

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147179.9 + phase: 1 /partial
(375 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
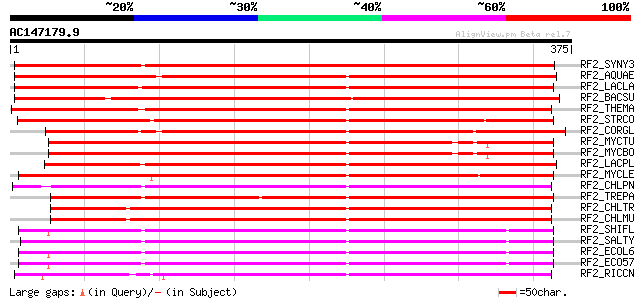
Score E
Sequences producing significant alignments: (bits) Value
RF2_SYNY3 (P74476) Peptide chain release factor 2 (RF-2) 426 e-119
RF2_AQUAE (O67695) Peptide chain release factor 2 (RF-2) 327 3e-89
RF2_LACLA (Q9CGX1) Peptide chain release factor 2 (RF-2) 309 9e-84
RF2_BACSU (P28367) Peptide chain release factor 2 (RF-2) 308 1e-83
RF2_THEMA (Q9X1R5) Peptide chain release factor 2 (RF-2) 299 9e-81
RF2_STRCO (Q53915) Peptide chain release factor 2 (RF-2) 288 1e-77
RF2_CORGL (Q8NS78) Peptide chain release factor 2 (RF-2) 285 1e-76
RF2_MYCTU (P66026) Peptide chain release factor 2 (RF-2) 283 4e-76
RF2_MYCBO (P66027) Peptide chain release factor 2 (RF-2) 283 4e-76
RF2_LACPL (Q88YL5) Peptide chain release factor 2 (RF-2) 282 1e-75
RF2_MYCLE (O32885) Peptide chain release factor 2 (RF-2) 281 1e-75
RF2_CHLPN (P56906) Peptide chain release factor 2 (RF-2) 271 2e-72
RF2_TREPA (O83585) Peptide chain release factor 2 (RF-2) 268 2e-71
RF2_CHLTR (O84465) Peptide chain release factor 2 (RF-2) 268 2e-71
RF2_CHLMU (P58105) Peptide chain release factor 2 (RF-2) 262 1e-69
RF2_SHIFL (P66025) Peptide chain release factor 2 (RF-2) 257 3e-68
RF2_SALTY (P28353) Peptide chain release factor 2 (RF-2) 257 3e-68
RF2_ECOL6 (P66023) Peptide chain release factor 2 (RF-2) 257 3e-68
RF2_ECO57 (P66024) Peptide chain release factor 2 (RF-2) 257 3e-68
RF2_RICCN (Q92IQ2) Peptide chain release factor 2 (RF-2) 256 9e-68
>RF2_SYNY3 (P74476) Peptide chain release factor 2 (RF-2)
Length = 372
Score = 426 bits (1094), Expect = e-119
Identities = 210/361 (58%), Positives = 279/361 (77%), Gaps = 2/361 (0%)
Query: 4 LRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEK 63
L++++E+ S R+ + ++ L L+ ++ LE+ A+ FWDD +AQQ L TL + K +
Sbjct: 8 LKRNLELISSRLGQTQDYLDLPGLKAKVQDLEQCAAQPDFWDDTDQAQQILQTLNETKSQ 67
Query: 64 IKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYD 123
++ ++ Q +D++ IV L E + D+ L EA + +++L K +DR+EL QLLSGPYD
Sbjct: 68 LEQWGIWQQQWQDSQAIVELLELED--DQALLTEAETTLEQLQKELDRWELQQLLSGPYD 125
Query: 124 KEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVE 183
+GA ++I AGAGGTDAQDWA+MLLRMY RW EKQ YK + E S G+EAG+KS T+E+E
Sbjct: 126 AKGATLTINAGAGGTDAQDWAEMLLRMYTRWSEKQGYKVHLAEISEGDEAGLKSVTLEIE 185
Query: 184 GRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISF 243
GRYAYGYL EKGTHR+VR SPFN+ G RQTSF+GVEVMPLL EE+++++IP++DL+IS
Sbjct: 186 GRYAYGYLKSEKGTHRLVRISPFNANGKRQTSFAGVEVMPLLGEEAISLDIPDKDLDIST 245
Query: 244 SRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQR 303
SRAGGKGGQNVNKVETAVRI H+PTG+ +RCT+ERSQL NK KAL+ LKAKLL++ EEQR
Sbjct: 246 SRAGGKGGQNVNKVETAVRIVHLPTGLAVRCTQERSQLQNKEKALAILKAKLLIVLEEQR 305
Query: 304 ATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLK 363
A +IRGD+V+A WG QIRNYVFHPY+LVKD+RT ET D+ V+DGEL FI++YL+
Sbjct: 306 AQAIAEIRGDMVEAAWGTQIRNYVFHPYQLVKDLRTNVETTDVGGVMDGELSDFIEAYLR 365
Query: 364 H 364
H
Sbjct: 366 H 366
>RF2_AQUAE (O67695) Peptide chain release factor 2 (RF-2)
Length = 373
Score = 327 bits (838), Expect = 3e-89
Identities = 168/362 (46%), Positives = 243/362 (66%), Gaps = 4/362 (1%)
Query: 4 LRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEK 63
L+ VE +R++++++ + LE EL +L+++ S +FW+D+ KA+Q + V+E
Sbjct: 6 LKGKVEELRKRLEDVKKILSPEKLESELKELDQKMSEPNFWEDQEKAKQVIQRRKWVEET 65
Query: 64 IKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYD 123
+ L + + V+D E +V +T E ++ + +E IKE+ +++ EL LSG D
Sbjct: 66 LNKLKNLEKSVKDLEELVEITSEEDTETWAMMDEE---IKEVERTLRELELKTYLSGEMD 122
Query: 124 KEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVE 183
+ A ++I AGAGGT+A DWADML RMY RW EK+ Y+ +++ + + AGIKS T+ V+
Sbjct: 123 AKNAYLTIQAGAGGTEACDWADMLFRMYKRWAEKKGYEVELIDITPDDVAGIKSVTVLVK 182
