ +L+++A C Q ERP
Sbjct: 495 ARLANDDDVMLLDWVKGLLKEKKLEALVDVDLQGNYKDEEVEQLIQVALLCTQSSPMERP 554
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 119 bits (297), Expect = 2e-26
Identities = 86/290 (29%), Positives = 146/290 (49%), Gaps = 32/290 (11%)
Query: 50 LNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQFLAEAA 109
L ++ AT FS + I+ + G VYK L + VAVK+ S+ ++F+AE
Sbjct: 907 LGDIVEATDHFSKKNIIGDGGF---GTVYKACLPGEKTVAVKKLSEAKTQGNREFMAEME 963
Query: 110 GVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDK------QPLPWEMRVR 163
+GKV+H +V+L+G C+ +E+LLV EYM N +L HW + + L W R++
Sbjct: 964 TLGKVKHPNLVSLLGYCSFSEEKLLVYEYMVNGSLD----HWLRNQTGMLEVLDWSKRLK 1019
Query: 164 VAYHVAQALD--HCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKNSRDGKSY-STNL 220
+A A+ L H I+ D+ A IL D D +P+++ FGL + +S+ ST +
Sbjct: 1020 IAVGAARGLAFLHHGFIPHIIHRDIKASNILLDGDFEPKVADFGLARLISACESHVSTVI 1079
Query: 221 A----YTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIP------------PSHALDLI 264
A Y PPE+ ++ R + +YS+G +LL+L++GK A+ I
Sbjct: 1080 AGTFGYIPPEYGQSARATTKGDVYSFGVILLELVTGKEPTGPDFKESEGGNLVGWAIQKI 1139
Query: 265 RGKNALLLMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLTAV 314
A+ ++D L + +L+++A CL +RP++ +L A+
Sbjct: 1140 NQGKAVDVIDPLLVSVALKNSQLRLLQIAMLCLAETPAKRPNMLDVLKAL 1189
>IRA4_HUMAN (Q9NWZ3) Interleukin-1 receptor-associated kinase-4 (EC
2.7.1.37) (IRAK-4) (NY-REN-64 antigen)
Length = 460
Score = 113 bits (283), Expect = 1e-24
Identities = 96/315 (30%), Positives = 155/315 (48%), Gaps = 37/315 (11%)
Query: 17 PPTALPDSKKTDPGDDGGDGDQESQVPVFKEYGLNELRRATHEFSTEYIV---SESGEKA 73
P ++ P++K + D F + EL+ T+ F I ++ GE
Sbjct: 148 PDSSSPENKSLEVSDTR-----------FHSFSFYELKNVTNNFDERPISVGGNKMGEGG 196
Query: 74 PNVVYKGKLENNRLVAVKRFSKQ----SWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEG 129
VVYKG + NN VAVK+ + + QQF E + K +H+ +V L+G ++G
Sbjct: 197 FGVVYKGYV-NNTTVAVKKLAAMVDITTEELKQQFDQEIKVMAKCQHENLVELLGFSSDG 255
Query: 130 DERLLVAEYMPNDTLSKHLFHWD-KQPLPWEMRVRVAYHVAQALDHCSMENRKIYHDLNA 188
D+ LV YMPN +L L D PL W MR ++A A ++ EN I+ D+ +
Sbjct: 256 DDLCLVYVYMPNGSLLDRLSCLDGTPPLSWHMRCKIAQGAANGINFLH-ENHHIHRDIKS 314
Query: 189 YRILFDEDGDPRLSSFGLMKNS-RDGKSYSTN-----LAYTPPEFLRTGRIIAESVIYSY 242
IL DE ++S FGL + S + ++ T+ AY PE LR G I +S IYS+
Sbjct: 315 ANILLDEAFTAKISDFGLARASEKFAQTVMTSRIVGTTAYMAPEALR-GEITPKSDIYSF 373
Query: 243 GTVLLDLLSG-----KHIPPSHALDLIRG-KNALLLMDSSLEGQYANDDATK---LVELA 293
G VLL++++G +H P LD+ ++ ++ ++ + + D+T + +A
