

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147011.10 - phase: 0
(165 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
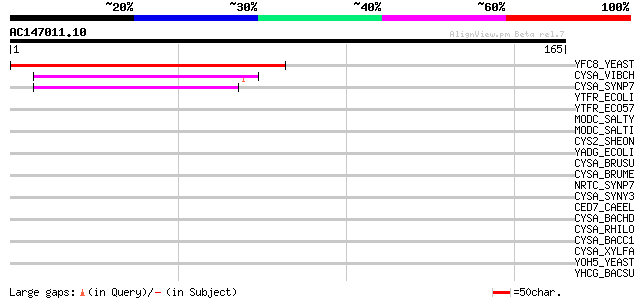
Score E
Sequences producing significant alignments: (bits) Value
YFC8_YEAST (P43569) Probable ATP-dependent transporter YFL028C 89 3e-18
CYSA_VIBCH (Q9KUI0) Sulfate/thiosulfate import ATP-binding prote... 42 6e-04
CYSA_SYNP7 (P14788) Sulfate/thiosulfate import ATP-binding prote... 42 8e-04
YTFR_ECOLI (Q6BEX0) Hypothetical ABC transporter ATP-binding pro... 40 0.002
YTFR_ECO57 (P63299) Hypothetical ABC transporter ATP-binding pro... 40 0.002
MODC_SALTY (Q8ZQR6) Molybdenum import ATP-binding protein modC (... 40 0.002
MODC_SALTI (Q8Z8A4) Molybdenum import ATP-binding protein modC (... 40 0.002
CYS2_SHEON (Q8E8K8) Sulfate/thiosulfate import ATP-binding prote... 40 0.002
YADG_ECOLI (P36879) Hypothetical ABC transporter ATP-binding pro... 40 0.002
CYSA_BRUSU (P63354) Sulfate/thiosulfate import ATP-binding prote... 40 0.002
CYSA_BRUME (P63353) Sulfate/thiosulfate import ATP-binding prote... 40 0.002
NRTC_SYNP7 (P38045) Nitrate transport ATP-binding protein nrtC 40 0.003
CYSA_SYNY3 (P74548) Sulfate/thiosulfate import ATP-binding prote... 40 0.003
CED7_CAEEL (P34358) ABC transporter ced-7 (Cell death protein 7) 40 0.003
CYSA_BACHD (Q9K876) Sulfate/thiosulfate import ATP-binding prote... 39 0.004
CYSA_RHILO (Q98K23) Sulfate/thiosulfate import ATP-binding prote... 39 0.005
CYSA_BACC1 (O31339) Sulfate/thiosulfate import ATP-binding prote... 39 0.005
CYSA_XYLFA (Q9PDN2) Sulfate/thiosulfate import ATP-binding prote... 39 0.007
YOH5_YEAST (Q08234) Probable ATP-dependent transporter YOL074C/Y... 38 0.009
YHCG_BACSU (P54591) Hypothetical ABC transporter ATP-binding pro... 38 0.009
>YFC8_YEAST (P43569) Probable ATP-dependent transporter YFL028C
Length = 289
Score = 89.4 bits (220), Expect = 3e-18
Identities = 42/82 (51%), Positives = 61/82 (74%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLLKP++VLLLDE+TVDLDV+ARA LL+FL+ E + R +++YATHIFDGL WP +
Sbjct: 161 MGLLKPWRVLLLDEVTVDLDVIARARLLEFLKWETETRRCSVVYATHIFDGLAKWPNQVY 220
Query: 61 YVAHGRLELAMPIEKVKETSKL 82
++ G++ + +K E S++
Sbjct: 221 HMKSGKIVDNLDYQKDVEFSEV 242
>CYSA_VIBCH (Q9KUI0) Sulfate/thiosulfate import ATP-binding protein
cysA (EC 3.6.3.25) (Sulfate-transporting ATPase)
Length = 376
Score = 42.0 bits (97), Expect = 6e-04
Identities = 25/68 (36%), Positives = 39/68 (56%), Gaps = 1/68 (1%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHGRL 67
+VLLLDE LD R +L ++LR DE G T ++ TH D + +V +++GR+
Sbjct: 156 EVLLLDEPFGALDAKVRKELRRWLRSLHDELGFTSVFVTHDQDEALELSDRVVVMSNGRI 215
Query: 68 E-LAMPIE 74
E + P+E
Sbjct: 216 EQIDSPVE 223
>CYSA_SYNP7 (P14788) Sulfate/thiosulfate import ATP-binding protein
cysA (EC 3.6.3.25) (Sulfate-transporting ATPase)
Length = 344
Score = 41.6 bits (96), Expect = 8e-04
Identities = 25/61 (40%), Positives = 33/61 (53%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHGRL 67
QVLLLDE LD R DL +LRK DE T ++ TH + + IV + HG++
Sbjct: 158 QVLLLDEPFGALDAKVRKDLRSWLRKLHDEVHVTTVFVTHDQEEAMEVADQIVVMNHGKV 217