Query: 184 GRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISF 243
G YAYGYL GE+G HR+VR SPF++ R TSF+ V VMP + +E + +EI EDL+I
Sbjct: 183 GPYAYGYLKGEQGVHRLVRISPFDANARRHTSFAAVSVMPQI-DEDIKIEIKPEDLKIET 241
Query: 244 SRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQR 303
RA G GGQ VNK +TAVRITHIPTG+T+ C +ERSQ NK KAL LKAKL + ++
Sbjct: 242 FRASGAGGQYVNKTDTAVRITHIPTGITVSCQQERSQYQNKRKALELLKAKLYQLEMKKL 301
Query: 304 ATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLK 363
+ KQ G+ WG QIR+YVFHPYKL+KD+RTG+ET ++ +V+DGE+D FI+SYLK
Sbjct: 302 EEKKKQYEGEKTDIGWGHQIRSYVFHPYKLIKDLRTGYETGNVEAVMDGEIDEFIESYLK 361
Query: 364 HK 365
K
Sbjct: 362 WK 363
>RF2_LACLA (Q9CGX1) Peptide chain release factor 2 (RF-2)
Length = 365
Score = 309 bits (791), Expect = 9e-84
Identities = 153/360 (42%), Positives = 236/360 (65%), Gaps = 3/360 (0%)
Query: 4 LRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEK 63
+R +E ++++ RES L+ LE+E+A LE + + FW+D+A AQ+ + +K K
Sbjct: 6 IRNLLEGYAEKINGFRESLDLERLEEEIALLENDMAQPEFWNDQAAAQKVIDESNALKAK 65
Query: 64 IKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYD 123
+E+A+T++ + +E D+ + E + L + I+ +EL +L+ PYD
Sbjct: 66 YDNYQAMNNMLEEAQTMLEMLQE--EADEEMQAELEEMTIALGQKIESYELEIMLNQPYD 123
Query: 124 KEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVE 183
AV+ I G+GGT++QDW ML+RMY RWGE +K +++ G+ AG+KS +
Sbjct: 124 HMNAVLEIHPGSGGTESQDWGSMLMRMYTRWGEAHGFKVEILDYQDGDVAGLKSVALRFV 183
Query: 184 GRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISF 243
GR AYG+L GEKG HR+VR SPF+S R TSF+ V+VMP L ++S+ V++ + D+++
Sbjct: 184 GRNAYGFLRGEKGVHRLVRISPFDSANRRHTSFTSVDVMPEL-DDSIEVDVRDADVKMDT 242
Query: 244 SRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQR 303
R+GG GGQNVNKV T VR+THIPTG+ ++ T +R+Q N+ KA++ LK+KL + +++
Sbjct: 243 FRSGGAGGQNVNKVSTGVRLTHIPTGIVVQSTMDRTQYGNRDKAMAMLKSKLYQLEMDKK 302
Query: 304 ATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLK 363
E ++RGD + WG QIR+YVF PY+LVKD RTG+ET I++V+DGELD FI +YL+
Sbjct: 303 QAEVDELRGDQSEISWGSQIRSYVFMPYQLVKDTRTGYETGQISNVMDGELDGFINAYLR 362
>RF2_BACSU (P28367) Peptide chain release factor 2 (RF-2)
Length = 366
Score = 308 bits (789), Expect = 1e-83
Identities = 159/365 (43%), Positives = 231/365 (62%), Gaps = 5/365 (1%)
Query: 4 LRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEK 63
+R ++E + R+ + R S L+ E +A+L+E+ + FW+D+ KAQ ++ +K+
Sbjct: 6 IRAELENMASRLADFRGSLDLESKEARIAELDEQMADPEFWNDQQKAQTVINEANGLKDY 65
Query: 64 IKLLNDYKTQVEDAETIVMLTEEM-ESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPY 122
+ N YK E E + M + + E D L E +K L K + FEL LLS PY
Sbjct: 66 V---NSYKKLNESHEELQMTHDLLKEEPDTDLQLELEKELKSLTKEFNEFELQLLLSEPY 122
Query: 123 DKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEV 182
DK A++ + GAGGT++QDW MLLRMY RWGE++ +K ++ G+EAGIKS T+ +
Sbjct: 123 DKNNAILELHPGAGGTESQDWGSMLLRMYTRWGERRGFKVETLDYLPGDEAGIKSVTLLI 182
Query: 183 EGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEIS 242
+G AYGYL EKG HR+VR SPF+S G R TSF EVMP +E ++++I ED+++
Sbjct: 183 KGHNAYGYLKAEKGVHRLVRISPFDSSGRRHTSFVSCEVMPEFNDE-IDIDIRTEDIKVD 241
Query: 243 FSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQ 302
RA G GGQ+VN ++AVRITH+PT V + C ERSQ+ N+ +A+ LKAKL E+
Sbjct: 242 TYRASGAGGQHVNTTDSAVRITHLPTNVVVTCQTERSQIKNRERAMKMLKAKLYQRRIEE 301
Query: 303 RATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYL 362
+ E +IRG+ + WG QIR+YVFHPY +VKD RT E ++ +V+DG++D FI +YL
Sbjct: 302 QQAELDEIRGEQKEIGWGSQIRSYVFHPYSMVKDHRTNTEMGNVQAVMDGDIDTFIDAYL 361
Query: 363 KHKYS 367
+ K S
Sbjct: 362 RSKLS 366
>RF2_THEMA (Q9X1R5) Peptide chain release factor 2 (RF-2)
Length = 369
Score = 299 bits (765), Expect = 9e-81
Identities = 151/361 (41%), Positives = 232/361 (63%), Gaps = 5/361 (1%)
Query: 2 YSLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVK 61
+ + +E ++ K++ + +EL ++E++ + S WDD+ KA++ L +K
