

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147002.8 + phase: 0 /pseudo
(1664 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
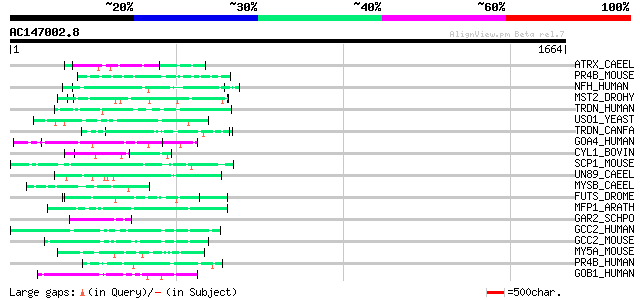
Score E
Sequences producing significant alignments: (bits) Value
ATRX_CAEEL (Q9U7E0) Transcriptional regulator ATRX homolog (X-li... 62 1e-08
PR4B_MOUSE (Q61136) Serine/threonine-protein kinase PRP4 homolog... 59 1e-07
NFH_HUMAN (P12036) Neurofilament triplet H protein (200 kDa neur... 58 2e-07
MST2_DROHY (Q08696) Axoneme-associated protein mst101(2) 57 4e-07
TRDN_HUMAN (Q13061) Triadin 55 1e-06
USO1_YEAST (P25386) Intracellular protein transport protein USO1 55 2e-06
TRDN_CANFA (P82179) Triadin 53 8e-06
GOA4_HUMAN (Q13439) Golgi autoantigen, golgin subfamily A member... 53 8e-06
CYL1_BOVIN (P35662) Cylicin I (Multiple-band polypeptide I) 52 1e-05
SCP1_MOUSE (Q62209) Synaptonemal complex protein 1 (SCP-1 protein) 52 1e-05
UN89_CAEEL (O01761) Muscle M-line assembly protein unc-89 (Uncoo... 51 3e-05
MYSB_CAEEL (P02566) Myosin heavy chain B (MHC B) 51 3e-05
FUTS_DROME (Q9W596) Microtubule-associated protein futsch 51 3e-05
MFP1_ARATH (Q9LW85) MAR binding filament-like protein 1 50 4e-05
GAR2_SCHPO (P41891) Protein gar2 50 5e-05
GCC2_HUMAN (Q8IWJ2) GRIP and coiled-coil domain-containing prote... 49 9e-05
GCC2_MOUSE (Q8CHG3) GRIP and coiled-coil domain-containing prote... 49 2e-04
MY5A_MOUSE (Q99104) Myosin Va (Myosin 5A) (Dilute myosin heavy c... 48 2e-04
PR4B_HUMAN (Q13523) Serine/threonine-protein kinase PRP4 homolog... 47 3e-04
GOB1_HUMAN (Q14789) Golgi autoantigen, golgin subfamily B member... 47 3e-04
>ATRX_CAEEL (Q9U7E0) Transcriptional regulator ATRX homolog
(X-linked nuclear protein-1)
Length = 1359
Score = 62.0 bits (149), Expect = 1e-08
Identities = 73/276 (26%), Positives = 119/276 (42%), Gaps = 35/276 (12%)
Query: 188 NEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDE-----DQSVKMAMLSNK 242
+E E KKSKS + K KS K S S +E D + ++ + K S
Sbjct: 96 DEEEDRKKSKSKKKVDQKKKEKSKKKRTTSSSEDEDSDEEREQKSKKKSKKTKKQTSSES 155
Query: 243 LEYLARKQKKFLSKRGSYKNFKK---------EDQKGCFNCKKPGHFIADCPDLQKEKFK 293
E ++K SK+ K+ KK ED+K KK K+K K
Sbjct: 156 SEESEEERKVKKSKKNKEKSVKKRAETSEESDEDEKPSKKSKKG----------LKKKAK 205
Query: 294 GKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSE 353
+S+ S + + +K KKS ++ +SE +EA + K + SSE SE
Sbjct: 206 SESESESEDEKEVKKSKKKSKKVVKKESESE-----DEAPEKKKTEKRKRSKTSSEESSE 260
Query: 354 AE-SDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLEL 412
+E SD E+E + S P+++ ++K+L S E +++ L +K K+ TL+
Sbjct: 261 SEKSDEEEEEKESSPKPKKKKPLAVKKLSSDEESEESDVEVLPQK----KKRGAVTLISD 316
Query: 413 KASEEELKGFNLISATYEDRLKSLCQKLQEKCDKGS 448
E++ K + S E+++ K QE + GS
Sbjct: 317 SEDEKDQKSESEAS-DVEEKVSKKKAKKQESSESGS 351
Score = 53.9 bits (128), Expect = 4e-06
Identities = 97/438 (22%), Positives = 174/438 (39%), Gaps = 54/438 (12%)
Query: 164 EAKDLNTLSVEDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEES 223
E D + + EDL + ++ + K K + +K KAS S E+
Sbjct: 9 EDSDGHVIEDEDLEMARQIENERKEKRAQKLKEKREREGKPPPKKRPAKKRKASSSEEDD 68
Query: 224 PDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIA- 282
D + +S K + K E + + + ++ S K+ KK DQK KK +
Sbjct: 69 DDEEESPRKSSKKSRKRAKSESESDESDEEEDRKKS-KSKKKVDQKKKEKSKKKRTTSSS 127
Query: 283 ---DCPDLQKEKFKGKSKKS-------SFNSSKFRKQIKKSLMATWEDLDSESGSDKEEA 332
D + +++K K KSKK+ S S+ +++KKS E + EE+
Sbjct: 128 EDEDSDEEREQKSKKKSKKTKKQTSSESSEESEEERKVKKS-KKNKEKSVKKRAETSEES 186
Query: 333 DDDAKAAVGLVATVSSEAVSEAESDSEDENEV-YSKIPRQELVDSLKELLSLFEHRTNEL 391
D+D K + + +A SE+ES+SEDE EV SK +++V E +E
Sbjct: 187 DEDEKPSKKSKKGLKKKAKSESESESEDEKEVKKSKKKSKKVVKKESE---------SED 237
Query: 392 TDLKEKYVDLMKQQKSTLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDKGSGNK 451
++K + K+ K++ E SE+ + +E+ + K
Sbjct: 238 EAPEKKKTEKRKRSKTSSEESSESEKS-------------------DEEEEEKESSPKPK 278
Query: 452 HEIALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTF 511
+ L ++ + S+ + + K K + + E KS + ++
Sbjct: 279 KKKPLAVKKLSSDEESEESDVEVLPQKKKRGAVTLISDSEDEKDQKSESE-------ASD 331
Query: 512 VPEGTNAKTAVQSKPEASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLI 571
V E + K A + + SGS + S N K K K P+ +K I+ S + + I
Sbjct: 332 VEEKVSKKKAKKQESSESGSDSSEGSITVNRKSK--KKEKPEKKKKGIIMDSSKLQKETI 389
Query: 572 ---KPESKIPKQKDQKNK 586
+ E + K+ ++K K
Sbjct: 390 DAERAEKERRKRLEKKQK 407
Score = 37.0 bits (84), Expect = 0.45
Identities = 69/331 (20%), Positives = 119/331 (35%), Gaps = 94/331 (28%)
Query: 290 EKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSE 349
+K K K ++ K R K+ ++ ED D E S ++ + K
Sbjct: 37 QKLKEKREREGKPPPKKRPAKKRKASSSEEDDDDEEESPRKSSKKSRK-----------R 85
Query: 350 AVSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTL 409
A SE+ESD DE E K +K VD K++KS
Sbjct: 86 AKSESESDESDEEEDRKK-------------------------SKSKKKVDQKKKEKSKK 120
Query: 410 LELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGIDRSKV 469
+S E+ ED + QK ++K K K + + + + S+
Sbjct: 121 KRTTSSSED-----------EDSDEEREQKSKKKSKK---TKKQTSSESS-----EESEE 161
Query: 470 ASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKPEAS 529
+ + KNK K + E S+E + K + +SK
Sbjct: 162 ERKVKKSKKNKEKSVKKRAETSEE--------------------SDEDEKPSKKSK---- 197