Sbjct: 374 GVVLLEIITGLPAVDEHREPQLLLDIKEEIEDEEKTIEDYIDKKMNDADSTSVEAMYSVA 433
Query: 294 SKCLQFEARERPDIK 308
S+CL + +RPDIK
Sbjct: 434 SQCLHEKKNKRPDIK 448
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase
receptor precursor (EC 2.7.1.37) (S-receptor kinase)
(SRK)
Length = 849
Score = 113 bits (282), Expect = 1e-24
Identities = 81/295 (27%), Positives = 147/295 (49%), Gaps = 33/295 (11%)
Query: 38 QESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQS 97
+E ++P+ + + + +AT FS+ ++ G+ +VYKG+L + + +AVKR SK S
Sbjct: 509 EELELPLIE---METVVKATENFSS---CNKLGQGGFGIVYKGRLLDGKEIAVKRLSKTS 562
Query: 98 WPDAQQFLAEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLF-HWDKQPL 156
+F+ E + +++H +V ++GCC EGDE++L+ EY+ N +L +LF + L
Sbjct: 563 VQGTDEFMNEVTLIARLQHINLVQVLGCCIEGDEKMLIYEYLENLSLDSYLFGKTRRSKL 622
Query: 157 PWEMRVRVAYHVAQALDHCSMEN--RKIYHDLNAYRILFDEDGDPRLSSFGLMK-NSRDG 213
W R + VA+ L + ++ R I+ DL IL D++ P++S FG+ + RD
Sbjct: 623 NWNERFDITNGVARGLLYLHQDSRFRIIHRDLKVSNILLDKNMIPKISDFGMARIFERDE 682
Query: 214 KSYST-----NLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALDL----- 263
+T Y PE+ G +S ++S+G ++L+++SGK + LD
Sbjct: 683 TEANTMKVVGTYGYMSPEYAMYGIFSEKSDVFSFGVIVLEIVSGKKNRGFYNLDYENDLL 742
Query: 264 ------IRGKNALLLMD-------SSLEGQYANDDATKLVELASKCLQFEARERP 305
+ AL ++D SS + + K +++ C+Q A RP
Sbjct: 743 SYVWSRWKEGRALEIVDPVIVDSLSSQPSIFQPQEVLKCIQIGLLCVQELAEHRP 797
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 107 bits (267), Expect = 7e-23
Identities = 79/271 (29%), Positives = 130/271 (47%), Gaps = 26/271 (9%)
Query: 70 GEKAPNVVYKGKLENNRLVAVKRF-SKQSWPDAQQFLAEAAGVGKVRHKRMVNLIGCCAE 128
G+ +VY+G + NN VA+KR + + F AE +G++RH+ +V L+G A
Sbjct: 699 GKGGAGIVYRGSMPNNVDVAIKRLVGRGTGRSDHGFTAEIQTLGRIRHRHIVRLLGYVAN 758
Query: 129 GDERLLVAEYMPNDTLSKHLFHWDKQPLPWEMRVRVAYHVAQALD--HCSMENRKIYHDL 186
D LL+ EYMPN +L + L L WE R RVA A+ L H ++ D+
Sbjct: 759 KDTNLLLYEYMPNGSLGELLHGSKGGHLQWETRHRVAVEAAKGLCYLHHDCSPLILHRDV 818
Query: 187 NAYRILFDEDGDPRLSSFGLMKNSRDG------KSYSTNLAYTPPEFLRTGRIIAESVIY 240
+ IL D D + ++ FGL K DG S + + Y PE+ T ++ +S +Y
Sbjct: 819 KSNNILLDSDFEAHVADFGLAKFLVDGAASECMSSIAGSYGYIAPEYAYTLKVDEKSDVY 878
Query: 241 SYGTVLLDLLSGKHIPP--SHALDLIR--------------GKNALLLMDSSLEGQYAND 284
S+G VLL+L++GK +D++R + ++D L G Y
Sbjct: 879 SFGVVLLELIAGKKPVGEFGEGVDIVRWVRNTEEEITQPSDAAIVVAIVDPRLTG-YPLT 937
Query: 285 DATKLVELASKCLQFEARERPDIKFLLTAVT 315
+ ++A C++ EA RP ++ ++ +T
Sbjct: 938 SVIHVFKIAMMCVEEEAAARPTMREVVHMLT 968
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 107 bits (267), Expect = 7e-23