Sbjct: 6 FETKTKIEELEKKYKDVLSVVNEDEINKELEEVEKKLTDPSVWDDQKKAREYTQKLKRLK 65
Query: 62 EKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGP 121
+ L ++ ED E + L++E D+ + + +++EL ++ + EL +L+G
Sbjct: 66 NISEDLKRVRSLFEDLEVAIELSDE----DQEMAQHVEEIVQELEGAVKKLELEIILNGK 121
Query: 122 YDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIE 181
YD A +S+ GAGGT++QDWA MLLRMY+RW E++ + +VE GEEAGIK ATI
Sbjct: 122 YDPNNAYLSVHPGAGGTESQDWAQMLLRMYMRWAERKGFDVEIVEFQPGEEAGIKDATIL 181
Query: 182 VEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEI 241
++G YAYGYL E G HR+VR SPF++ R TSF+ V V+P + ++ +++EI EDL+I
Sbjct: 182 IKGEYAYGYLKHESGVHRLVRISPFDAARRRHTSFASVNVIPEI-DDDVDIEIRPEDLKI 240
Query: 242 SFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEE 301
RA G GGQ VNK E+AVRITH+PTG+ + C ERSQ NK AL LKAKL + E
Sbjct: 241 ETFRASGHGGQYVNKTESAVRITHLPTGIVVSCQNERSQHQNKQTALKILKAKLYQLEME 300
Query: 302 QRATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSY 361
++ E ++I+G++ WG QIR+Y+FHPY +VKD RTG ET ++ +V+DG++D FI++
Sbjct: 301 KKRREIQEIQGELKDISWGNQIRSYIFHPYTMVKDHRTGVETANVDAVMDGDIDMFIEAE 360
Query: 362 L 362
L
Sbjct: 361 L 361
>RF2_STRCO (Q53915) Peptide chain release factor 2 (RF-2)
Length = 368
Score = 288 bits (738), Expect = 1e-77
Identities = 148/358 (41%), Positives = 231/358 (64%), Gaps = 4/358 (1%)
Query: 6 KDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIK 65
++++ S ++ I L L ++A LEE+A+ S WDD AQ+ S L+ ++ +++
Sbjct: 8 EELKSLSSTMESIEAVLDLDRLRADIAVLEEQAAAPSLWDDPEAAQKITSKLSHLQAEVR 67
Query: 66 LLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKE 125
+ +++D + + EE + D EA S + + K++D E+ LLSG YD
Sbjct: 68 KAEALRGRIDDLGVLFEMAEEEDDPDTRA--EAESELAAVRKALDEMEVRTLLSGEYDAR 125
Query: 126 GAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGR 185
A+++I A AGG DA D+A+ L RMY+RW E+ YKT V E S EEAGIKS T V+
Sbjct: 126 EALVNIRAEAGGVDAADFAEKLQRMYLRWAEQHGYKTEVYETSYAEEAGIKSTTFAVQSP 185
Query: 186 YAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSR 245
YAYG LS E+GTHR+VR SPF+++G RQTSF+GVE++P++ E++ ++EI E +L + R
Sbjct: 186 YAYGTLSVEQGTHRLVRISPFDNQGRRQTSFAGVEILPVV-EQTDHIEIDESELRVDVYR 244
Query: 246 AGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRAT 305
+ G GGQ VN ++AVR+THIPTG+ + C ERSQ+ NK A++ L+AKLL ++
Sbjct: 245 SSGPGGQGVNTTDSAVRLTHIPTGIVVSCQNERSQIQNKATAMNVLQAKLLERRRQEEQA 304
Query: 306 EFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLK 363
+ ++GD + WG Q+R+YV HPY++VKD+RT HE + +V +GE+D F+++ ++
Sbjct: 305 KMDALKGDGGNS-WGNQMRSYVLHPYQMVKDLRTEHEVGNPEAVFNGEIDGFLEAGIR 361
>RF2_CORGL (Q8NS78) Peptide chain release factor 2 (RF-2)
Length = 368
Score = 285 bits (730), Expect = 1e-76
Identities = 149/347 (42%), Positives = 222/347 (63%), Gaps = 6/347 (1%)
Query: 25 QLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDAETIVMLT 84
Q + + +LE +A+ S WDD AQQ S L+ V+ +++ + D + ++ED +V L
Sbjct: 26 QEMSDRVRELEAQAADPSLWDDPDHAQQVTSELSHVQAELRKITDLRQRIEDLPIMVELA 85
Query: 85 EEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWA 144
EE E D + EE + +L ID E+ +LSG YD AVI+I +GAGG DA DWA
Sbjct: 86 EE-EDGDTSIAEEE---LADLRSLIDALEVKTMLSGEYDAREAVINIRSGAGGVDAADWA 141
Query: 145 DMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQS 204
+ML+RMY RW EK +K + + S EEAGIKSAT V G Y YG LS E+G HR+VR S
Sbjct: 142 EMLMRMYTRWAEKNGHKVDIYDISYAEEAGIKSATFVVHGDYMYGQLSVEQGAHRLVRIS 201
Query: 205 PFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRIT 264
PF+++G RQTSF+ VEV+P++ E+ +++IP+ D+ + R+ G GGQ+VN ++AVR+T
Sbjct: 202 PFDNQGRRQTSFAEVEVLPVV-EKVDSIDIPDADVRVDVYRSSGPGGQSVNTTDSAVRLT 260
Query: 265 HIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKAEWGQQIR 324
HIPTG+ + C E+SQ+ NK A+ L+AKLL ++ E + G A WG Q+R
Sbjct: 261 HIPTGIVVTCQNEKSQIQNKASAMRVLQAKLLERKRQEERAEMDAL-GAGGNASWGNQMR 319
Query: 325 NYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLKHKYSMTMS 371
+YV HPY++VKD+RT E D V+DG++D +++ ++ + + + S
