

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147179.5 - phase: 0
(706 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
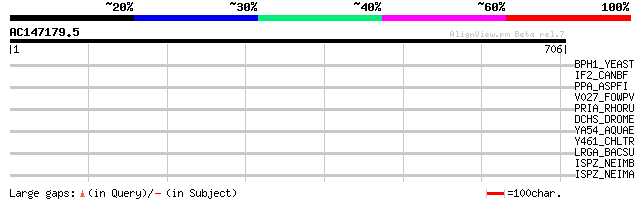
Score E
Sequences producing significant alignments: (bits) Value
BPH1_YEAST (P25356) Beige protein homolog 1 35 0.52
IF2_CANBF (Q7VQM3) Translation initiation factor IF-2 35 0.68
PPA_ASPFI (Q12546) Acid phosphatase precursor (EC 3.1.3.2) (pH 6... 34 1.5
V027_FOWPV (Q9J5H4) G-protein coupled receptor homolog FPV027 33 3.4
PRIA_RHORU (P05445) Primosomal protein N' (Replication factor Y) 32 4.4
DCHS_DROME (Q05733) Histidine decarboxylase (EC 4.1.1.22) (HDC) 32 7.5
YA54_AQUAE (O67153) Hypothetical UPF0151 protein AQ_1054 31 9.8
Y461_CHLTR (O84467) Hypothetical UPF0151 protein CT461 precursor 31 9.8
LRGA_BACSU (P94515) Antiholin-like protein lrgA 31 9.8
ISPZ_NEIMB (P65196) Probable intracellular septation protein 31 9.8
ISPZ_NEIMA (P65195) Probable intracellular septation protein 31 9.8
>BPH1_YEAST (P25356) Beige protein homolog 1
Length = 2167
Score = 35.4 bits (80), Expect = 0.52
Identities = 26/95 (27%), Positives = 49/95 (51%), Gaps = 8/95 (8%)
Query: 349 DFKSGAKHAIILPTFPLDSRFMQTSSWHHNYECQSVASSSYETIRALVFSASPV--ESVV 406
DF +K +IIL T ++ + W+ N + S+AS E IR L F S + S+
Sbjct: 552 DFNDISK-SIILDTDAYENIVLNLDLWYMNEDQSSLASGGLEIIRFLFFQISSLMEASIY 610
Query: 407 ARVYDSRYGDLVLVVE-----AHMTKRADENSRGN 436
++ +++ D+ ++ + +TKR ++NS+ N
Sbjct: 611 SKFNSNKFNDMNILEKLCLSYQAVTKRENQNSKFN 645
>IF2_CANBF (Q7VQM3) Translation initiation factor IF-2
Length = 897
Score = 35.0 bits (79), Expect = 0.68
Identities = 26/95 (27%), Positives = 42/95 (43%), Gaps = 6/95 (6%)
Query: 341 GHVSYVDLDFKSGAKHAIILPTFPLDSRFMQTSSWHHNYECQSVASSSYETIRALVFSAS 400
GH +V++ SG +L + + S ++ S HH V SS + S
Sbjct: 540 GHTQFVNVSAVSGEGVNDLLDSILVQSEILELKSMHHGLAKAVVIESSLDR------SRG 593
Query: 401 PVESVVARVYDSRYGDLVLVVEAHMTKRADENSRG 435
PV +V+ R + + GD+VL + RA +S G
Sbjct: 594 PVVTVLVRSGELKCGDIVLCGTEYGRVRAMRDSLG 628
>PPA_ASPFI (Q12546) Acid phosphatase precursor (EC 3.1.3.2) (pH
6-optimum acid phosphatase) (APase6)
Length = 614
Score = 33.9 bits (76), Expect = 1.5
Identities = 24/77 (31%), Positives = 34/77 (43%), Gaps = 10/77 (12%)
Query: 205 FGHPTDQLLKDLDLELSHWDSQSEKPVTK------ISFGHFPLSFSAPSSSGRTLKEVF- 257
FG+ + + E HW Q V + I H P+ SA SS ++E F