Query: 68 E 68
E
Sbjct: 218 E 218
>YTFR_ECOLI (Q6BEX0) Hypothetical ABC transporter ATP-binding
protein ytfR
Length = 500
Score = 40.4 bits (93), Expect = 0.002
Identities = 35/133 (26%), Positives = 56/133 (41%), Gaps = 22/133 (16%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHGRL 67
+VL+LDE T LD +LL L ++ +RG ++I+ TH D + I + +G
Sbjct: 165 KVLILDEPTASLDT-QEVELLFDLMRQLRDRGVSLIFVTHFLDQVYQVSDRITVLRNG-- 221
Query: 68 ELAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKER---------------KAAGLP 112
+ET +L + V++ L +E D QR R K +
Sbjct: 222 ----SFVGCRETCELPQIELVKMMLGRELDTHALQRAGRTLLSDKPVAAFKNYGKKGTIA 277
Query: 113 EFDKQVDGSRVVG 125
FD +V +VG
Sbjct: 278 PFDLEVRPGEIVG 290
>YTFR_ECO57 (P63299) Hypothetical ABC transporter ATP-binding
protein ytfR
Length = 500
Score = 40.4 bits (93), Expect = 0.002
Identities = 35/133 (26%), Positives = 56/133 (41%), Gaps = 22/133 (16%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHGRL 67
+VL+LDE T LD +LL L ++ +RG ++I+ TH D + I + +G
Sbjct: 165 KVLILDEPTASLDT-QEVELLFDLMRQLRDRGVSLIFVTHFLDQVYQVSDRITVLRNG-- 221
Query: 68 ELAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKER---------------KAAGLP 112
+ET +L + V++ L +E D QR R K +
Sbjct: 222 ----SFVGCRETCELPQIELVKMMLGRELDTHALQRAGRTLLSDKPVAAFKNYGKKGTIA 277
Query: 113 EFDKQVDGSRVVG 125
FD +V +VG
Sbjct: 278 PFDLEVRPGEIVG 290
>MODC_SALTY (Q8ZQR6) Molybdenum import ATP-binding protein modC (EC
3.6.3.29)
Length = 352
Score = 40.4 bits (93), Expect = 0.002
Identities = 25/94 (26%), Positives = 48/94 (50%), Gaps = 6/94 (6%)
Query: 3 LLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYV 62
LL ++LLLDE LD+ + +LL +L++ E ++Y +H D + ++ +
Sbjct: 143 LLTAPELLLLDEPLASLDIPRKRELLPYLQRLAREINIPMLYVSHSLDEILHLADKVMVL 202
Query: 63 AHGRLELAMPIEKVKETSKLSLMRTVEVWLRKER 96
G+++ P+E+V +S + WL KE+
Sbjct: 203 EDGQVKAFGPLEEVWGSS------VMHPWLPKEQ 230
>MODC_SALTI (Q8Z8A4) Molybdenum import ATP-binding protein modC (EC
3.6.3.29)
Length = 352
Score = 40.4 bits (93), Expect = 0.002
Identities = 25/94 (26%), Positives = 48/94 (50%), Gaps = 6/94 (6%)
Query: 3 LLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYV 62
LL ++LLLDE LD+ + +LL +L++ E ++Y +H D + ++ +
Sbjct: 143 LLTAPELLLLDEPLASLDIPRKRELLPYLQRLAREINIPMLYVSHSLDEILHLADKVMVL 202
Query: 63 AHGRLELAMPIEKVKETSKLSLMRTVEVWLRKER 96
G+++ P+E+V +S + WL KE+
Sbjct: 203 EDGQVKAFGPLEEVWGSS------VMHPWLPKEQ 230
>CYS2_SHEON (Q8E8K8) Sulfate/thiosulfate import ATP-binding protein
cysA 2 (EC 3.6.3.25) (Sulfate-transporting ATPase 2)
Length = 354
Score = 40.4 bits (93), Expect = 0.002
Identities = 26/72 (36%), Positives = 39/72 (54%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHGRL 67
+VLLLDE LD RA+L ++LR+ DE T ++ TH + + IV + GR+
Sbjct: 156 KVLLLDEPFGALDAKVRAELRRWLRRLHDEINVTTVFVTHDQEEALEVADKIVVMNKGRI 215
Query: 68 ELAMPIEKVKET 79
E E+V +T
Sbjct: 216 EQQGTPEEVYDT 227
>YADG_ECOLI (P36879) Hypothetical ABC transporter ATP-binding
protein yadG
Length = 308
Score = 40.0 bits (92), Expect = 0.002
Identities = 22/60 (36%), Positives = 34/60 (56%), Gaps = 1/60 (1%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHGRL 67