Sbjct: 320 SYVLHPYQMVKDLRTNFEVNDPQKVLDGDIDGLLEAGIRWRMAESQS 366
>RF2_MYCTU (P66026) Peptide chain release factor 2 (RF-2)
Length = 371
Score = 283 bits (725), Expect = 4e-76
Identities = 147/342 (42%), Positives = 226/342 (65%), Gaps = 11/342 (3%)
Query: 27 LEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDAETIVMLT-E 85
L + KLE EAS WDD+ +AQ+ S L+ + +++ + + + +++D + L E
Sbjct: 28 LRSRIEKLEHEASDPHLWDDQTRAQRVTSELSHTQGELRRVEELRRRLDDLPVLYELAAE 87
Query: 86 EMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWAD 145
E + EA + +K L I+ E+ LLSG YD+ A+++I +GAGG DA DWA+
Sbjct: 88 EAGAAAADAVAEADAELKSLRADIEATEVRTLLSGEYDEREALVTIRSGAGGVDAADWAE 147
Query: 146 MLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSP 205
ML+RMY+RW E+ +Y V + S EEAGIKSAT V +AYG LS E+GTHR+VR SP
Sbjct: 148 MLMRMYIRWAEQHKYPVEVFDTSYAEEAGIKSATFAVHAPFAYGTLSVEQGTHRLVRISP 207
Query: 206 FNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITH 265
F+++ RQTSF+ VEV+P++ E + +++IPE D+ + R+ G GGQ+VN ++AVR+TH
Sbjct: 208 FDNQSRRQTSFAEVEVLPVV-ETTDHIDIPEGDVRVDVYRSSGPGGQSVNTTDSAVRLTH 266
Query: 266 IPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKAE----WGQ 321
IP+G+ + C E+SQL NKI A+ L+AKLL E +R E ++ D +KA+ WG
Sbjct: 267 IPSGIVVTCQNEKSQLQNKIAAMRVLQAKLL---ERKRLEERAEL--DALKADGGSSWGN 321
Query: 322 QIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLK 363
Q+R+YV HPY++VKD+RT +E + +V+DG+LD F+++ ++
Sbjct: 322 QMRSYVLHPYQMVKDLRTEYEVGNPAAVLDGDLDGFLEAGIR 363
>RF2_MYCBO (P66027) Peptide chain release factor 2 (RF-2)
Length = 371
Score = 283 bits (725), Expect = 4e-76
Identities = 147/342 (42%), Positives = 226/342 (65%), Gaps = 11/342 (3%)
Query: 27 LEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDAETIVMLT-E 85
L + KLE EAS WDD+ +AQ+ S L+ + +++ + + + +++D + L E
Sbjct: 28 LRSRIEKLEHEASDPHLWDDQTRAQRVTSELSHTQGELRRVEELRRRLDDLPVLYELAAE 87
Query: 86 EMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWAD 145
E + EA + +K L I+ E+ LLSG YD+ A+++I +GAGG DA DWA+
Sbjct: 88 EAGAAAADAVAEADAELKSLRADIEATEVRTLLSGEYDEREALVTIRSGAGGVDAADWAE 147
Query: 146 MLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSP 205
ML+RMY+RW E+ +Y V + S EEAGIKSAT V +AYG LS E+GTHR+VR SP
Sbjct: 148 MLMRMYIRWAEQHKYPVEVFDTSYAEEAGIKSATFAVHAPFAYGTLSVEQGTHRLVRISP 207
Query: 206 FNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITH 265
F+++ RQTSF+ VEV+P++ E + +++IPE D+ + R+ G GGQ+VN ++AVR+TH
Sbjct: 208 FDNQSRRQTSFAEVEVLPVV-ETTDHIDIPEGDVRVDVYRSSGPGGQSVNTTDSAVRLTH 266
Query: 266 IPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKAE----WGQ 321
IP+G+ + C E+SQL NKI A+ L+AKLL E +R E ++ D +KA+ WG
Sbjct: 267 IPSGIVVTCQNEKSQLQNKIAAMRVLQAKLL---ERKRLEERAEL--DALKADGGSSWGN 321
Query: 322 QIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLK 363
Q+R+YV HPY++VKD+RT +E + +V+DG+LD F+++ ++
Sbjct: 322 QMRSYVLHPYQMVKDLRTEYEVGNPAAVLDGDLDGFLEAGIR 363
>RF2_LACPL (Q88YL5) Peptide chain release factor 2 (RF-2)
Length = 357
Score = 282 bits (721), Expect = 1e-75
Identities = 149/343 (43%), Positives = 212/343 (61%), Gaps = 5/343 (1%)
Query: 24 LQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVED-AETIVM 82
L L + + + E + FWDD+A AQ+ + +K K +V D A +
Sbjct: 12 LDALNESIQENEARMAEPGFWDDQAAAQKVIDENNVLKGKYDTFKQLADEVGDLAVAYEL 71
Query: 83 LTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQD 142
L+EE D + E + + + ++ L LL GPYD+ A++ I GAGGT++QD
Sbjct: 72 LSEEP---DAEMQAEFETDFQHAEHDLQQYRLNLLLDGPYDRNNAILEIHPGAGGTESQD 128
Query: 143 WADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVR 202
W MLLRMY RW +K V+ G+EAGIKS T+ + G AYGYL EKG HR+VR
Sbjct: 129 WGAMLLRMYTRWAASHNFKVETVDYQAGDEAGIKSVTLLISGHNAYGYLRSEKGVHRLVR 188
Query: 203 QSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVR 262
SPF++ G R TSF+ V+VMP L +++++V+I EDL+I RA G GGQ+VNK +AVR
Sbjct: 189 ISPFDAAGRRHTSFASVDVMPEL-DDTVDVDIRPEDLKIDVYRASGAGGQHVNKTSSAVR 247