Identities = 76/230 (33%), Positives = 122/230 (53%), Gaps = 17/230 (7%)
Query: 52 ELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGV 111
+L AT+ F + +V G VYK +L++ +VA+K+ S ++F AE +
Sbjct: 880 DLLEATNGFHNDSLVGSGGF---GDVYKAQLKDGSVVAIKKLIHVSGQGDREFTAEMETI 936
Query: 112 GKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDKQ--PLPWEMRVRVAYHVA 169
GK++H+ +V L+G C G+ERLLV EYM +L L K L W R ++A A
Sbjct: 937 GKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKTGIKLNWPARRKIAIGAA 996
Query: 170 QALD--HCSMENRKIYHDLNAYRILFDEDGDPRLSSFGL--MKNSRDGKSYSTNLA---- 221
+ L H + I+ D+ + +L DE+ + R+S FG+ + ++ D + LA
Sbjct: 997 RGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGTPG 1056
Query: 222 YTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALDLIRGKNALL 271
Y PPE+ ++ R + +YSYG VLL+LL+GK P+ + D G N L+
Sbjct: 1057 YVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQ--PTDSADF--GDNNLV 1102
>IRA4_MOUSE (Q8R4K2) Interleukin-1 receptor-associated kinase-4 (EC
2.7.1.37) (IRAK-4)
Length = 459
Score = 107 bits (266), Expect = 9e-23
Identities = 97/324 (29%), Positives = 147/324 (44%), Gaps = 47/324 (14%)
Query: 12 HSPEDPPTALPDSKKTDPGDDGGDGDQESQVPVFKEYGLNELRRATHEFSTEYIVS---E 68
HS E P ++ PD++ + D F + +EL+ T+ F + +
Sbjct: 143 HSCEPPDSSSPDNRSVESSDTR-----------FHSFSFHELKSITNNFDEQPASAGGNR 191
Query: 69 SGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDA----QQFLAEAAGVGKVRHKRMVNLIG 124
GE VVYKG + NN +VAVK+ QQF E + +H+ +V L+G
Sbjct: 192 MGEGGFGVVYKGCV-NNTIVAVKKLGAMVEISTEELKQQFDQEIKVMATCQHENLVELLG 250
Query: 125 CCAEGDERLLVAEYMPNDTLSKHLFHWD-KQPLPWEMRVRVAYHVAQALDHCSMENRKIY 183
++ D LV YMPN +L L D PL W R +VA A + EN I+
Sbjct: 251 FSSDSDNLCLVYAYMPNGSLLDRLSCLDGTPPLSWHTRCKVAQGTANGIRFLH-ENHHIH 309
Query: 184 HDLNAYRILFDEDGDPRLSSFGLMK-NSRDGKSYSTN-----LAYTPPEFLRTGRIIAES 237
D+ + IL D+D ++S FGL + ++R ++ T+ AY PE LR G I +S
Sbjct: 310 RDIKSANILLDKDFTAKISDFGLARASARLAQTVMTSRIVGTTAYMAPEALR-GEITPKS 368
Query: 238 VIYSYGTVLLDLLSGKHIPPSHALDLIRGKNALLLMDSSLEGQ------YAND------- 284
IYS+G VLL+L++G A+D R LL + +E + Y ++
Sbjct: 369 DIYSFGVVLLELITG-----LAAVDENREPQLLLDIKEEIEDEEKTIEDYTDEKMSDADP 423
Query: 285 -DATKLVELASKCLQFEARERPDI 307
+ AS+CL + RPDI
Sbjct: 424 ASVEAMYSAASQCLHEKKNRRPDI 447
>IRA1_HUMAN (P51617) Interleukin-1 receptor-associated kinase 1 (EC
2.7.1.37) (IRAK-1)
Length = 712
Score = 107 bits (266), Expect = 9e-23
Identities = 79/264 (29%), Positives = 124/264 (46%), Gaps = 26/264 (9%)