++L+LDE T +D+ R + FL K+ +++G TII TH + E NI + HG L
Sbjct: 156 KLLILDEPTAGVDIELRRSMWGFL-KDLNDKGTTIILTTHYLEEAEMLCRNIGIIQHGEL 214
>CYSA_BRUSU (P63354) Sulfate/thiosulfate import ATP-binding protein
cysA (EC 3.6.3.25) (Sulfate-transporting ATPase)
Length = 359
Score = 40.0 bits (92), Expect = 0.002
Identities = 22/61 (36%), Positives = 35/61 (57%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHGRL 67
+VLLLDE LD R +L ++LR+ D+ G T ++ TH D + +V ++ GR+
Sbjct: 156 RVLLLDEPFGALDAQVRKELRRWLREIHDKTGHTTVFVTHDQDEALELADRVVVMSQGRI 215
Query: 68 E 68
E
Sbjct: 216 E 216
>CYSA_BRUME (P63353) Sulfate/thiosulfate import ATP-binding protein
cysA (EC 3.6.3.25) (Sulfate-transporting ATPase)
Length = 359
Score = 40.0 bits (92), Expect = 0.002
Identities = 22/61 (36%), Positives = 35/61 (57%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHGRL 67
+VLLLDE LD R +L ++LR+ D+ G T ++ TH D + +V ++ GR+
Sbjct: 156 RVLLLDEPFGALDAQVRKELRRWLREIHDKTGHTTVFVTHDQDEALELADRVVVMSQGRI 215
Query: 68 E 68
E
Sbjct: 216 E 216
>NRTC_SYNP7 (P38045) Nitrate transport ATP-binding protein nrtC
Length = 659
Score = 39.7 bits (91), Expect = 0.003
Identities = 31/110 (28%), Positives = 52/110 (47%), Gaps = 8/110 (7%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHG-- 65
++LLLDE LD L R +L + L + C E G T + TH D +V + +G
Sbjct: 158 KLLLLDEPFGALDALTRGNLQEQLMRICQEAGVTAVMVTHDVDEALLLSDRVVMLTNGPA 217
Query: 66 -RLELAMPIEKVKETSKLSLMRTVEVW-LRKE----RDEDRRQRKERKAA 109
++ + ++ + +L +M T + LR E + RR ++ KAA
Sbjct: 218 AQIGQILEVDFPRPRQRLEMMETPHYYDLRNELINFLQQQRRAKRRAKAA 267
>CYSA_SYNY3 (P74548) Sulfate/thiosulfate import ATP-binding protein
cysA (EC 3.6.3.25) (Sulfate-transporting ATPase)
Length = 355
Score = 39.7 bits (91), Expect = 0.003
Identities = 25/71 (35%), Positives = 38/71 (53%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHGRL 67
QVLLLDE LD R +L +LRK DE T ++ TH + + IV +++G++
Sbjct: 152 QVLLLDEPFGALDAKVRKELRAWLRKLHDEVHLTSVFVTHDQEEAMEVADEIVVMSNGKI 211
Query: 68 ELAMPIEKVKE 78
E E++ E
Sbjct: 212 EQVGTAEEIYE 222
>CED7_CAEEL (P34358) ABC transporter ced-7 (Cell death protein 7)
Length = 1704
Score = 39.7 bits (91), Expect = 0.003
Identities = 27/88 (30%), Positives = 44/88 (49%), Gaps = 2/88 (2%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
M L+ +V+LLDE T +D AR D+ K + +E R TI+ TH D E +
Sbjct: 691 MALIGDSEVVLLDEPTAGMDPGARQDVQKLVEREKANR--TILLTTHYMDEAERLGDWVF 748
Query: 61 YVAHGRLELAMPIEKVKETSKLSLMRTV 88
++HG+L + + +K+ + TV
Sbjct: 749 IMSHGKLVASGTNQYLKQKFGTGYLLTV 776
>CYSA_BACHD (Q9K876) Sulfate/thiosulfate import ATP-binding protein
cysA (EC 3.6.3.25) (Sulfate-transporting ATPase)
Length = 357
Score = 39.3 bits (90), Expect = 0.004
Identities = 26/74 (35%), Positives = 38/74 (51%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHGRL 67
+VLLLDE LD R DL K+LRK +E T ++ TH + D +V + G++
Sbjct: 156 KVLLLDEPFGALDAKVRKDLRKWLRKLHNEFQVTSVFVTHDQEEALDVSDRVVVMNQGKI 215
Query: 68 ELAMPIEKVKETSK 81
E ++V E K
Sbjct: 216 EQVGSPDEVYEQPK 229
>CYSA_RHILO (Q98K23) Sulfate/thiosulfate import ATP-binding protein
cysA (EC 3.6.3.25) (Sulfate-transporting ATPase)
Length = 346