Query: 263 ITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKAEWGQQ 322
ITH+PTG+ + +RSQL N+ AL+ L+AKL EE++A E I+G+ + WG Q
Sbjct: 248 ITHVPTGIVVASQAQRSQLQNRQTALNMLRAKLYEREEEKKAKERAAIQGEQMDIGWGSQ 307
Query: 323 IRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLKHK 365
IR+YVFHPY +VKD RT +E+ +V+DG+LDPF+ +YL+ K
Sbjct: 308 IRSYVFHPYTMVKDHRTNYESHHGQAVMDGDLDPFMDAYLQWK 350
>RF2_MYCLE (O32885) Peptide chain release factor 2 (RF-2)
Length = 374
Score = 281 bits (720), Expect = 1e-75
Identities = 141/361 (39%), Positives = 228/361 (63%), Gaps = 6/361 (1%)
Query: 7 DVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKL 66
D+ + + ++ L + KLE EAS WDD+ +AQ+ S L+ + +++
Sbjct: 8 DIAALDSTLTTVERVLDVEGLRTRIEKLEHEASDPKLWDDQVRAQRVTSELSHAQGELRR 67
Query: 67 LNDYKTQVEDAETIVMLTEEMESVDKG----LYEEASSLIKELNKSIDRFELTQLLSGPY 122
+ + + +++D + L E + + EA + +K L I+ E+ LLSG Y
Sbjct: 68 IEELRRRLDDLPVLYELAAEERAAAAASGMEAFAEADAELKALRVDIEATEVRTLLSGEY 127
Query: 123 DKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEV 182
D+ A+I+I +GAGG DA DWA+ML+RMY+RW E+ +Y V++ S EEAG+KSAT V
Sbjct: 128 DEREALITIRSGAGGVDAADWAEMLMRMYIRWAEQHKYGVEVLDTSYAEEAGVKSATFAV 187
Query: 183 EGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEIS 242
+AYG L+ E+GTHR+VR SPF+++ RQTSF+ VEV+P++ E + +++IPE D+ +
Sbjct: 188 HAPFAYGTLASEQGTHRLVRISPFDNQSRRQTSFAEVEVLPVV-EITDHIDIPEGDVRVD 246
Query: 243 FSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQ 302
R+ G GGQ+VN ++AVR+TH+PTG+ + C E+SQL NK+ A+ L+AKLL +
Sbjct: 247 VYRSSGPGGQSVNTTDSAVRLTHVPTGLVVTCQNEKSQLQNKVSAMRVLQAKLLERKRLE 306
Query: 303 RATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYL 362
E ++G + WG QIR+YV HPY++VKD+R +E + T+V+DG++D F+++ +
Sbjct: 307 ERAELDALKGR-GGSSWGNQIRSYVLHPYQMVKDLRNEYEVGNPTAVLDGDIDGFLEAGI 365
Query: 363 K 363
+
Sbjct: 366 R 366
>RF2_CHLPN (P56906) Peptide chain release factor 2 (RF-2)
Length = 369
Score = 271 bits (693), Expect = 2e-72
Identities = 144/361 (39%), Positives = 218/361 (59%), Gaps = 10/361 (2%)
Query: 3 SLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKE 62
+LR ++ +A++ + ++ + ++EL LEEE+S +FW D A + + ++
Sbjct: 11 ALRTEISLAARSLFDLDKK------QKELQVLEEESSEENFWQDSVHAGKISEQIVSLRR 64
Query: 63 KIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPY 122
+I+ + K++++ E + + +E D + E+ K + +E +LLSG
Sbjct: 65 QIQEYQELKSKIDAIEFFLEDADALE--DPAICEDLEKEFLFCEKKLAVWETQRLLSGEA 122
Query: 123 DKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEV 182
DK ++I AGAGGT++ DW +ML RMY RW K ++ VV++ GE GIK T++
Sbjct: 123 DKNSCFLTINAGAGGTESCDWVEMLFRMYSRWATKHQWALEVVDRLDGEVVGIKHVTVKF 182
Query: 183 EGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEIS 242
G YAYGY E+G HR+VR SPF+S G R TSF+ V+V P + ++ + +EI DL I
Sbjct: 183 SGMYAYGYAKAERGVHRLVRISPFDSNGKRHTSFASVDVFPEI-DDQIKIEIRPNDLRID 241
Query: 243 FSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQ 302
R+ G GGQ+VN E+AVRITH+P+GV + C ERSQ+ N+ + L+AKL ++
Sbjct: 242 TFRSSGAGGQHVNVTESAVRITHLPSGVVVSCQNERSQIQNRESCMKMLQAKLYQQVLQE 301
Query: 303 RATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGE-LDPFIKSY 361
R + R D + WG QIRNYVF PY LVKDVRTGHET ++ +++DGE LD FIK+Y
Sbjct: 302 RLEKQSLDRKDKKEIAWGSQIRNYVFQPYTLVKDVRTGHETGNVQAMLDGELLDEFIKAY 361
Query: 362 L 362
L
Sbjct: 362 L 362
>RF2_TREPA (O83585) Peptide chain release factor 2 (RF-2)
Length = 368
Score = 268 bits (684), Expect = 2e-71
Identities = 136/336 (40%), Positives = 209/336 (61%), Gaps = 4/336 (1%)
Query: 28 EQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDAETIVMLTEEM 87
E +A LE A+ FW +RA+A+ L+ L ++ ++ + + D + L E
Sbjct: 25 EARIATLEAAAAAPDFWSERARAEALLAELKTLRATLEPWRALRRESADLRALYELAREA 84
Query: 88 ESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWADML 147
+ D L E SSL +++ + LT+LL D+ A ++I +GAGG +A DWA ML
Sbjct: 85 Q--DASLEPELSSLFSDISARFEEASLTRLLHEEVDRLDAFVTIHSGAGGVEACDWAQML 142
Query: 148 LRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSPFN 207