Query: 17 PPTALPDSKKTDPGDDGGDGDQESQVPVFKEYGLNELRRATHEFSTEYIVSESGEKAPNV 76
PP P T PG + + P + L E+ R TH FS E + E G
Sbjct: 169 PPPPSPAPSSTKPGPESSVSLLQGARPFPFCWPLCEISRGTHNFSEELKIGEGGF---GC 225
Query: 77 VYKGKLENNRLVAVKRFSKQS---WPDAQQ-FLAEAAGVGKVRHKRMVNLIGCCAEGDER 132
VY+ + N + AVKR + + W +Q FL E + + RH +V+ G CA+
Sbjct: 226 VYRAVMRNT-VYAVKRLKENADLEWTAVKQSFLTEVEQLSRFRHPNIVDFAGYCAQNGFY 284
Query: 133 LLVAEYMPNDTLSKHLFHWDKQ---PLPWEMRVRVAYHVAQALDHCSMENRKIYH-DLNA 188
LV ++PN +L L H Q PL W R+ + A+A+ ++ + H D+ +
Sbjct: 285 CLVYGFLPNGSLEDRL-HCQTQACPPLSWPQRLDILLGTARAIQFLHQDSPSLIHGDIKS 343
Query: 189 YRILFDEDGDPRLSSFGLMKNSR-DGKSYSTN------------LAYTPPEFLRTGRIIA 235
+L DE P+L FGL + SR G S S + LAY P E+++TGR+
Sbjct: 344 SNVLLDERLTPKLGDFGLARFSRFAGSSPSQSSMVARTQTVRGTLAYLPEEYIKTGRLAV 403
Query: 236 ESVIYSYGTVLLDLLSGKHIPPSH 259
++ +S+G V+L+ L+G+ +H
Sbjct: 404 DTDTFSFGVVVLETLAGQRAVKTH 427
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 106 bits (265), Expect = 1e-22
Identities = 76/230 (33%), Positives = 122/230 (53%), Gaps = 17/230 (7%)
Query: 52 ELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGV 111
+L AT+ F + +V G VYK +L++ +VA+K+ S ++F AE +
Sbjct: 880 DLLEATNGFHNDSLVGSGGF---GDVYKAQLKDGSVVAIKKLIHVSGQGDREFTAEMETI 936
Query: 112 GKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDK--QPLPWEMRVRVAYHVA 169
GK++H+ +V L+G C G+ERLLV EYM +L L K L W R ++A A
Sbjct: 937 GKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKIGIKLNWPARRKIAIGAA 996
Query: 170 QALD--HCSMENRKIYHDLNAYRILFDEDGDPRLSSFGL--MKNSRDGKSYSTNLA---- 221
+ L H + I+ D+ + +L DE+ + R+S FG+ + ++ D + LA
Sbjct: 997 RGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGTPG 1056
Query: 222 YTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALDLIRGKNALL 271
Y PPE+ ++ R + +YSYG VLL+LL+GK P+ + D G N L+
Sbjct: 1057 YVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQ--PTDSADF--GDNNLV 1102
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 106 bits (265), Expect = 1e-22
Identities = 84/291 (28%), Positives = 144/291 (48%), Gaps = 30/291 (10%)
Query: 52 ELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQFLAEAAGV 111
+L +AT+ F + ++ G VYK L++ VA+K+ S ++F+AE +
Sbjct: 875 DLLQATNGFHNDSLIGSGGF---GDVYKAILKDGSAVAIKKLIHVSGQGDREFMAEMETI 931
Query: 112 GKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDKQ---PLPWEMRVRVAYHV 168
GK++H+ +V L+G C GDERLLV E+M +L + + H K+ L W R ++A
Sbjct: 932 GKIKHRNLVPLLGYCKVGDERLLVYEFMKYGSL-EDVLHDPKKAGVKLNWSTRRKIAIGS 990