Query: 530 GSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLIKPESKIP-----KQKDQK 584
+ LK K ++S+ +S+ K +K+S+ + ++K ES+ K+K +K
Sbjct: 198 ----------KGLKKKAKSESESESEDEKEVKKSKKKSKKVVKKESESEDEAPEKKKTEK 247
Query: 585 NKAATASEKTIPKGVKPKVLNDQKPLSIHPK 615
K + S + + K ++K S PK
Sbjct: 248 RKRSKTSSEESSESEKSDEEEEEKESSPKPK 278
>PR4B_MOUSE (Q61136) Serine/threonine-protein kinase PRP4 homolog
(EC 2.7.1.37) (PRP4 pre-mRNA processing factor 4
homolog) (Pre-mRNA protein kinase)
Length = 1007
Score = 58.9 bits (141), Expect = 1e-07
Identities = 105/464 (22%), Positives = 181/464 (38%), Gaps = 58/464 (12%)
Query: 202 PSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYK 261
PS + ++ A + +SV E +G+ EDQS NK +K+ K SK +K
Sbjct: 7 PSLREQAEMDDADNSEKSVNEE-NGEVSEDQS------QNKHSRHKKKKHKHRSKHKKHK 59
Query: 262 NFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDL 321
+ +ED+ KK H +KE + K+ + K+ K +A EDL
Sbjct: 60 HSSEEDRD-----KKHKHKHKHKKHKRKEVIEASDKEGLSPA----KRTKLDDLALLEDL 110
Query: 322 DSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPRQELVDSLKELL 381
+ + K E D++ G V + + ES SE+E E++ K +
Sbjct: 111 EKQRALIKAELDNELME--GKVQSGMGLILQGYESGSEEEGEIHEKARN----GNRSSTR 164
Query: 382 SLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASEEELKGFNLISATYEDRLKSLCQKLQ 441
S E+TD K K + ++ + + KG + +R KS ++
Sbjct: 165 SSSTRGKLEITDNKNSAKKRSKSRSKERTRHRSDKRKSKGAGEMLREKANRSKSKERRKS 224
Query: 442 EKCDKGSGNKHEIALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCD 501
+ K S ++ + +SK + + + GK +EEK K
Sbjct: 225 KSPSKRSKSQDQAR----------KSKSPPLRRRSQEKVGKARSPAEEKMKSEEKGK--- 271
Query: 502 CIKDGLKSTFVPEGTNAKTAVQSKPEASGSQAKITSKPENLKIKVMTKSDPKSQKIKILK 561
IKD KS V N ++ +SK S + SK + K + KS++ +I K
Sbjct: 272 -IKDRKKSPIV----NERSRDRSKKSKSPVDLRDKSKDRRSRSK-----ERKSKRSEIDK 321
Query: 562 RSEPVHQNLIKPESKIPKQKDQKNKAATASEKTIPKGVKPKVLNDQKPLSIHPKVQGRKS 621
+P+ P K+ ++ + + + PK L D+ S P + R+S
Sbjct: 322 EKKPIK----SPSKDASSGKENRSPSRRPGRSPKRRSLSPK-LRDKSRRSRSPLLNDRRS 376
Query: 622 KTSKANPKGPMKIWVPKSELAKNAGVLKGKRE---TKVMVPRQR 662
K SK+ P + P AK+ + + +RE ++ PR R
Sbjct: 377 KQSKS----PSRTLSP-GRRAKSRSLERKRREPERRRLSSPRTR 415
>NFH_HUMAN (P12036) Neurofilament triplet H protein (200 kDa
neurofilament protein) (Neurofilament heavy polypeptide)
(NF-H)
Length = 1026
Score = 57.8 bits (138), Expect = 2e-07
Identities = 116/536 (21%), Positives = 209/536 (38%), Gaps = 56/536 (10%)
Query: 158 KVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKAS 217
KV + E+ K + E ++ + E + E T ++ K A +GK + +
Sbjct: 437 KVKSEEKIKVVEKSEKETVIVEEQTEETQVTEEVTEEEEKE-AKEEEGKEEEGGE----- 490
Query: 218 ESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKP 277
EE +G +E +S ++ + K+ K+ +KE+ K K P
Sbjct: 491 ---EEEAEGGEEETKSPPAEEAASPEKEAKSPVKEEAKSPAEAKSPEKEEAKSPAEVKSP 547
Query: 278 GHFIADCPDLQKEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAK 337
A P ++ K ++K +K ++K A + + ++ + AK
Sbjct: 548 EK--AKSPAKEEAKSPPEAKSPEKEEAKSPAEVKSPEKAKSPAKEEAKSPAEAKSPEKAK 605
Query: 338 AAVGLVATVSSEAVS----EAESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTD 393
+ V A +EA S EA+S +E ++ +K P +E S ++ S + ++ E +
Sbjct: 606 SPVKEEAKSPAEAKSPVKEEAKSPAEVKSPEKAKSPTKEEAKSPEKAKSPEKAKSPEKEE 665
Query: 394 LKEKYVDLMKQQKSTLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDKG-SGNKH 452
K ++ KS + S E+ K A ++ KS ++ + +K S K
Sbjct: 666 AKSP-----EKAKSPVKAEAKSPEKAKSPVKAEAKSPEKAKSPVKEEAKSPEKAKSPVKE 720
Query: 453 EIALDDFIMAGI-DRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTF 511
E + + + + +K S K + K S EK+K
Sbjct: 721 EAKSPEKAKSPVKEEAKTPEKAKSPVKEEAK----SPEKAKS------------------ 758
Query: 512 VPEGTNAKTAVQSKPEASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLI 571
PE AKT PEA + AK ++ K KS K + K P+ ++
Sbjct: 759 -PE--KAKTLDVKSPEAK-TPAKEEARSPADKFPEKAKSPVKEEVKSPEKAKSPLKEDAK 814
Query: 572 KPESKIPKQKDQKNKAATASEKTIPKGVKPKVLNDQKPLSIHPKVQGRKSKTSKANPKGP 631
PE +IPK+++ K+ + K +P +++ PK + +K + PK
Sbjct: 815 APEKEIPKKEEVKSPVKEEEKPQEVKVKEPPKKAEEEKAPATPKTEEKKDSKKEEAPK-- 872
Query: 632 MKIWVPKSELAKNAGVLKGKRETKVMVPRQRMFKAHDWRESFVPYPYNERWRRSEV 687
+ PK E K V K K E+KV ++ +A D ++ VP P E + EV
Sbjct: 873 KEAPKPKVEEKKEPAVEKPK-ESKVEAKKE---EAEDKKK--VPTPEKEAPAKVEV 922
Score = 50.4 bits (119), Expect = 4e-05
Identities = 103/480 (21%), Positives = 181/480 (37%), Gaps = 46/480 (9%)
Query: 189 EHETSKKSKSIALPSKGKTSKSSKAYKASE--SVEESPDGDSDEDQSVKMAMLSNKLEYL 246
E E +K + P K K+ +A +E S E++ +E +S A K E
Sbjct: 568 EKEEAKSPAEVKSPEKAKSPAKEEAKSPAEAKSPEKAKSPVKEEAKSPAEAKSPVKEEAK 627
Query: 247 ARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSKKSSFNSSKF 306
+ + K K K+ KE+ K K P A P+ ++ K K+K +K
Sbjct: 628 SPAEVKSPEKA---KSPTKEEAKSPEKAKSPEK--AKSPEKEEAKSPEKAKSPVKAEAKS 682
Query: 307 RKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYS 366
++ K + A + + KEEA KA + V EA S ++ S + E +
Sbjct: 683 PEKAKSPVKAEAKSPEKAKSPVKEEAKSPEKAK----SPVKEEAKSPEKAKSPVKEE--A 736
Query: 367 KIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKAS----------- 415
K P + +E S + ++ E K K +D+ + T + +A
Sbjct: 737 KTPEKAKSPVKEEAKSPEKAKSPE----KAKTLDVKSPEAKTPAKEEARSPADKFPEKAK 792
Query: 416 ---EEELKGFNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGIDRSKVASM 472