+RMY RW E++ + +V+ L E G+KS T+++ G +A+G+L GE G HR+VR SPF+
Sbjct: 143 MRMYTRWAERRSFCVHIVDL-LESEGGVKSVTLKICGSHAFGFLKGETGVHRLVRISPFD 201
Query: 208 SKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITHIP 267
S R TSF+ V P+L ++ + V I ED+ + R+GG GGQ+VNK ++AVRITH+P
Sbjct: 202 SAARRHTSFTSTYVFPVL-DDHVEVHIRSEDMRVDTYRSGGAGGQHVNKTDSAVRITHLP 260
Query: 268 TGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKAEWGQQIRNYV 327
TG+ + C ERSQ++N+ ALS L+A+L +++ E ++ + WG QIR+YV
Sbjct: 261 TGIVVTCQNERSQISNRATALSLLRARLYAYERQKKQQEHQRFASEKKDISWGNQIRSYV 320
Query: 328 FHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLK 363
FHPY +VKD R+ ET +I +V+DG L+PFI+SYL+
Sbjct: 321 FHPYTMVKDHRSKCETGNIHAVMDGALEPFIRSYLE 356
>RF2_CHLTR (O84465) Peptide chain release factor 2 (RF-2)
Length = 369
Score = 268 bits (684), Expect = 2e-71
Identities = 141/336 (41%), Positives = 205/336 (60%), Gaps = 4/336 (1%)
Query: 28 EQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDAETIVMLTEEM 87
E EL +LE++A FWDD +A + +A +K+++ N+ K +V + L +E
Sbjct: 30 ENELKELEQQAVQDGFWDDVVRAGKISERIARLKQQLSEFNELKNKVSAIQ--FFLEDEE 87
Query: 88 ESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWADML 147
S D + +E K I +E +LLSG D+ +SI AGAGGT++ DW +ML
Sbjct: 88 SSKDLEMQKELEKEFVFCEKKITEWETLRLLSGELDRNSCFLSINAGAGGTESCDWVEML 147
Query: 148 LRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSPFN 207
LRMY+RW ++ V+++ GE AGIK T+++ G YAYGY E G HR+VR SPF+
Sbjct: 148 LRMYMRWASSHSWRIEVIDRLDGEVAGIKHITLKLVGEYAYGYAKAESGVHRLVRISPFD 207
Query: 208 SKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITHIP 267
S R TSF+ VEV P + ++ + VEI D+ I R+ G GGQ+VN ++AVRITH P
Sbjct: 208 SNAKRHTSFASVEVFPEI-DDKIEVEIRPGDIRIDTYRSSGAGGQHVNVTDSAVRITHFP 266
Query: 268 TGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKAEWGQQIRNYV 327
TG+ + C ERSQ+ N+ ++ L+A++ ++R + R + + WG QIRNYV
Sbjct: 267 TGIVVSCQNERSQIQNREACMNMLRARIYQKLLQERLEKQNIDRKNKKEISWGSQIRNYV 326
Query: 328 FHPYKLVKDVRTGHETPDITSVIDGE-LDPFIKSYL 362
F PY LVKDVRTG+E +I +++DGE LD FIK+YL
Sbjct: 327 FQPYTLVKDVRTGYEVGNIQAMMDGELLDAFIKAYL 362
>RF2_CHLMU (P58105) Peptide chain release factor 2 (RF-2)
Length = 369
Score = 262 bits (669), Expect = 1e-69
Identities = 139/336 (41%), Positives = 203/336 (60%), Gaps = 4/336 (1%)
Query: 28 EQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDAETIVMLTEEM 87
E EL LE+ + SFWDD ++A + +A +K+++ N+ K ++ + L +E
Sbjct: 30 ETELKNLEQLSIQDSFWDDVSRAGRISEKIARLKQQLLEYNELKNKINSIK--FFLEDEE 87
Query: 88 ESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWADML 147
S D + +E K I +E +LL+G D+ +SI AGAGGT++ DW +ML
Sbjct: 88 SSKDLEIQKELEKEFIFCEKKIAEWETLRLLAGELDRNSCFLSINAGAGGTESCDWVEML 147
Query: 148 LRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSPFN 207
RMY RW +K V+++ GE AGIK T+++ G YAYGY E G HR+VR SPF+
Sbjct: 148 FRMYSRWANSHGWKVEVIDRLDGEVAGIKHITLKLIGEYAYGYAKAESGVHRLVRISPFD 207
Query: 208 SKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITHIP 267
S R TSF+ VEV P + ++ + VEI D+ I R+ G GGQ+VN ++AVRITH P
Sbjct: 208 SNAKRHTSFASVEVFPEI-DDKIEVEIRPGDIRIDTYRSSGAGGQHVNVTDSAVRITHFP 266
Query: 268 TGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKAEWGQQIRNYV 327
TG+ + C ERSQ+ N+ ++ L+A++ ++R + R + + WG QIRNYV
Sbjct: 267 TGIVVSCQSERSQIQNREACMNMLRARIYQKLLQERLEKQNTNRKEKKEISWGSQIRNYV 326
Query: 328 FHPYKLVKDVRTGHETPDITSVIDGE-LDPFIKSYL 362
F PY LVKDVRTG+E +I +++DGE LD FIK+YL
Sbjct: 327 FQPYTLVKDVRTGYEVGNIQAMMDGELLDAFIKAYL 362
>RF2_SHIFL (P66025) Peptide chain release factor 2 (RF-2)
Length = 365
Score = 257 bits (657), Expect = 3e-68
Identities = 138/364 (37%), Positives = 218/364 (58%), Gaps = 11/364 (3%)
Query: 7 DVEIASQRVKEIRESSGL-------QLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLAD 59
++ + R++++ E S + ++ L ++ E W++ +AQ +
Sbjct: 3 EINPVNNRIQDLTERSDVLRGYLDYDAKKERLEEVNAELEQPDVWNEPERAQALGKERSS 62
