

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
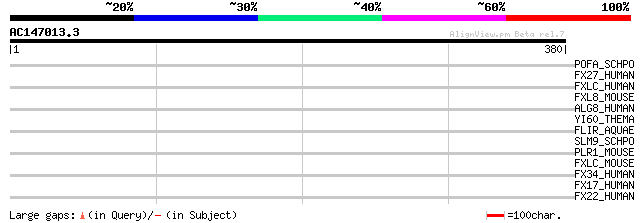
Query= AC147013.3 + phase: 0
(380 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
POFA_SCHPO (Q9P7W4) F-box/WD-repeat protein pof10 (Skp1-binding ... 42 0.003
FX27_HUMAN (Q8NI29) F-box only protein 27 (F-box/G-domain protei... 33 1.2
FXLC_HUMAN (Q9NXK8) F-box/LRR-repeat protein 12 (F-box and leuci... 32 2.7
FXL8_MOUSE (Q8CIG9) F-box/LRR-repeat protein 8 (F-box and leucin... 32 2.7
ALG8_HUMAN (Q9BVK2) Probable dolichyl pyrophosphate Glc1Man9GlcN... 32 3.5
YI60_THEMA (Q9X2H4) Hypothetical UPF0097 protein TM1860 31 4.6
FLIR_AQUAE (O67773) Flagellar biosynthetic protein fliR 31 4.6
SLM9_SCHPO (O74309) Mitotic control protein slm9 31 6.0
PLR1_MOUSE (Q922V4) Pleiotropic regulator 1 31 6.0
FXLC_MOUSE (Q9EPX5) F-box/LRR-repeat protein 12 (F-box and leuci... 31 6.0
FX34_HUMAN (Q9NWN3) F-box only protein 34 31 6.0
FX17_HUMAN (Q96EF6) F-box only protein 17 (F-box only protein 26) 31 6.0
FX22_HUMAN (Q8NEZ5) F-box only protein 22 (F-box protein FBX22p44) 30 7.8
>POFA_SCHPO (Q9P7W4) F-box/WD-repeat protein pof10 (Skp1-binding
protein 2)
Length = 662
Score = 41.6 bits (96), Expect = 0.003
Identities = 19/45 (42%), Positives = 27/45 (59%)
Query: 16 AETTANKPLPFLPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSD 60
+ N+ + LP+E+++II L RSLL +C CK WK L SD
Sbjct: 23 SSNNTNRTVLNLPKEILIIIFSFLDPRSLLSAQCTCKYWKKLLSD 67
>FX27_HUMAN (Q8NI29) F-box only protein 27 (F-box/G-domain protein
5)
Length = 283
Score = 33.1 bits (74), Expect = 1.2
Identities = 19/60 (31%), Positives = 31/60 (51%), Gaps = 3/60 (5%)
Query: 1 MGGENVNNTDSTLPVAETTANKPLPF--LPEELIVIILLRLPVRSLL-RFKCVCKSWKTL 57
MG + +P E + L LP EL++++L +P R+LL R + VC+ W+ L
Sbjct: 1 MGASVSRGRAARVPAPEPEPEEALDLSQLPPELLLVVLSHVPPRTLLGRCRQVCRGWRAL 60
>FXLC_HUMAN (Q9NXK8) F-box/LRR-repeat protein 12 (F-box and
leucine-rich repeat protein 12) (F-box protein FBL12)
Length = 326
Score = 32.0 bits (71), Expect = 2.7
Identities = 14/34 (41%), Positives = 19/34 (55%)
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSD 60
LP+ +++ I LPVR +R VC WK L D
Sbjct: 7 LPDSVLLEIFSYLPVRDRIRISRVCHRWKRLVDD 40
>FXL8_MOUSE (Q8CIG9) F-box/LRR-repeat protein 8 (F-box and
leucine-rich repeat protein 8) (F-box protein FBL8)
Length = 374
Score = 32.0 bits (71), Expect = 2.7