+EE+K + ++ K+ +++ +K + S K E + + +
Sbjct: 793 SPVKEEVKSPEKAKSPLKEDAKAPEKEIPKKEEVKSPVKEEEKPQEVKVKEPPKKAEEEK 852
Query: 473 IYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKPEASGSQ 532
+T K + K EE K+ + K + K V + +K V++K E + +
Sbjct: 853 APATPKTEEKKDSKKEEAPKKEAPKPKVE----EKKEPAVEKPKESK--VEAKKEEAEDK 906
Query: 533 AKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLIKPESKIPKQKDQKNKAATASE 592
K+ + + KV K D K ++ + + EP +P K++ K T E
Sbjct: 907 KKVPTPEKEAPAKVEVKEDAKPKEKTEVAKKEPDDAKAKEPSKPAEKKEAAPEKKDTKEE 966
Query: 593 KTIPKGVKP----KVLNDQKPLSIHP-----KVQGRKSKTSKANPKGPMKIWVPKSELAK 643
K KP K D K LS P + + S T + + K P K K+ K
Sbjct: 967 KAKKPEEKPKTEAKAKEDDKTLSKEPSKPKAEKAEKSSSTDQKDSKPPEKATEDKAAKGK 1026
>MST2_DROHY (Q08696) Axoneme-associated protein mst101(2)
Length = 1391
Score = 57.0 bits (136), Expect = 4e-07
Identities = 118/559 (21%), Positives = 208/559 (37%), Gaps = 66/559 (11%)
Query: 142 DHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHETSKKSKSIAL 201
+ ++K ++ + + K A +E + E+L +K ET+KK K +A
Sbjct: 432 EELAKNIKKAAEKKKCKEAAKKEKEAAERKKCEELAKKIKKAAEKKKCEETAKKGKEVA- 490
Query: 202 PSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYK 261
+ K + +K K +E ++ E ++ + K E A+K+K+ K+ K
Sbjct: 491 -ERKKCEELAKKIKKAEIKKKCKKLAKKEKETAE----KKKCEKAAKKRKEAAEKKKCEK 545
Query: 262 NFKKE----DQKGCFNCKKPGHFIAD---CPDLQKEKFKGKSKKSSFNSSKFRKQI---- 310
KK ++K C K A+ C KE+ + KK ++K K++
Sbjct: 546 AAKKRKEAAEKKKCEKSAKKRKEAAEKKKCEKAAKERKEAAEKKKCEEAAKKEKEVAERK 605
Query: 311 KKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPR 370
K +A +E KE A + +AA ++ + +A E + K+ +
Sbjct: 606 KCEELAKKIKKAAEKKKCKEAAKKEKEAAEREKCGELAKKIKKAA-----EKKKCKKLAK 660
Query: 371 QELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASE---------EELKG 421
+E + K+ + E + K+K + K++K + K E E K
Sbjct: 661 KEKETAEKKKCEKAAKKRKEAAE-KKKCAEAAKKEKEAAEKKKCEEAAKKEKEAAERKKC 719
Query: 422 FNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGIDRSKVASMIYSTYKNKG 481
L + K C+KL +K G NK + A ++ K + K
Sbjct: 720 EELAKKIKKAAEKKKCKKLAKKKKAGEKNKLKKGNKKGKKALKEKKKCRELAKKKAAEK- 778
Query: 482 KGIGYSEEKSKEYSLKSYCD--------------CIKDGLKSTFVPEGTNA-KTAVQSKP 526
K + +K KE + K C+ C K K E K A + K
Sbjct: 779 KKCKEAAKKEKEAAEKKKCEKTAKKRKEEAEKKKCEKTAKKRKEAAEKKKCEKAAKKRKE 838
Query: 527 EASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLIKPESKIPKQKDQKNK 586
EA + + T+K K ++ K + KR + + + +K K+ +K K
Sbjct: 839 EAEKKKCEKTAK------KRKETAEKKKCEKAAKKRKQAAEKKKCEKAAKKRKEAAEKKK 892
Query: 587 AATASEKTIPKGVKPK---VLNDQKPLSIHPKVQGRKSKTSKANPKGPMKIWV------- 636
A A++K K K +K ++ K + K KA K K
Sbjct: 893 CAEAAKKEKELAEKKKCEEAAKKEKEVAERKKCEELAKKIKKAAEKKKCKKLAKKEKKAG 952
Query: 637 PKSELAKNAGVLKGKRETK 655
K++L K AG KGK++ K
Sbjct: 953 EKNKLKKKAG--KGKKKCK 969
Score = 53.1 bits (126), Expect = 6e-06
Identities = 101/470 (21%), Positives = 167/470 (35%), Gaps = 57/470 (12%)
Query: 191 ETSKKSKSIALPSK-GKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLARK 249
+ +KK K A K + +K K + EE+ + + + K L+ K++ A K
Sbjct: 879 KAAKKRKEAAEKKKCAEAAKKEKELAEKKKCEEAAKKEKEVAERKKCEELAKKIKKAAEK 938
Query: 250 QK-----KFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSKKSSFNSS 304
+K K K G KK+ KG CKK G + +K K +K +
Sbjct: 939 KKCKKLAKKEKKAGEKNKLKKKAGKGKKKCKKLGKKSKRAAEKKKCAEAAKKEKEAATKK 998
Query: 305 KFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEV 364
K ++ KK + ++K++ ++ AK + + E ++ E
Sbjct: 999 KCEERAKK----------QKEAAEKKQCEERAK------KLKEAAEQKQCEERAKKLKEA 1042
Query: 365 YSKIPRQELVDSLKELL--SLFEHRTNELTDLKE-KYVDLMKQQKSTLLELKASEEELKG 421
K +E LKE E R +L + E K + +++ E K EE K
Sbjct: 1043 AEKKQCEERAKKLKEAAEQKQCEERAKKLKEAAEKKQCEERAKKEKEAAEKKQCEERAK- 1101
Query: 422 FNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGIDRSKVASMIYSTYKNKG 481
+LK +K Q C++ + + E A R K A+ +
Sbjct: 1102 ----------KLKEAAEKKQ--CEERAKKEKEAAEKKRCEEAAKREKEAA--------EK 1141
Query: 482 KGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKPEASGSQAKITSKPEN 541
K + +K KE + K C + K E K A +K E +Q K K +
Sbjct: 1142 KKCAEAAKKEKEATEKQ--KCAEAAKKEKEAAE--KKKCAEAAKREKEAAQKK---KCAD 1194
Query: 542 LKIKVMTKSDPKSQKIKILKRSEPVHQNLIKPESKIPKQKDQKNKAATASEKTIPKGVKP 601
L K ++ K + K E + +K K+ +K K A A++K K
Sbjct: 1195 LAKKEQEPAEMKKCEEAAKKEKEAAEKQKCAKAAKKEKEAAEKKKCAEAAKKEQEAAEKK 1254
Query: 602 KVLNDQKPLSIHPKVQGRKSKTSKANPKGPMKIWVPKSELAKNAGVLKGK 651
K K K +K K KA +K K + L+ K
Sbjct: 1255 KCAEAAK----KEKEAEKKRKCEKAEKAAALKRQCAKLVIRAKEAALRKK 1300
Score = 47.0 bits (110), Expect = 4e-04
Identities = 110/525 (20%), Positives = 189/525 (35%), Gaps = 69/525 (13%)
Query: 174 EDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDED-- 231
ED + K E + KK + A K K ++ K KA++ +E+ + E+
Sbjct: 361 EDEKKACKELAKKKKEADEKKKCEEAANKEK-KAAEKKKCEKAAKERKEAAEKKKCEEAA 419
Query: 232 QSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKE----DQKGCFNCKKPGHFIADCPDL 287
+ K A K E LA+ KK K+ + KKE ++K C K A+
Sbjct: 420 KKEKEAAERKKCEELAKNIKKAAEKKKCKEAAKKEKEAAERKKCEELAKKIKKAAEKKKC 479
Query: 288 QKEKFKGKSKKSSFNSSKFRKQIKKSLM------------ATWEDLDSESGSDK------ 329
++ KGK + K+IKK+ + T E E + K
Sbjct: 480 EETAKKGKEVAERKKCEELAKKIKKAEIKKKCKKLAKKEKETAEKKKCEKAAKKRKEAAE 539