Query: 169 AQALD--HCSMENRKIYHDLNAYRILFDEDGDPRLSSFGL--MKNSRDGKSYSTNLA--- 221
A+ L H + I+ D+ + +L DE+ + R+S FG+ + ++ D + LA
Sbjct: 991 ARGLAFLHHNCSPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLAGTP 1050
Query: 222 -YTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKHIPPSHALDLIRG-------KNALLLM 273
Y PPE+ ++ R + +YSYG VLL+LL+GK P+ + D ++A L +
Sbjct: 1051 GYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKR--PTDSPDFGDNNLVGWVKQHAKLRI 1108
Query: 274 DSSLEGQYANDDATKLVEL------ASKCLQFEARERPDIKFLLTAVTPLQ 318
+ + +D +EL A CL A RP + ++ +Q
Sbjct: 1109 SDVFDPELMKEDPALEIELLQHLKVAVACLDDRAWRRPTMVQVMAMFKEIQ 1159
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 97.4 bits (241), Expect = 7e-20
Identities = 76/287 (26%), Positives = 135/287 (46%), Gaps = 23/287 (8%)
Query: 46 KEYGLNELRRATHEFSTEYIVSESGEKAPNVVYKGKLENNRLVAVKRFSKQSWPDAQQFL 105
KE ++L +T+ F I+ G +VYK L + + VA+K+ S ++F
Sbjct: 720 KELSYDDLLDSTNSFDQANIIGCGGF---GMVYKATLPDGKKVAIKKLSGDCGQIEREFE 776
Query: 106 AEAAGVGKVRHKRMVNLIGCCAEGDERLLVAEYMPNDTLSKHLFHWDKQP--LPWEMRVR 163
AE + + +H +V L G C ++RLL+ YM N +L L + P L W+ R+R
Sbjct: 777 AEVETLSRAQHPNLVLLRGFCFYKNDRLLIYSYMENGSLDYWLHERNDGPALLKWKTRLR 836
Query: 164 VAYHVAQAL--DHCSMENRKIYHDLNAYRILFDEDGDPRLSSFGLMKNSRDGKSYST--- 218
+A A+ L H + ++ D+ + IL DE+ + L+ FGL + +++ +
Sbjct: 837 IAQGAAKGLLYLHEGCDPHILHRDIKSSNILLDENFNSHLADFGLARLMSPYETHVSTDL 896
Query: 219 --NLAYTPPEFLRTGRIIAESVIYSYGTVLLDLLSGKH----IPPSHALDLI-------R 265
L Y PPE+ + + +YS+G VLL+LL+ K P DLI
Sbjct: 897 VGTLGYIPPEYGQASVATYKGDVYSFGVVLLELLTDKRPVDMCKPKGCRDLISWVVKMKH 956
Query: 266 GKNALLLMDSSLEGQYANDDATKLVELASKCLQFEARERPDIKFLLT 312
A + D + + + + +++E+A CL ++RP + L++
Sbjct: 957 ESRASEVFDPLIYSKENDKEMFRVLEIACLCLSENPKQRPTTQQLVS 1003
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.134 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 58,798,962
Number of Sequences: 164201
Number of extensions: 2492652
Number of successful extensions: 7539
Number of sequences better than 10.0: 804
Number of HSP's better than 10.0 without gapping: 148
Number of HSP's successfully gapped in prelim test: 656
Number of HSP's that attempted gapping in prelim test: 6934
Number of HSP's gapped (non-prelim): 900
length of query: 498
length of database: 59,974,054
effective HSP length: 114
effective length of query: 384
effective length of database: 41,255,140
effective search space: 15841973760
effective search space used: 15841973760
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC147404.5