Score = 38.9 bits (89), Expect = 0.005
Identities = 22/72 (30%), Positives = 38/72 (52%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHGRL 67
+VLLLDE LD R +L ++LR+ D G T ++ TH + + +V ++ GR+
Sbjct: 156 KVLLLDEPFGALDAQVRRELRRWLREIHDRTGHTTVFVTHDQEEALELADRVVVMSQGRI 215
Query: 68 ELAMPIEKVKET 79
E + + +T
Sbjct: 216 EQVGTADDIYDT 227
>CYSA_BACC1 (O31339) Sulfate/thiosulfate import ATP-binding protein
cysA (EC 3.6.3.25) (Sulfate-transporting ATPase)
Length = 357
Score = 38.9 bits (89), Expect = 0.005
Identities = 26/71 (36%), Positives = 37/71 (51%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHGRL 67
++LLLDE LD R +L ++LRK DE T ++ TH + D IV + GR+
Sbjct: 156 KILLLDEPFGALDAKVRKELRRWLRKLHDEFQITSVFVTHDQEEALDVADRIVVMNEGRI 215
Query: 68 ELAMPIEKVKE 78
E E+V E
Sbjct: 216 EQMGTPEEVYE 226
>CYSA_XYLFA (Q9PDN2) Sulfate/thiosulfate import ATP-binding protein
cysA (EC 3.6.3.25) (Sulfate-transporting ATPase)
Length = 348
Score = 38.5 bits (88), Expect = 0.007
Identities = 24/68 (35%), Positives = 36/68 (52%), Gaps = 1/68 (1%)
Query: 8 QVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIVYVAHGRL 67
+VLLLDE LD R DL ++LR+ D G T ++ TH + + + + GR+
Sbjct: 156 RVLLLDEPFGALDAQVRRDLRRWLREIHDRTGLTTVFVTHDQEEALELADRVAILNQGRI 215
Query: 68 E-LAMPIE 74
E +A P E
Sbjct: 216 EQVASPAE 223
>YOH5_YEAST (Q08234) Probable ATP-dependent transporter
YOL074C/YOL075C
Length = 1294
Score = 38.1 bits (87), Expect = 0.009
Identities = 21/45 (46%), Positives = 25/45 (54%)
Query: 3 LLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATH 47
LL +LLLDE T LD A +L+ L K C E+G TII H
Sbjct: 853 LLNDPPILLLDEPTSGLDSFTSATILEILEKLCREQGKTIIITIH 897
Score = 28.1 bits (61), Expect = 9.4
Identities = 14/39 (35%), Positives = 21/39 (52%)
Query: 9 VLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATH 47
++ LDE T LD + ++K L+K E G T I + H
Sbjct: 206 IMFLDEPTTGLDAYSAFLVIKTLKKLAKEDGRTFIMSIH 244
>YHCG_BACSU (P54591) Hypothetical ABC transporter ATP-binding
protein yhcG
Length = 232
Score = 38.1 bits (87), Expect = 0.009
Identities = 21/86 (24%), Positives = 41/86 (47%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
+ L + V+LLDE LD + R ++ L D ++ ATH D +E ++
Sbjct: 138 LALARRADVILLDEPFSGLDPMVRDSIVNSLVSYIDFEQQIVVIATHEIDEIETLLDEVI 197
Query: 61 YVAHGRLELAMPIEKVKETSKLSLMR 86
+A+G +E ++E +S+++
Sbjct: 198 ILANGEKVAQREVEDIREQEGMSVLQ 223
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.139 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,728,159
Number of Sequences: 164201
Number of extensions: 756372
Number of successful extensions: 2945
Number of sequences better than 10.0: 208
Number of HSP's better than 10.0 without gapping: 129
Number of HSP's successfully gapped in prelim test: 79
Number of HSP's that attempted gapping in prelim test: 2793
Number of HSP's gapped (non-prelim): 225
length of query: 165
length of database: 59,974,054
effective HSP length: 102
effective length of query: 63
effective length of database: 43,225,552
effective search space: 2723209776
effective search space used: 2723209776
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147011.10