Query: 330 -----------EEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPRQELVDSLK 378
+EA + K + + E +++ E K +E K
Sbjct: 540 KKKCEKAAKKRKEAAEKKKCEKSAKKRKEAAEKKKCEKAAKERKEAAEKKKCEEAAKKEK 599
Query: 379 ELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASEEELKG---FNLISATYEDRLKS 435
E+ + EL +K + K +++ E +A+E E G + A + + K
Sbjct: 600 EVAE--RKKCEELAKKIKKAAEKKKCKEAAKKEKEAAEREKCGELAKKIKKAAEKKKCKK 657
Query: 436 LCQKLQE-----KCDKGSGNKHEIALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSEEK 490
L +K +E KC+K + + E A + K A+ + K + +K
Sbjct: 658 LAKKEKETAEKKKCEKAAKKRKEAAEKKKCAEAAKKEKEAA--------EKKKCEEAAKK 709
Query: 491 SKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKPEASGSQAKITSKPENLKIKVMTKS 550
KE + + C+ + +K K +K + +G + K+ K N K K K+
Sbjct: 710 EKEAAERKKCEELAKKIKKA----AEKKKCKKLAKKKKAGEKNKL--KKGNKKGK---KA 760
Query: 551 DPKSQKIKILKRSEPVHQNLIKPESKIPKQKDQKNKAATASEKTIPKGVKPKVLNDQKPL 610
+ +K + L + + + K +K K+ +K K ++K + K K K
Sbjct: 761 LKEKKKCRELAKKKAAEKKKCKEAAKKEKEAAEKKKCEKTAKKRKEEAEKKKCEKTAK-- 818
Query: 611 SIHPKVQGRKSKTSKANPKGPMKIWVPKSELAKNAGVLKGKRETK 655
K K K KA K K K + K A K E K
Sbjct: 819 --KRKEAAEKKKCEKAAKK--RKEEAEKKKCEKTAKKRKETAEKK 859
Score = 38.1 bits (87), Expect = 0.20
Identities = 82/433 (18%), Positives = 159/433 (35%), Gaps = 26/433 (6%)
Query: 191 ETSKKSKSIALPSK----GKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLS-----N 241
E +KK K A K K K + K E + + +++ + K+A N
Sbjct: 689 EAAKKEKEAAEKKKCEEAAKKEKEAAERKKCEELAKKIKKAAEKKKCKKLAKKKKAGEKN 748
Query: 242 KLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSKKSSF 301
KL+ +K KK L ++ + K+ CK+ + + +K + K +K
Sbjct: 749 KLKKGNKKGKKALKEKKKCRELAKKKAAEKKKCKEAAKKEKEAAEKKKCEKTAKKRKEEA 808
Query: 302 NSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDE 361
K K KK A + ++ ++E + K + + E ++
Sbjct: 809 EKKKCEKTAKKRKEAAEKKKCEKAAKKRKEEAEKKKCEKTAKKRKETAEKKKCEKAAKKR 868
Query: 362 NEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASEEELKG 421
+ K ++ KE E + KEK + K+ + + K E K
Sbjct: 869 KQAAEKKKCEKAAKKRKEAA---EKKKCAEAAKKEKELAEKKKCEEAAKKEKEVAERKKC 925
Query: 422 FNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGIDRSKVASM-IYSTYKNK 480
L + K C+KL +K +K +G K+++ AG + K + S +
Sbjct: 926 EELAKKIKKAAEKKKCKKLAKK-EKKAGEKNKLKK----KAGKGKKKCKKLGKKSKRAAE 980
Query: 481 GKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKPEASGSQAKITSKPE 540
K + +K KE + K C+ + K E + + EA+ Q + + +
Sbjct: 981 KKKCAEAAKKEKEAATKKKCE--ERAKKQKEAAEKKQCEERAKKLKEAA-EQKQCEERAK 1037
Query: 541 NLKIKVMTKS-DPKSQKIKILKRSEPVHQNLIK----PESKIPKQKDQKNKAATASEKTI 595
LK K + +++K+K + + K E K +++ +K K A ++
Sbjct: 1038 KLKEAAEKKQCEERAKKLKEAAEQKQCEERAKKLKEAAEKKQCEERAKKEKEAAEKKQCE 1097
Query: 596 PKGVKPKVLNDQK 608
+ K K ++K
Sbjct: 1098 ERAKKLKEAAEKK 1110
>TRDN_HUMAN (Q13061) Triadin
Length = 728
Score = 55.5 bits (132), Expect = 1e-06
Identities = 121/562 (21%), Positives = 206/562 (36%), Gaps = 88/562 (15%)
Query: 134 LKKSYVSSDHVSKILRSLPSRW-RPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHET 192
L+K + D K + P R + KVT E+ K + ++ H+ + + E
Sbjct: 137 LRKKEIHKDKTEK--QEKPERKIQTKVTHKEKEKGKEKVREKEKPEKKATHKEKIEKKE- 193
Query: 193 SKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKK 252
++K++A K K+ A K+ E ++ G E A + + ++ ++K
Sbjct: 194 KPETKTVAKEQK----KAKTAEKSEEKTKKEVKGGKQEKVKQTAAKVKEVQKTPSKPKEK 249
Query: 253 FLSKRGSYKNFKKEDQKGCFNCK-----------KPGHFIADCPDLQKEKFKGKSKKSSF 301
++ + +++DQ C+ KPG A P L E+ + S
Sbjct: 250 EDKEKAAVSKHEQKDQYAF--CRYMIDIFVHGDLKPGQSPAIPPPLPTEQASRPTPASPA 307
Query: 302 NSSKFRKQIKKSLMATWEDL-----DSESGSDKEEADDDAKAAVGLVA-----TVSSEAV 351
K ++ K T E D + S+KE A D K G + TV A
Sbjct: 308 LEEKEGEKKKAEKKVTSETKKKEKEDIKKKSEKETAIDVEKKEPGKASETKQGTVKIAAQ 367
Query: 352 SEAESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLE 411
+ A+ D + E+ +K P + E + + KEK+V+ K K
Sbjct: 368 AAAKKDEKKEDSKKTKKPAE------------VEQPKGKKQEKKEKHVEPAKSPKKEH-- 413
Query: 412 LKASEEELKGFNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGIDRSKVAS 471
S + ++K+ ++ +E+ S K + G K
Sbjct: 414 --------------SVPSDKQVKAKTERAKEEIGAVSSKK--------AVPGKKEEKTTK 451
Query: 472 MIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKPEASGS 531
+ + + G S K KE +K K VP K K E
Sbjct: 452 TVEQEIRKEKSGKTSSILKDKE-PIKG---------KEEKVPASLKEKEPETKKDEKMSK 501
Query: 532 QAK-ITSKPENLKIKVMTKSDPKSQKIKILKRSEPVH---QNLIKPESKI----PKQKDQ 583
K + KP L+ K K +P+ +K SE V Q+++KPE + P++K
Sbjct: 502 AGKEVKPKPPQLQGKKEEKPEPQIKKEAKPAISEKVQIHKQDIVKPEKTVSHGKPEEKVL 561
Query: 584 KNKAATASEKTI-PKGVKPKVLNDQKPLSIH-PKVQGRKSKTSKANPKGPMKIWVPKSEL 641
K A EKT PK K +++P SI K + TS+ G K + + E
Sbjct: 562 KQVKAVTIEKTAKPKPTKKAEHREREPPSIKTDKPKPTPKGTSEVTESGKKKTEISEKE- 620
Query: 642 AKNAGVLKGKRETKVMVPRQRM 663
+K +K RE KV ++ +
Sbjct: 621 SKEKADMKHLREEKVSTRKESL 642
>USO1_YEAST (P25386) Intracellular protein transport protein USO1
Length = 1790
Score = 54.7 bits (130), Expect = 2e-06
Identities = 107/558 (19%), Positives = 215/558 (38%), Gaps = 80/558 (14%)
Query: 70 KMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDESIEEMYSRFQTLVS 129
K + TA + S E S K E ++ Q + K ++ I +M L
Sbjct: 819 KDKENQTALLEYKSTIHKQEDSIKTLEKGLETILSQ----KKKAEDGINKMGKDLFALSR 874
