

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147006.6 + phase: 0 /pseudo
(797 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
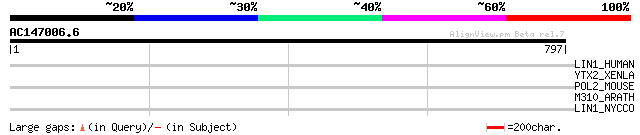
Score E
Sequences producing significant alignments: (bits) Value
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 44 0.001
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 36 0.46
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 34 1.7
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 33 3.9
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 32 6.6
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 44.3 bits (103), Expect = 0.001
Identities = 38/181 (20%), Positives = 80/181 (43%), Gaps = 30/181 (16%)
Query: 29 RVELLVEDIKMLDEKGEEGTLTE*EVGLHKGKLVELWRLLKAKDTLIT----QRSRSRWL 84
+++ L+ +K L+++ + + + +++++ LK +T T SRS +
Sbjct: 312 KIDTLISQLKELEKQEQTNSKAS-----RRQEIIKIRAELKEIETQKTLQKINESRSWFF 366
Query: 85 KKGDANSKFFHKCIKLRKSINSIKALEENEGWVVS-PFEVCRKVVNYFTNHFAE------ 137
+K + + + IK ++ N I ++ + G + + P E+ + Y+ + +A
Sbjct: 367 EKINKIDRPLARLIKKKREKNQIDTIKNDRGDITTDPTEIQTTIREYYKHLYANKLENLE 426
Query: 138 ------DRWDRPRLDRVDFESLTEVENGLLVAPFSLLEIEEVVRDSDGGKSPGPNGYNFS 191
D + PRL++ + ESL P + EIE ++ KSPGP G+
Sbjct: 427 EMDKFLDTYTLPRLNQEEVESLNR--------PITSSEIEAIINSLPNKKSPGPEGFTAE 478
Query: 192 F 192
F
Sbjct: 479 F 479
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 35.8 bits (81), Expect = 0.46
Identities = 42/187 (22%), Positives = 79/187 (41%), Gaps = 8/187 (4%)
Query: 11 LKVILKE*IKKEYGGMEDRVELLVEDIKMLDEK--GEEGTLTE*EVGLHKGKLVELWRLL 68
LK++ +E K G +E L ++ L+++ G E + E L + + +
Sbjct: 293 LKLLCQEYTKSVSGQRNAEIEALNGEVLDLEQRLSGSEDQALQCEY-LERKEALRNMEQR 351
Query: 69 KAKDTLITQRSRSRWLKKGDANSKFFHKCIKLRKSINSIKALEENEGWVVSPFEVCR-KV 127
+A+ + RSR + L D S+FF+ K + + I L +G + E R +
Sbjct: 352 QARGAFV--RSRMQLLCDMDRGSRFFYALEKKKGNRKQITCLFAEDGTPLEDPEAIRDRA 409
Query: 128 VNYFTNHFAEDRWDRPRLDRV--DFESLTEVENGLLVAPFSLLEIEEVVRDSDGGKSPGP 185
+++ N F+ D + + ++E L P +L E+ + +R KSPG
Sbjct: 410 RSFYQNLFSPDPISPDACEELWDGLPVVSERRKERLETPITLDELSQALRLMPHNKSPGL 469
Query: 186 NGYNFSF 192
+G F
Sbjct: 470 DGLTIEF 476
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 33.9 bits (76), Expect = 1.7
Identities = 32/142 (22%), Positives = 56/142 (38%), Gaps = 17/142 (11%)
Query: 67 LLKAKDTLITQRSRSRWLKKGDANSKFFHKCIKLRKSINSI-----------KALEENEG 115
++K + + +R + S FF K K+ K + + K E
Sbjct: 366 IIKLRGEINQVETRRTIQRINQTRSWFFEKINKIDKPLARLTKGHRDKILINKIRNEKGD 425
Query: 116 WVVSPFEVCRKVVNYFTNHFAE-----DRWDRPRLDRVDFESLTEVENGLLVAPFSLLEI 170
P E+ + +++ ++ D D+ LDR L + + L +P S EI
Sbjct: 426 ITTDPEEIQNTIRSFYKRLYSTKLENLDEMDK-FLDRYQVPKLNQDQVDHLNSPISPKEI 484
Query: 171 EEVVRDSDGGKSPGPNGYNFSF 192
E V+ KSPGP+G++ F
Sbjct: 485 EAVINSLPTKKSPGPDGFSAEF 506
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 32.7 bits (73), Expect = 3.9
Identities = 14/36 (38%), Positives = 22/36 (60%), Gaps = 1/36 (2%)
Query: 549 KFYGEEWEGVKKISWVKWELVCQNKR-NGSLGVNSL 583
+F+ E +KISWV W+ +C++K +G LG L
Sbjct: 27 EFWWSSCENKRKISWVAWQKLCKSKEDDGGLGFRDL 62
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 32.0 bits (71), Expect = 6.6
Identities = 23/99 (23%), Positives = 47/99 (47%), Gaps = 8/99 (8%)
Query: 99 KLRKSINSIKALEENEGWVVSPFEVCRKVVNYFTNHFAEDRWDRPR-----LDRVDFESL 153
+++ I+SI+ N+ P E+ +K++N + +++ + L+ L
Sbjct: 384 RVKSLISSIR--NGNDEITTDPSEI-QKILNEYYKKLYSHKYENLKEIDQYLEACHLPRL 440
Query: 154 TEVENGLLVAPFSLLEIEEVVRDSDGGKSPGPNGYNFSF 192
++ E +L P S EI +++ KSPGP+G+ F
Sbjct: 441 SQKEVEMLNRPISSSEIASTIQNLPKKKSPGPDGFTSEF 479
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.356 0.160 0.625
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 84,954,078
Number of Sequences: 164201
Number of extensions: 3251514
Number of successful extensions: 14212
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 14211
Number of HSP's gapped (non-prelim): 5
length of query: 797
length of database: 59,974,054
effective HSP length: 118
effective length of query: 679
effective length of database: 40,598,336
effective search space: 27566270144
effective search space used: 27566270144
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 70 (31.6 bits)
Medicago: description of AC147006.6