Query: 60 VKEKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLS 119
++ + L+ K +ED ++ L E + D+ + EA + + L + + + E ++ S
Sbjct: 63 LEAVVDTLDQMKQGLEDVSGLLELAVEAD--DEETFNEAVAELDALEEKLAQLEFRRMFS 120
Query: 120 GPYDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSAT 179
G YD + I AG+GGT+AQDWA ML RMY+RW E + +KT ++E+S GE AGIKS T
Sbjct: 121 GEYDSADCYLDIQAGSGGTEAQDWASMLERMYLRWAESRGFKTEIIEESEGEVAGIKSVT 180
Query: 180 IEVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDL 239
I++ G YAYG+L E G HR+VR+SPF+S G R TSFS V P + ++ +++EI DL
Sbjct: 181 IKISGDYAYGWLRTETGVHRLVRKSPFDSGGRRHTSFSSAFVYPEV-DDDIDIEINPADL 239
Query: 240 EISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIA 299
I RA G GGQ+VN+ E+AVRITHIPTG+ +C +RSQ NK +A+ ++KAKL +
Sbjct: 240 RIDVYRASGAGGQHVNRTESAVRITHIPTGIVTQCQNDRSQHKNKDQAMKQMKAKLYELE 299
Query: 300 EEQRATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIK 359
+++ E + + + WG QIR+YV + +KD+RTG ET + +V+DG LD FI+
Sbjct: 300 MQKKNAEKQAMEDNKSDIGWGSQIRSYVLDDSR-IKDLRTGVETRNTQAVLDGSLDQFIE 358
Query: 360 SYLK 363
+ LK
Sbjct: 359 ASLK 362
>RF2_SALTY (P28353) Peptide chain release factor 2 (RF-2)
Length = 365
Score = 257 bits (657), Expect = 3e-68
Identities = 137/356 (38%), Positives = 214/356 (59%), Gaps = 4/356 (1%)
Query: 8 VEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLL 67
++ ++R +R ++ L ++ E W++ +AQ + ++ + L
Sbjct: 11 IQDLTERTNVLRGYLDYDAKKERLEEVNAELEQPDVWNEPERAQALGKERSSLEAIVDTL 70
Query: 68 NDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGA 127
+ ++D ++ L E + D+ + EA + + L + + + E ++ SG YD
Sbjct: 71 DQMTQGLDDVSGLLELAVEAD--DEETFNEAVAELNTLEEKLAQLEFRRMFSGEYDSADC 128
Query: 128 VISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYA 187
+ I AG+GGT+AQDWA MLLRMY+RW E + +KT V+E+S GE AGIKSATI++ G YA
Sbjct: 129 YLDIQAGSGGTEAQDWASMLLRMYLRWAEARGFKTEVIEESEGEVAGIKSATIKISGEYA 188
Query: 188 YGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAG 247
YG+L E G HR+VR+SPF+S G R TSFS V P + ++ ++++I DL I RA
Sbjct: 189 YGWLRTETGVHRLVRKSPFDSGGRRHTSFSSAFVYPEV-DDDIDIDINPADLRIDVYRAS 247
Query: 248 GKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEF 307
G GGQ+VN+ E+AVRITHIPTG+ +C +RSQ NK +A+ ++KAKL + +++ E
Sbjct: 248 GAGGQHVNRTESAVRITHIPTGIVTQCQNDRSQHKNKDQAMKQMKAKLYELEMQKKNAEK 307
Query: 308 KQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLK 363
+ + WG QIR+YV + +KD+RTG ET + +V+DG LD FI++ LK
Sbjct: 308 QAMEDTKSDIGWGSQIRSYVLDDSR-IKDLRTGVETRNTQAVLDGSLDQFIEASLK 362
>RF2_ECOL6 (P66023) Peptide chain release factor 2 (RF-2)
Length = 365
Score = 257 bits (657), Expect = 3e-68
Identities = 138/364 (37%), Positives = 218/364 (58%), Gaps = 11/364 (3%)
Query: 7 DVEIASQRVKEIRESSGL-------QLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLAD 59
++ + R++++ E S + ++ L ++ E W++ +AQ +
Sbjct: 3 EINPVNNRIQDLTERSDVLRGYLDYDAKKERLEEVNAELEQPDVWNEPERAQALGKERSS 62
Query: 60 VKEKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLS 119
++ + L+ K +ED ++ L E + D+ + EA + + L + + + E ++ S
Sbjct: 63 LEAVVDTLDQMKQGLEDVSGLLELAVEAD--DEETFNEAVAELDALEEKLAQLEFRRMFS 120
Query: 120 GPYDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSAT 179
G YD + I AG+GGT+AQDWA ML RMY+RW E + +KT ++E+S GE AGIKS T
Sbjct: 121 GEYDSADCYLDIQAGSGGTEAQDWASMLERMYLRWAESRGFKTEIIEESEGEVAGIKSVT 180
Query: 180 IEVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDL 239
I++ G YAYG+L E G HR+VR+SPF+S G R TSFS V P + ++ +++EI DL
Sbjct: 181 IKISGDYAYGWLRTETGVHRLVRKSPFDSGGRRHTSFSSAFVYPEV-DDDIDIEINPADL 239
Query: 240 EISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIA 299
I RA G GGQ+VN+ E+AVRITHIPTG+ +C +RSQ NK +A+ ++KAKL +
Sbjct: 240 RIDVYRASGAGGQHVNRTESAVRITHIPTGIVTQCQNDRSQHKNKDQAMKQMKAKLYELE 299
Query: 300 EEQRATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIK 359
+++ E + + + WG QIR+YV + +KD+RTG ET + +V+DG LD FI+
Sbjct: 300 MQKKNAEKQAMEDNKSDIGWGSQIRSYVLDDSR-IKDLRTGVETRNTQAVLDGSLDQFIE 358