Query: 130 GLQIL--------KKSYVSSDHVSKILRSLPSRWRPKVTAI-------EEAK-DLNTLSV 173
+Q + K+ S+ + K +SL K+T I EE K N LS
Sbjct: 875 EMQAVEENCKNLQKEKDKSNVNHQKETKSLKEDIAAKITEIKAINENLEEMKIQCNNLSK 934
Query: 174 ED--LVSSLKVHEMSLNEHET-----SKKSKSIALPSKGKTSKSSKAYKASESVEESPDG 226
E + L ++ H+ ++K KS+A K +++ KA E
Sbjct: 935 EKEHISKELVEYKSRFQSHDNLVAKLTEKLKSLANNYKDMQAENESLIKAVE-------- 986
Query: 227 DSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPD 286
+S + S++++ L NK++ ++++++ F +RGS + N ++ I+D
Sbjct: 987 ESKNESSIQLSNLQNKIDSMSQEKENFQIERGSIEK----------NIEQLKKTISDLEQ 1036
Query: 287 LQKEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATV 346
++E KS + ++ QI L E A+D+ + +
Sbjct: 1037 TKEEII----SKSDSSKDEYESQI---------SLLKEKLETATTANDENVNKISELTKT 1083
Query: 347 SSEAVSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQK 406
E +E + +NE+ +K+ E +LKE+ EH E L+++ + +Q
Sbjct: 1084 REELEAELAAYKNLKNELETKLETSE--KALKEVKENEEHLKEEKIQLEKEATETKQQLN 1141
Query: 407 STLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGIDR 466
S L++ E+E + YE+++ + ++ E+ + L+D I +
Sbjct: 1142 SLRANLESLEKEHEDLAAQLKKYEEQIANKERQYNEEISQ---------LNDEITSTQQE 1192
Query: 467 SKVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKP 526
++ + + K + + E+ KS D + +K TN + ++S
Sbjct: 1193 NESIKKKNDELEGEVKAMKSTSEEQSNLK-KSEIDALNLQIKELKKKNETNEASLLESIK 1251
Query: 527 EASGSQAKI----------TSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLIKPESK 576
KI + L+ K+ D S+ +++ K SE + + L ++
Sbjct: 1252 SVESETVKIKELQDECNFKEKEVSELEDKLKASEDKNSKYLELQKESEKIKEELDAKTTE 1311
Query: 577 IPKQKDQKNKAATASEKT 594
+ Q ++ + A EK+
Sbjct: 1312 LKIQLEKITNLSKAKEKS 1329
Score = 43.9 bits (102), Expect = 0.004
Identities = 114/562 (20%), Positives = 217/562 (38%), Gaps = 92/562 (16%)
Query: 64 PRTEYMKMSDKSTAKAMFASLCANFEGS---KKVKEAKALMLVHQYE-------LFRMKD 113
PR EY + K+ K +AS F+ KV + +L + + F
Sbjct: 641 PRKEYFEFITKTLGKDNYASRIKQFKKDSYFSKVDMNEDSILTPELDETGLPKVYFSTYF 700
Query: 114 DESIEEMYSRFQTLVSGLQILKK-SYVSSDHVSKILRSLPSRWRPKVTAIE-EAKDLNTL 171
+ E R +T +S + S +S + V K+ R ++ + ++T+++ E + +
Sbjct: 701 IQLFNENIYRIRTALSHDPDEEPISKISFEEVEKLQRQC-TKLKGEITSLQTETESTHEN 759
Query: 172 SVEDLVSSLKVHE------MSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPD 225
E L++ H+ LN +S K L ++ K + S + + + D
Sbjct: 760 LTEKLIALTNEHKELDEKYQILNSSHSSLKENFSILETELKNVRDS-----LDEMTQLRD 814
Query: 226 GDSDEDQSVKMAMLSNK---------LEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKK 276
+D+ + A+L K ++ L + + LS++ ++ + K F +
Sbjct: 815 VLETKDKENQTALLEYKSTIHKQEDSIKTLEKGLETILSQKKKAEDGINKMGKDLFALSR 874
Query: 277 PGHFIAD-CPDLQKEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDD 335
+ + C +LQKEK KS+ N K K +K+ + A ++
Sbjct: 875 EMQAVEENCKNLQKEK-----DKSNVNHQKETKSLKEDIAAKITEI-------------- 915
Query: 336 AKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLK 395
+A++E + + + SK ++ + L E S F+ N + L
Sbjct: 916 -------------KAINENLEEMKIQCNNLSK-EKEHISKELVEYKSRFQSHDNLVAKLT 961
Query: 396 EKYVDLMKQQKSTLLELKASEEELKGFNLISATYEDRLKSLCQ--KLQEKCDKGSGNKHE 453
EK L K +++A E +LI A E + +S Q LQ K D S K
Sbjct: 962 EKLKSLANNYK----DMQAENE-----SLIKAVEESKNESSIQLSNLQNKIDSMSQEKEN 1012
Query: 454 IALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVP 513
I+R + I + K I E+ +E KS D KD +S
Sbjct: 1013 FQ--------IERGSIEKNI----EQLKKTISDLEQTKEEIISKS--DSSKDEYESQISL 1058
Query: 514 EGTNAKTAVQSKPEASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLIKP 573
+TA + E +++T E L+ ++ + K++ L+ SE + + +
Sbjct: 1059 LKEKLETATTANDENVNKISELTKTREELEAELAAYKNLKNELETKLETSEKALKEVKEN 1118
Query: 574 ESKIPKQKDQKNKAATASEKTI 595
E + ++K Q K AT +++ +
Sbjct: 1119 EEHLKEEKIQLEKEATETKQQL 1140
Score = 42.7 bits (99), Expect = 0.008
Identities = 90/430 (20%), Positives = 171/430 (38%), Gaps = 39/430 (9%)
Query: 46 KKLYKKHHKIRGIIVASIPRTE-----YMKMSDKSTAKAMFASLCANFEGSKKVKEAKAL 100
K+L KK+ ++ SI E ++ D+ K S + + + K +K L
Sbjct: 1233 KELKKKNETNEASLLESIKSVESETVKIKELQDECNFKEKEVSELEDKLKASEDKNSKYL 1292
Query: 101 MLVHQYELFRMKDDESIEEMYSRFQTLV----------SGLQILKK-SYVSSDHVSKILR 149
L + E + + D E+ + + + S L LKK S + + L
Sbjct: 1293 ELQKESEKIKEELDAKTTELKIQLEKITNLSKAKEKSESELSRLKKTSSEERKNAEEQLE 1352
Query: 150 SLPSRWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSK 209
L + + K A E+ + L + +++ E E + L +K +
Sbjct: 1353 KLKNEIQIKNQAFEKERKLLNEGSSTITQEYS-EKINTLEDELIRLQNENELKAKEIDNT 1411
Query: 210 SSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLS-KRGSYKNFK--KE 266
S+ K S S +E + + +S++ +LS K + + R +K LS +R + ++ + KE
Sbjct: 1412 RSELEKVSLSNDELLEEKQNTIKSLQDEILSYK-DKITRNDEKLLSIERDNKRDLESLKE 1470
Query: 267 DQKGCFNCKKPGHFIADCPDLQKEKFKGKSKKSSFNSSKFRKQIK------KSLMATWED 320
+ K + + K K + +KS K I+ KS M T
Sbjct: 1471 QLRAAQESKAKVEEGLKKLEEESSKEKAELEKSKEMMKKLESTIESNETELKSSMETIRK 1530
Query: 321 LDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPRQEL----VDS 376
D + K+ A++D K + S+ +S +D E+ SK+ + +++
Sbjct: 1531 SDEKLEQSKKSAEEDIKN----LQHEKSDLISRINESEKDIEELKSKLRIEAKSGSELET 1586
Query: 377 LKELLSLFEHR----TNELTDLKEKYVDLMKQQKSTLLELKASEEELKGFNLISATYEDR 432
+K+ L+ + + E T LK K D+ ++ K E+K+++EE + E
Sbjct: 1587 VKQELNNAQEKIRINAEENTVLKSKLEDIERELKDKQAEIKSNQEEKELLTSRLKELEQE 1646
Query: 433 LKSLCQKLQE 442
L S QK Q+
Sbjct: 1647 LDSTQQKAQK 1656