Sbjct: 425 FGNVNGSVHETKSYEQWHWLQQDLAKVDRSKTPWVIVMSHRPMYSSAYSSYQLHVREAFE 484
Query: 258 ---LKHSISAYLCGHLH 271
LK+ + AYL GH+H
Sbjct: 485 GLLLKYGVDAYLSGHIH 501
>V027_FOWPV (Q9J5H4) G-protein coupled receptor homolog FPV027
Length = 336
Score = 32.7 bits (73), Expect = 3.4
Identities = 23/77 (29%), Positives = 36/77 (45%), Gaps = 5/77 (6%)
Query: 487 WSWKEFFVMGCQWASLYYPLLWSALGFMFSFLLVPKALLFFQMNL-YTYRNFIANKGVVN 545
W EF C+ +S ++ A F+ +F+ + K L L Y YR + N V
Sbjct: 92 WYLGEFL---CRVSSFFFTTNMYASMFLLTFISIDKYLTLTSHRLVYKYRKY-RNYYVCI 147
Query: 546 GALWILQELCRVPTLWF 562
GA+W + VPTL++
Sbjct: 148 GAIWCISIALGVPTLYY 164
>PRIA_RHORU (P05445) Primosomal protein N' (Replication factor Y)
Length = 811
Score = 32.3 bits (72), Expect = 4.4
Identities = 32/116 (27%), Positives = 47/116 (39%), Gaps = 17/116 (14%)
Query: 313 GAPPQEFWEWEIGDWRKSRAIRILAIDRGHVSYVDLDFKSGAKHAIILPTFPL------- 365
GA P + W ++G + RA R +A+ R V GA+ A+ LP L
Sbjct: 299 GAAPVQ-WHSQMGAAARRRAWRAVALGRAPVVV-------GARSALFLPYPDLGLIIVDE 350
Query: 366 --DSRFMQTSSWHHNYECQSVASSSYETIRALVFSASPVESVVARVYDSRYGDLVL 419
DS F Q +N +V + A++ SA+P + RY LVL
Sbjct: 351 EHDSAFKQEEGVPYNARDMAVVRARLGGFPAVLASATPSLETIENARQGRYRHLVL 406
>DCHS_DROME (Q05733) Histidine decarboxylase (EC 4.1.1.22) (HDC)
Length = 847
Score = 31.6 bits (70), Expect = 7.5
Identities = 33/151 (21%), Positives = 59/151 (38%), Gaps = 32/151 (21%)
Query: 112 EWVEYRNVMDGVIERSGL---HKSLFFDLRGNHDSFGVP-----------VVGGSF---- 153
+W+E R V V+E + ++ + + +++FG VV GSF
Sbjct: 462 DWMEIRQVASTVLEEMNITISNRVYLKETKEKNEAFGSSLLLSNSPLSPKVVNGSFAAIF 521
Query: 154 ---DFFSKYSVNGQLGRNRS------VNSVTLETKE---RKHLFVGIDTTMSAGAGLR-- 199
+F +K ++ S V + + K+ H+ V + TT+ AG G
Sbjct: 522 DADEFLAKTYAGVRIAHQESPSMRRRVRGILMSGKQFSLDSHMDVVVQTTLDAGNGATRT 581
Query: 200 GPTNVFGHPTDQLLKDLDLELSHWDSQSEKP 230
TN +GH T + + + S + E P
Sbjct: 582 STTNSYGHTTSAAQANSERQASIQEDNEESP 612
>YA54_AQUAE (O67153) Hypothetical UPF0151 protein AQ_1054
Length = 261
Score = 31.2 bits (69), Expect = 9.8
Identities = 21/65 (32%), Positives = 30/65 (45%), Gaps = 6/65 (9%)
Query: 39 ETVIEPKGTPDSAVWVVQLSDLHLSVHHPNRALDFTNLVGHALSFINPSLVLITGDLTDG 98
ET PKGT + ++ SD+HL P D +V P +++ TGD DG
Sbjct: 26 ETEKLPKGTE---IKIMNASDMHLG---PVMREDRVEMVKRVYEREKPDILVATGDTVDG 79
Query: 99 KSKDL 103
K+L
Sbjct: 80 NMKNL 84
>Y461_CHLTR (O84467) Hypothetical UPF0151 protein CT461 precursor
Length = 329