Query: 360 SYLK 363
+ LK
Sbjct: 359 ASLK 362
>RF2_ECO57 (P66024) Peptide chain release factor 2 (RF-2)
Length = 365
Score = 257 bits (657), Expect = 3e-68
Identities = 138/364 (37%), Positives = 218/364 (58%), Gaps = 11/364 (3%)
Query: 7 DVEIASQRVKEIRESSGL-------QLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLAD 59
++ + R++++ E S + ++ L ++ E W++ +AQ +
Sbjct: 3 EINPVNNRIQDLTERSDVLRGYLDYDAKKERLEEVNAELEQPDVWNEPERAQALGKERSS 62
Query: 60 VKEKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLS 119
++ + L+ K +ED ++ L E + D+ + EA + + L + + + E ++ S
Sbjct: 63 LEAVVDTLDQMKQGLEDVSGLLELAVEAD--DEETFNEAVAELDALEEKLAQLEFRRMFS 120
Query: 120 GPYDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSAT 179
G YD + I AG+GGT+AQDWA ML RMY+RW E + +KT ++E+S GE AGIKS T
Sbjct: 121 GEYDSADCYLDIQAGSGGTEAQDWASMLERMYLRWAESRGFKTEIIEESEGEVAGIKSVT 180
Query: 180 IEVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDL 239
I++ G YAYG+L E G HR+VR+SPF+S G R TSFS V P + ++ +++EI DL
Sbjct: 181 IKISGDYAYGWLRTETGVHRLVRKSPFDSGGRRHTSFSSAFVYPEV-DDDIDIEINPADL 239
Query: 240 EISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIA 299
I RA G GGQ+VN+ E+AVRITHIPTG+ +C +RSQ NK +A+ ++KAKL +
Sbjct: 240 RIDVYRASGAGGQHVNRTESAVRITHIPTGIVTQCQNDRSQHKNKDQAMKQMKAKLYELE 299
Query: 300 EEQRATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIK 359
+++ E + + + WG QIR+YV + +KD+RTG ET + +V+DG LD FI+
Sbjct: 300 MQKKNAEKQAMEDNKSDIGWGSQIRSYVLDDSR-IKDLRTGVETRNTQAVLDGSLDQFIE 358
Query: 360 SYLK 363
+ LK
Sbjct: 359 ASLK 362
>RF2_RICCN (Q92IQ2) Peptide chain release factor 2 (RF-2)
Length = 368
Score = 256 bits (653), Expect = 9e-68
Identities = 140/366 (38%), Positives = 217/366 (59%), Gaps = 13/366 (3%)
Query: 4 LRKDVEIASQRVKEIRE----SSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLAD 59
+R ++E +++++ E S ++ + L LEE + S W+D+A AQ+ L ++
Sbjct: 1 MRAEIENYVKKIEQSLELLWRSLDVEASTERLNALEELTADPSLWNDQANAQKLLREKSN 60
Query: 60 VKEKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSL---IKELNKSIDRFELTQ 116
++EK+ N K+ ++DA + EEM + L E S + +K L+ +FE
Sbjct: 61 LEEKLNAFNKLKSNLKDALEL----EEMAEAENDL-ETLSQIEQDLKNLSIIAAKFETEC 115
Query: 117 LLSGPYDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIK 176
L SG D + I AGAGGT++ DWA +++RMY+R+ E+ +KT ++ GEEAGIK
Sbjct: 116 LFSGEADGNNCFLEINAGAGGTESHDWASIMMRMYLRFAERLGFKTEIINMINGEEAGIK 175
Query: 177 SATIEVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPE 236
S TI + G+ AYG+ E G HR+VR SPFN+ G R TSF+ V P + ++++ + I +
Sbjct: 176 SCTIRIIGKRAYGWFKTETGVHRLVRISPFNAAGKRMTSFASSWVYPEI-DDNIAITIED 234
Query: 237 EDLEISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLL 296
+DL I RA G GGQ+VN ++AVRITHIPTG +C +RSQ NK +A+ L+AKL
Sbjct: 235 KDLRIDTFRASGAGGQHVNTTDSAVRITHIPTGTVTQCQSDRSQHKNKAQAMKMLQAKLY 294
Query: 297 VIAEEQRATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDP 356
+ ++R + WG QIR+YV PY +VKD+RT +ET D V+DG+L+
Sbjct: 295 ELEMQKRTDSVNEQNAAKTDNSWGHQIRSYVLQPYHMVKDLRTDYETSDTKGVLDGDLEE 354
Query: 357 FIKSYL 362
F+ ++L
Sbjct: 355 FVSAHL 360
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.131 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,000,137
Number of Sequences: 164201
Number of extensions: 1688125
Number of successful extensions: 5841
Number of sequences better than 10.0: 254
Number of HSP's better than 10.0 without gapping: 162
Number of HSP's successfully gapped in prelim test: 92
Number of HSP's that attempted gapping in prelim test: 5389
Number of HSP's gapped (non-prelim): 296
length of query: 375
length of database: 59,974,054
effective HSP length: 112
effective length of query: 263
effective length of database: 41,583,542
effective search space: 10936471546
effective search space used: 10936471546
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 66 (30.0 bits)
Medicago: description of AC147179.9