Score = 38.1 bits (87), Expect = 0.20
Identities = 72/356 (20%), Positives = 139/356 (38%), Gaps = 25/356 (7%)
Query: 13 DEELWDILEDGVDDLDLDEEGAAIDRKIHTPAQ---KKLYKKHHKIRGIIVASIPRTEYM 69
DE+L I D DL+ +E ++ + KKL ++ K + + S + +
Sbjct: 1451 DEKLLSIERDNKRDLESLKEQLRAAQESKAKVEEGLKKLEEESSKEKAELEKSKEMMKKL 1510
Query: 70 KMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDD--ESIEEMYSRFQTL 127
+ + +S + +S+ + +K++++K L K D I E + L
Sbjct: 1511 ESTIESNETELKSSMETIRKSDEKLEQSKKSAEEDIKNLQHEKSDLISRINESEKDIEEL 1570
Query: 128 VSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSL 187
S L+I KS + V + L + + K+ E + +ED+ LK + +
Sbjct: 1571 KSKLRIEAKSGSELETVKQELNNA----QEKIRINAEENTVLKSKLEDIERELKDKQAEI 1626
Query: 188 NEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLA 247
++ K+ + L + S++ KA +S EE V+ + L K L
Sbjct: 1627 KSNQEEKELLTSRLKELEQELDSTQQ-KAQKSEEERRA--EVRKFQVEKSQLDEKAMLLE 1683
Query: 248 RKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSKKSSFNSSKFR 307
K ++K ++K + +K + ++ + L KE K++ S +
Sbjct: 1684 TKYNDLVNKEQAWKRDEDTVKKTTDSQRQ------EIEKLAKELDNLKAENSKLKEANED 1737
Query: 308 KQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENE 363
+ LM DLD ++ + + D L +SS+ + E D EDE E
Sbjct: 1738 RSEIDDLMLLVTDLDEKNAKYRSKLKD-------LGVEISSDEEDDEEDDEEDEEE 1786
Score = 33.1 bits (74), Expect = 6.6
Identities = 62/318 (19%), Positives = 133/318 (41%), Gaps = 47/318 (14%)
Query: 134 LKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEH-ET 192
L+ + S V + L+ L + +E++K++ ++ L S+++ +E L ET
Sbjct: 1472 LRAAQESKAKVEEGLKKLEEESSKEKAELEKSKEM----MKKLESTIESNETELKSSMET 1527
Query: 193 SKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKK 252
+KS SK + K + +S S +S++D + L +KL A+ +
Sbjct: 1528 IRKSDEKLEQSKKSAEEDIKNLQHEKSDLISRINESEKD----IEELKSKLRIEAKSGSE 1583
Query: 253 FLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSKKSSFNSSKFRKQIKK 312
+ + N +++ + N ++ + D+++E K K++ S++ K++
Sbjct: 1584 LETVKQELNNAQEKIR---INAEENTVLKSKLEDIEREL---KDKQAEIKSNQEEKELLT 1637
Query: 313 SLMATWEDLDSESGSDKEEADDDAKAAVG--------------LVATVSSEAVSEAESDS 358
S + E + ++++++ +A V L+ T ++ V++ ++
Sbjct: 1638 SRLKELEQELDSTQQKAQKSEEERRAEVRKFQVEKSQLDEKAMLLETKYNDLVNKEQAWK 1697
Query: 359 EDENEVYSKIP--RQELVDSLKELLSLFEHRT-----NE-----------LTDLKEKYVD 400
DE+ V RQE+ KEL +L + NE +TDL EK
Sbjct: 1698 RDEDTVKKTTDSQRQEIEKLAKELDNLKAENSKLKEANEDRSEIDDLMLLVTDLDEKNAK 1757
Query: 401 LMKQQKSTLLELKASEEE 418
+ K +E+ + EE+
Sbjct: 1758 YRSKLKDLGVEISSDEED 1775
>TRDN_CANFA (P82179) Triadin
Length = 700
Score = 52.8 bits (125), Expect = 8e-06
Identities = 90/386 (23%), Positives = 147/386 (37%), Gaps = 80/386 (20%)
Query: 288 QKEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVS 347
++EK K + K +S K ++ +KK D D+ +K+E + G + V+
Sbjct: 306 EEEKKKAEKKVTSETKKKEKEDVKKK-----SDKDTAIDVEKKEPGKAPETKQGTIKVVA 360
Query: 348 SEAVSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKS 407
A A+ D + E+ +K P +E H + + KEKYV+ K K
Sbjct: 361 QAA---AKKDEKKEDSKKTKTPVEE-------------HPKGKKQEKKEKYVEPAKSSKK 404
Query: 408 TLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGIDRS 467
SA E ++K+ ++ +E+ S K + G
Sbjct: 405 E----------------HSAPSEKQVKAKTERAKEETSAASTKK--------AVPGKKEE 440
Query: 468 KVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKPE 527
K + + + G + K KE +K K +P K + KP+
Sbjct: 441 KTTKTVEQEIRKEKSGKTSTASKDKEPEIK----------KDEKMP-----KADKEVKPK 485
Query: 528 ASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLIKPESKIPKQKDQKNKA 587
SQ K K E+ K + K ++ I K + V KPE K+ KQ KA
Sbjct: 486 PPQSQVKKEEKSES-----QVKKEAKPEQ-DIAKPEKTVSHG--KPEEKVVKQVKATEKA 537
Query: 588 ATASEKTIPKGVKPKVLNDQKPLSIHPKVQGRKSK-TSKANPK----GPMKIWVPKSELA 642
A EKT+ K + +K P ++ K K TSK P+ G KI + E +
Sbjct: 538 AI--EKTVKPKPAKKAEHQEKE---SPTIKTDKPKPTSKETPEVTESGKKKIEKSEKE-S 591
Query: 643 KNAGVLKGKRETKVMVPRQRMFKAHD 668
K +K +E KV R+ ++H+
Sbjct: 592 KEKAEMKHLKEEKVST-RKESLQSHN 616
Score = 46.2 bits (108), Expect = 7e-04
Identities = 104/468 (22%), Positives = 176/468 (37%), Gaps = 79/468 (16%)
Query: 216 ASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCK 275
+S+ E+ DGD D D+ ++E KQK+ ++ +KE++
Sbjct: 114 SSDGDEDDDDGDEDTDK--------GEIEEPPLKQKEIHKEKA-----EKEEKPERKILA 160
Query: 276 KPGHFIADCPDLQKEKFKGKSKKSSFNSSKFRKQIK--KSLMATWEDLDSESGSDKEEAD 333
K H + +K K K KS+K + + K K+ K MA E K+E
Sbjct: 161 KVAH-----REKEKVKEKEKSEKKATHKEKIEKKEKPETKTMAKEERKAKTEEKIKKEVK 215
Query: 334 DDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPRQELVDS---LKELLSLFEH---R 387
+ V A E E E + + + + E D + ++ +F H R
Sbjct: 216 GGKQEKVKPTAAKVKEVQKTPPKAKEKEGKETAAVAKHEQKDQYAFCRYMIDMFVHGDLR 275
Query: 388 TNELTDLKEKYVDLMKQQ---KSTLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKC 444
+ L + + S LE K EE+ K +++ + + K + +++K
Sbjct: 276 PGQSPALPPPLPTVQASRPTPASPTLEGKEEEEKKKAEKKVTSETKKKEK---EDVKKKS 332
Query: 445 DKGSGNKHEIALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIK 504
DK + A+D + + K T K + +EK K
Sbjct: 333 DK------DTAID---VEKKEPGKAPETKQGTIKVVAQAAAKKDEK-------------K 370
Query: 505 DGLKSTFVPEGTNAKTAVQSKPEASGSQAKITSK----PENLKIKVMTKSDPKSQKIKIL 560
+ K T P + K Q K E AK + K P ++K T+ +
Sbjct: 371 EDSKKTKTPVEEHPKGKKQEKKEKYVEPAKSSKKEHSAPSEKQVKAKTERAKEETSAAST 430