Score = 31.2 bits (69), Expect = 9.8
Identities = 15/41 (36%), Positives = 25/41 (60%), Gaps = 3/41 (7%)
Query: 54 VVQLSDLHLSVHHPNRALDFTNLVGHALSFINPSLVLITGD 94
+VQ+SDLHL+ P+ F V +S ++P +++ TGD
Sbjct: 54 IVQISDLHLNHSTPDA---FLKKVSRKISSLSPDILVFTGD 91
>LRGA_BACSU (P94515) Antiholin-like protein lrgA
Length = 146
Score = 31.2 bits (69), Expect = 9.8
Identities = 15/53 (28%), Positives = 29/53 (54%)
Query: 624 LFVVLPAILVTGALTAERAIYRERVLALSGKKKDDIDLNSRRPLLNGNHNSTI 676
L +VL +L T L ++ + +L+LSGK+K + D+ ++ N+N +
Sbjct: 92 LQIVLVILLATIILLGATGLFSQLILSLSGKRKTEADMKTKTVQSPQNNNELV 144
>ISPZ_NEIMB (P65196) Probable intracellular septation protein
Length = 176
Score = 31.2 bits (69), Expect = 9.8
Identities = 24/110 (21%), Positives = 52/110 (46%), Gaps = 14/110 (12%)
Query: 485 LSWSWKEFFVMGCQWASLYYPLLWSALGFMFS---FLLVPKALLFFQMNLYTYRNFIANK 541
L W +K+ M QW L +++ + F++ ++LF+ L+ + + +A K
Sbjct: 41 LYWKYKKLDTM--QWVGLVLIVVFGGATIVLGDSRFIMWKPSVLFWLGALFLWGSHLAGK 98
Query: 542 GVVNGALWILQELCRVPTLW----FGWIGYLFYLILFPWFMGQVFTEGKS 587
+ ++ +E+ +W + W+G+L ++ + WF VFT +S
Sbjct: 99 NGLKASIG--REIQLPDAVWAKLTYMWVGFLIFMGIANWF---VFTRFES 143
>ISPZ_NEIMA (P65195) Probable intracellular septation protein
Length = 176
Score = 31.2 bits (69), Expect = 9.8
Identities = 24/110 (21%), Positives = 52/110 (46%), Gaps = 14/110 (12%)
Query: 485 LSWSWKEFFVMGCQWASLYYPLLWSALGFMFS---FLLVPKALLFFQMNLYTYRNFIANK 541
L W +K+ M QW L +++ + F++ ++LF+ L+ + + +A K
Sbjct: 41 LYWKYKKLDTM--QWVGLVLIVVFGGATIVLGDSRFIMWKPSVLFWLGALFLWGSHLAGK 98
Query: 542 GVVNGALWILQELCRVPTLW----FGWIGYLFYLILFPWFMGQVFTEGKS 587
+ ++ +E+ +W + W+G+L ++ + WF VFT +S
Sbjct: 99 NGLKASIG--REIQLPDAVWAKLTYMWVGFLIFMGIANWF---VFTRFES 143
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.138 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 86,163,845
Number of Sequences: 164201
Number of extensions: 3757177
Number of successful extensions: 9867
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 9863
Number of HSP's gapped (non-prelim): 13
length of query: 706
length of database: 59,974,054
effective HSP length: 117
effective length of query: 589
effective length of database: 40,762,537
effective search space: 24009134293
effective search space used: 24009134293
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 69 (31.2 bits)
Medicago: description of AC147179.5