Identities = 19/60 (31%), Positives = 31/60 (51%), Gaps = 6/60 (10%)
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSW------KTLFSDTHFANNHFLISTVYPQLVAC 80
LPEE++ +I LP+R L VC++W T++SD + + L + P L +C
Sbjct: 8 LPEEVLALIFRDLPLRDLAVATRVCRAWAAAAANSTVWSDKSISCDCELEDLLPPYLSSC 67
>ALG8_HUMAN (Q9BVK2) Probable dolichyl pyrophosphate Glc1Man9GlcNAc2
alpha-1,3-glucosyltransferase (EC 2.4.1.-)
(Dolichyl-P-Glc:Glc1Man9GlcNAc2-PP-dolichyl
glucosyltransferase) (HUSSY-02)
Length = 526
Score = 31.6 bits (70), Expect = 3.5
Identities = 29/98 (29%), Positives = 46/98 (46%), Gaps = 7/98 (7%)
Query: 23 PLPFLPEELIVIILLRL--PVRSLLRFKCVCKSWKTLFSDTHFANNHFLISTVYPQLVAC 80
PL F EL + ILL L + S+ K + + K LF+ + +L+ + P V C
Sbjct: 419 PLLFTAPELPIKILLMLLFTIYSISSLKTLFRKEKPLFN---WMETFYLLG-LGPLEVCC 474
Query: 81 ESVSAYRTWEIKTYPIESLLENSSTTVIPVSNTGHQRY 118
E V + +W++K YP LL S + V+ + Y
Sbjct: 475 EFVFPFTSWKVK-YPFIPLLLTSVYCAVGVTYAWFKLY 511
>YI60_THEMA (Q9X2H4) Hypothetical UPF0097 protein TM1860
Length = 187
Score = 31.2 bits (69), Expect = 4.6
Identities = 27/113 (23%), Positives = 48/113 (41%), Gaps = 11/113 (9%)
Query: 147 LKSKSSPTIDRFIYYGFGYDQVNHKYKLLAVKAFSRITETMIYTFGENSCKNVEVKDFPR 206
+K ++S I++ + GFG V+ + L + + E I E+ C+ V+ FP
Sbjct: 13 VKKQASEIIEKLMKRGFGATWVSEENMHLTLFFLGEVDEQKISEIAEHLCR--RVRGFPS 70
Query: 207 YPPNRKHLGKF---VSGTLNWIVDERDGRATILSFDIEKETYRQVLLPQHGYA 256
+ K G F +S + W+ E R L ++ E L HG++
Sbjct: 71 FSFTVKGFGYFKRKMSPRVFWLGVENTDRLMKLYEELRNE------LSHHGFS 117
>FLIR_AQUAE (O67773) Flagellar biosynthetic protein fliR
Length = 258
Score = 31.2 bits (69), Expect = 4.6
Identities = 14/55 (25%), Positives = 29/55 (52%), Gaps = 3/55 (5%)
Query: 320 FISENGVVLLMNTSSSQLILYNLNSRGLYFPRIPPSYLNIYYIEPKLDLHIYHES 374
F + G ++ ++ + I+ L F R+P NI+Y+ P++ L+ ++ES
Sbjct: 125 FFTLYGTLIFLSLGGGEFIILGLAES---FKRLPVGSFNIFYLNPEVFLNFFYES 176
>SLM9_SCHPO (O74309) Mitotic control protein slm9
Length = 807
Score = 30.8 bits (68), Expect = 6.0
Identities = 15/58 (25%), Positives = 26/58 (43%), Gaps = 4/58 (6%)
Query: 268 CICVCTSFLDTRWQLWMMKKYGVAESWTKLMSIPHENLLISNISLWPCVEPLFISENG 325
C V T L +W +K + S + + H N +S I+ P +E F+++ G
Sbjct: 553 CCIVSTGLL----YIWNIKNFEAIHSPVSTLPLFHSNFSVSKIARGPSIEQFFVTKQG 606
>PLR1_MOUSE (Q922V4) Pleiotropic regulator 1
Length = 513
Score = 30.8 bits (68), Expect = 6.0
Identities = 30/129 (23%), Positives = 47/129 (36%), Gaps = 30/129 (23%)