Query: 561 KRSEPVHQNLIKPESKIPK------QKDQKNKAATASEKTIPKGVK----PKVLNDQKPL 610
K++ P K E K K +K++ K +TAS+ P+ K PK + KP
Sbjct: 431 KKAVPG-----KKEEKTTKTVEQEIRKEKSGKTSTASKDKEPEIKKDEKMPKADKEVKPK 485
Query: 611 SIHPKVQGRKSKTSKANPKGPMKIWVPKSELAK-NAGVLKGKRETKVM 657
P+ Q +K + S++ K K P+ ++AK V GK E KV+
Sbjct: 486 P--PQSQVKKEEKSESQVKKEAK---PEQDIAKPEKTVSHGKPEEKVV 528
Score = 33.5 bits (75), Expect = 5.0
Identities = 59/263 (22%), Positives = 99/263 (37%), Gaps = 25/263 (9%)
Query: 135 KKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHETSK 194
K+ YV SK S PS + K ++ + S + V K + + + +
Sbjct: 392 KEKYVEPAKSSKKEHSAPSEKQVKAKTERAKEETSAASTKKAVPGKKEEKTTKTVEQEIR 451
Query: 195 KSKS--IALPSKGK---TSKSSKAYKASESVEESPDGD--SDEDQSVKMAMLSNKLEY-L 246
K KS + SK K K K KA + V+ P E++S K E +
Sbjct: 452 KEKSGKTSTASKDKEPEIKKDEKMPKADKEVKPKPPQSQVKKEEKSESQVKKEAKPEQDI 511
Query: 247 ARKQKKFLSKRGSYKNFK--KEDQKGCFN-------CKKPGHFIADCPDLQKEKFKGKSK 297
A+ +K + K K K +K KK H + P ++ +K K SK
Sbjct: 512 AKPEKTVSHGKPEEKVVKQVKATEKAAIEKTVKPKPAKKAEHQEKESPTIKTDKPKPTSK 571
Query: 298 KSSFNSSKFRKQIKKSLMATWEDL------DSESGSDKEEADDDAKAAVGLVATVSSEAV 351
++ + +K+I+KS + E + + + KE A VS E +
Sbjct: 572 ETPEVTESGKKKIEKSEKESKEKAEMKHLKEEKVSTRKESLQSHNVTKAEKPARVSREDL 631
Query: 352 SE--AESDSEDENEVYSKIPRQE 372
+ A +++E E S RQ+
Sbjct: 632 EDVSASKKAKEEAEDVSSTKRQK 654
>GOA4_HUMAN (Q13439) Golgi autoantigen, golgin subfamily A member 4
(Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (72.1
protein)
Length = 2230
Score = 52.8 bits (125), Expect = 8e-06
Identities = 99/463 (21%), Positives = 203/463 (43%), Gaps = 52/463 (11%)
Query: 12 LDEELWDILEDGVDDLDLDEEGAAIDRKIHTPAQKKLYKKHHKIRGIIVASIPRTEYMKM 71
L+EEL +L+D D + + E ++K H Q K H++ SI RTE
Sbjct: 703 LEEEL-SVLKDQTDKMKQELEAKMDEQKNHHQQQVDSIIKEHEV------SIQRTEKALK 755
Query: 72 SDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDESIEEMYSRFQTLVSGL 131
+ + + E K +KE HQ + ++ D I+ Q + L
Sbjct: 756 DQINQLELLLK------ERDKHLKE-------HQAHVENLEAD--IKRSEGELQQASAKL 800
Query: 132 QILKKSYVSSDHV-SKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEH 190
+ + SY S+ H +K ++ + K+ +E + L T V ++ + K L+ H
Sbjct: 801 DVFQ-SYQSATHEQTKAYEEQLAQLQQKLLDLETERILLTKQVAEVEAQKKDVCTELDAH 859
Query: 191 ETSKKSKSIALPSKGKTSKSSKAYKASESVEESP--DGDSDEDQSVKMAMLSNKLEYLAR 248
+ + L + S+ + K+ V ES DG+ +++Q+ ++ + + R
Sbjct: 860 KIQVQDLMQQLEKQN--SEMEQKVKSLTQVYESKLEDGNKEQEQTKQILVEKENMILQMR 917
Query: 249 KQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSKKSSFNSSKFRK 308
+ +K + + K KED N + F ++K K K K + + +
Sbjct: 918 EGQKKEIEILTQKLSAKEDSIHILNEEYETKFKNQEKKMEKVKQKAKEMQETLKKKLLDQ 977
Query: 309 Q--IKKSLMATWEDLDSESGSDKEEADDDAKA-AVGLVATVSSEAVSEAESDSEDENEVY 365
+ +KK L T +L + + + A+A + G+ S+AVS E++ +++ E
Sbjct: 978 EAKLKKELENTALELSQKEKQFNAKMLEMAQANSAGI-----SDAVSRLETNQKEQIESL 1032
Query: 366 SKIPRQELVDSLKELLSLFEHRTN-ELTDLKEKYVDLMKQQKSTLLELK-------ASEE 417
+++ R+EL D ++S++E + N + +L+E + +++++ + ELK +E
Sbjct: 1033 TEVHRRELND----VISIWEKKLNQQAEELQEIHEIQLQEKEQEVAELKQKILLFGCEKE 1088
Query: 418 ELKGFNLISATYEDRLK--SLCQKLQEKCDKGSGNKHEIALDD 458
E+ I+ E+ +K + +LQE+ + S + + +A D+
Sbjct: 1089 EMN--KEITWLKEEGVKQDTTLNELQEQLKQKSAHVNSLAQDE 1129
Score = 51.6 bits (122), Expect = 2e-05
Identities = 111/497 (22%), Positives = 212/497 (42%), Gaps = 54/497 (10%)
Query: 95 KEAKALMLVHQYELFRMKDDES-IEEMYSRFQTLVSGLQILKKSYVSSDHVSKILRSLPS 153
+E L + Q ++++ E+ I+ M + ++LV+ + L+K + + S +
Sbjct: 1283 EEQNTLNISFQQATHQLEEKENQIKSMKADIESLVTEKEALQKEGGNQQQAASEKESCIT 1342
Query: 154 RWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHETS---KKSKSIALPSKGKTSKS 210
+ + +++ E + TL E+L KV SL++ T + SI+L K S
Sbjct: 1343 QLKKELS---ENINAVTLMKEELKEK-KVEISSLSKQLTDLNVQLQNSISLSEKEAAISS 1398
Query: 211 SKAYKASESVEESPDGDSDEDQSVKMAMLSNK----LEYL-------ARKQKKFLSKRGS 259
+ E E D +D S K+ LS + LE + + +KK S+
Sbjct: 1399 LRKQYDEEKCELL---DQVQDLSFKVDTLSKEKISALEQVDDWSNKFSEWKKKAQSRFTQ 1455
Query: 260 YKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSK-----KSSFNSSKFRKQIKKSL 314
++N KE Q K + + +L KE+ ++K K K + + K+S
Sbjct: 1456 HQNTVKELQIQLELKSKEAYEKDEQINLLKEELDQQNKRFDCLKGEMEDDKSKMEKKESN 1515
Query: 315 MATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPRQELV 374
+ T +L S++ E D + T+ E+++E + + ++ K ELV
Sbjct: 1516 LET--ELKSQTARIMELEDHITQK------TIEIESLNEVLKNYNQQKDIEHK----ELV 1563
Query: 375 DSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASEEELKGFNLISATYEDRLK 434
L+ L E + N + + +EK + L Q S EL+ ++EL+ NL + E+ LK
Sbjct: 1564 QKLQHFQELGEEKDNRVKEAEEKILTLENQVYSMKAELETKKKELEHVNLSVKSKEEELK 1623
Query: 435 SLCQKLQ-EKCDKGSGNKHEIALDDFIMAGIDRSKVASMIYSTYK-NKGKGIGYSEEKSK 492
+L +L+ E K + K + + +A I + ++ M + KG SE +K
Sbjct: 1624 ALEDRLESESAAKLAELKRKA---EQKIAAIKKQLLSQMEEKEEQYKKGTESHLSELNTK 1680
Query: 493 EYSLKSYCDCIKDGLKS-------TFVPEGTNAKTAVQSKPEASGSQAKITSKPENLKIK 545
+ +++ LKS T + + A ++ E + SQ + E KI
Sbjct: 1681 LQEREREVHILEEKLKSVESSQSETLIVPRSAKNVAAYTEQEEADSQGCVQKTYEE-KIS 1739
Query: 546 VMTKSDPKSQKIKILKR 562
V+ ++ ++K K+L+R
Sbjct: 1740 VLQRN--LTEKEKLLQR 1754
Score = 41.2 bits (95), Expect = 0.024
Identities = 76/335 (22%), Positives = 134/335 (39%), Gaps = 62/335 (18%)
Query: 308 KQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSK 367