Query: 77 LVACESVSAYRTWEIKTYPIESLLENSSTTVIPVSNTGHQRYTILGSCNGFLCLYDNYQR 136
LV C S R W+++T L + V V + I GS +
Sbjct: 302 LVTCSRDSTARIWDVRTKASVHTLSGHTNAVATVRCQAAEPQIITGS----------HDT 351
Query: 137 CVRLWNPSINLKSKSSPTIDRFIYYGFGYDQVNHKYKLLAVKAFSRITETMIYTFGENSC 196
+RLW+ + K+ T+ NHK + AV + ++YTF S
Sbjct: 352 TIRLWD---LVAGKTRVTL------------TNHKKSVRAV-----VLHPLLYTFASGSP 391
Query: 197 KNVEVKDFP 205
N++ FP
Sbjct: 392 DNIKQWKFP 400
>FXLC_MOUSE (Q9EPX5) F-box/LRR-repeat protein 12 (F-box and
leucine-rich repeat protein 12) (F-box protein FBL12)
Length = 326
Score = 30.8 bits (68), Expect = 6.0
Identities = 14/34 (41%), Positives = 19/34 (55%)
Query: 27 LPEELIVIILLRLPVRSLLRFKCVCKSWKTLFSD 60
LP+ +++ I LPVR +R VC WK L D
Sbjct: 7 LPDLVLLEIFSYLPVRDRIRISRVCHRWKRLVDD 40
>FX34_HUMAN (Q9NWN3) F-box only protein 34
Length = 305
Score = 30.8 bits (68), Expect = 6.0
Identities = 20/71 (28%), Positives = 31/71 (43%), Gaps = 22/71 (30%)
Query: 7 NNTDSTLPVAETTANKP----------------------LPFLPEELIVIILLRLPVRSL 44
++ +STLPV E ++ K + FLP ++V I LP +SL
Sbjct: 130 SSVESTLPVLEASSWKKQVSHDFLETRFKIQQLLEPQQYMAFLPHHIMVKIFRLLPTKSL 189
Query: 45 LRFKCVCKSWK 55
+ KC C +K
Sbjct: 190 VALKCTCCYFK 200
>FX17_HUMAN (Q96EF6) F-box only protein 17 (F-box only protein 26)
Length = 278
Score = 30.8 bits (68), Expect = 6.0
Identities = 15/35 (42%), Positives = 24/35 (67%), Gaps = 1/35 (2%)
Query: 24 LPFLPEELIVIILLRLPVRSLL-RFKCVCKSWKTL 57
L LP EL+V +L +P RSL+ R + VC++W+ +
Sbjct: 18 LDALPPELLVQVLSHVPPRSLVTRCRPVCRAWRDI 52
>FX22_HUMAN (Q8NEZ5) F-box only protein 22 (F-box protein
FBX22p44)
Length = 403
Score = 30.4 bits (67), Expect = 7.8
Identities = 11/26 (42%), Positives = 18/26 (68%)
Query: 30 ELIVIILLRLPVRSLLRFKCVCKSWK 55
E++ +L LP ++LLR CVC+ W+
Sbjct: 28 EVVERVLTFLPAKALLRVACVCRLWR 53
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.138 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,526,796
Number of Sequences: 164201
Number of extensions: 1983096
Number of successful extensions: 4081
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 4073
Number of HSP's gapped (non-prelim): 15
length of query: 380
length of database: 59,974,054
effective HSP length: 112
effective length of query: 268
effective length of database: 41,583,542
effective search space: 11144389256
effective search space used: 11144389256
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC147013.3