K ++ L A + L+SES + E A+ + A + + +S+ E ++ E Y K
Sbjct: 1616 KSKEEELKALEDRLESESAAKLAELKRKAEQKI---AAIKKQLLSQME----EKEEQYKK 1668
Query: 368 IPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKAS--------EEEL 419
L EL + + R E+ L+EK + Q TL+ +++ +EE
Sbjct: 1669 GTESHL----SELNTKLQEREREVHILEEKLKSVESSQSETLIVPRSAKNVAAYTEQEEA 1724
Query: 420 KGFNLISATYEDRLKSLCQKLQEK---CDKGSGNKHEIALDDFIMAGIDRSKVASMIYST 476
+ TYE+++ L + L EK + K E F M + ++ + ++
Sbjct: 1725 DSQGCVQKTYEEKISVLQRNLTEKEKLLQRVGQEKEETVSSHFEMRCQYQERLIKLEHAE 1784
Query: 477 YKNKGKG--IGYS----EEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKPEASG 530
K IG+ EEK+K+YSL K+G K +Q+K
Sbjct: 1785 AKQHEDQSMIGHLQEELEEKNKKYSLIVAQHVEKEG-----------GKNNIQAKQNLEN 1833
Query: 531 --SQAKITSKPENLKIKVMTKSDPKSQKIK----ILKRSEPVH----QNLIKPESKIP-- 578
+ T + + L +++ QKIK L R + VH + L K+
Sbjct: 1834 VFDDVQKTLQEKELTCQIL------EQKIKELDSCLVRQKEVHRVEMEELTSKYEKLQAL 1887
Query: 579 KQKDQKNKAA-----TASEKTIPKGVKPKVLNDQK 608
+Q D +NK EK+ V+PK+L++ +
Sbjct: 1888 QQMDGRNKPTELLEENTEEKSKSHLVQPKLLSNME 1922
Score = 40.8 bits (94), Expect = 0.031
Identities = 112/563 (19%), Positives = 229/563 (39%), Gaps = 85/563 (15%)
Query: 44 AQKKLYKKHHKIRGIIVASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLV 103
A + L ++ K++GI+ S DKS + A L + ++ K+
Sbjct: 163 AYQMLQREKKKLQGILSQS---------QDKSLRR--IAELREELQMDQQAKK------- 204
Query: 104 HQYELFRMKDDESIEEMYSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKV---T 160
H E F D S+EE L + + +LK+ + +L+ LP + P+ T
Sbjct: 205 HLQEEF----DASLEEKDQYISVLQTQVSLLKQRLRNGPMNVDVLKPLP-QLEPQAEVFT 259
Query: 161 AIEEAKDLNTLSVED--LVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASE 218
E + VED V +L+ + + E K + S + + K E
Sbjct: 260 KEENPESDGEPVVEDGTSVKTLETLQQRVKRQENLLKRCKETIQSHKEQCTLLTSEK--E 317
Query: 219 SVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNF--KKEDQKGCF--NC 274
+++E D E + +K ++ K K +++ KN + E KG
Sbjct: 318 ALQEQLDERLQELEKIKDLHMAEKT--------KLITQLRDAKNLIEQLEQDKGMVIAET 369
Query: 275 KKPGHFIADCPDLQKEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDS--ESGSDKEEA 332
K+ H + + + + + + K+ + + R+Q +KS A +E+L+ + EEA
Sbjct: 370 KRQMHETLEMKEEEIAQLRSRIKQMTTQGEELREQKEKSERAAFEELEKALSTAQKTEEA 429
Query: 333 DDDAKAAVGLVATVSSEAVSEAESDSEDE----NEVYSKIPRQELVDSLKELLSLFEHRT 388
KA + E + E SE+E + S++ +QE+VD +K+ E +
Sbjct: 430 RRKLKAEM-------DEQIKTIEKTSEEERISLQQELSRV-KQEVVDVMKK---SSEEQI 478
Query: 389 NELTDLKEKYVDLMKQQKSTLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDKGS 448
+L L EK +L ++++ +L+ E E +++++K +K Q + K S
Sbjct: 479 AKLQKLHEK--ELARKEQELTKKLQTRERE----------FQEQMKVALEKSQSEYLKIS 526
Query: 449 GNKHEIALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLK 508
K + + + ++ K A I + +NK + + E + L+ ++ L+
Sbjct: 527 QEKEQ--QESLALEELELQKKA--ILTESENKLRDLQQEAETYRTRILE-----LESSLE 577
Query: 509 STFVPEGTNAK-TAVQSKPEASGSQAKITSKPENLKIKVMTKSDPK----SQKIKILKRS 563
+ +K AV + E + +IT E K ++ + + ++K+++LK+
Sbjct: 578 KSLQENKNQSKDLAVHLEAEKNKHNKEITVMVEKHKTELESLKHQQDALWTEKLQVLKQQ 637
Query: 564 EPVHQNLIKPESKIPKQKDQKNK 586
++ + + K+ K+K
Sbjct: 638 YQTEMEKLREKCEQEKETLLKDK 660
Score = 39.3 bits (90), Expect = 0.092
Identities = 113/574 (19%), Positives = 216/574 (36%), Gaps = 76/574 (13%)
Query: 73 DKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKDDESIEEMYSRFQTLVSGLQ 132
D++ KA L + S K LV + ++ +D + E+ S+ +T Q
Sbjct: 1128 DETKLKAHLEKLEVDLNKSLKENTFLQEQLV-ELKMLAEEDKRKVSELTSKLKTTDEEFQ 1186
Query: 133 ILKKSYVSSD--------HVSKILRSLPSRWR---PKVTAIEEAKDLNTLSVEDLVSSLK 181
LK S+ S+ K+ L + K A+ EAK +++ ++
Sbjct: 1187 SLKSSHEKSNKSLEDKSLEFKKLSEELAIQLDICCKKTEALLEAKTNELINISSSKTNAI 1246
Query: 182 VHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASE-------SVEESPDGDSDEDQSV 234
+ +S +H T+K +++ + + + ++ + +E S +++ +++ +
Sbjct: 1247 LSRISHCQHRTTKVKEALLIKTCTVSELEAQLRQLTEEQNTLNISFQQATHQLEEKENQI 1306
Query: 235 KMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCF-NCKKPGHFIADCPDLQKEKFK 293
K + +E L +K+ L K G + +++ C KK + L KE+ K
Sbjct: 1307 K--SMKADIESLV-TEKEALQKEGGNQQQAASEKESCITQLKKELSENINAVTLMKEELK 1363
Query: 294 GKSKKSSFNSSK---FRKQIKKSLM-----ATWEDLDSESGSDKEEADDDAKAAVGLVAT 345
K + S S + Q++ S+ A L + +K E D + V T
Sbjct: 1364 EKKVEISSLSKQLTDLNVQLQNSISLSEKEAAISSLRKQYDEEKCELLDQVQDLSFKVDT 1423
Query: 346 VSSEAVSEAE-------SDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKY 398
+S E +S E SE + + S+ + + +++KEL E ++ E + K++
Sbjct: 1424 LSKEKISALEQVDDWSNKFSEWKKKAQSRFTQHQ--NTVKELQIQLELKSKEAYE-KDEQ 1480
Query: 399 VDLMK----QQKSTLLELKASEEELKG-FNLISATYEDRLKSLCQKLQEKCDKGSGNKHE 453
++L+K QQ LK E+ K + E LKS ++ E D + E
Sbjct: 1481 INLLKEELDQQNKRFDCLKGEMEDDKSKMEKKESNLETELKSQTARIMELEDHITQKTIE 1540
Query: 454 IALDDFIMAGIDRSK-------VASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDG 506
I + ++ ++ K V + + + K E + K +L++ +K
Sbjct: 1541 IESLNEVLKNYNQQKDIEHKELVQKLQHFQELGEEKDNRVKEAEEKILTLENQVYSMKAE 1600
Query: 507 LKSTFVPEGTNAKTAVQSKPEASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPV 566
L++ K E + SK E LK + K+ LKR
Sbjct: 1601 LET--------------KKKELEHVNLSVKSKEEELKALEDRLESESAAKLAELKR---- 1642
Query: 567 HQNLIKPESKIPKQKDQKNKAATASEKTIPKGVK 600
K E KI K Q E+ KG +
Sbjct: 1643 -----KAEQKIAAIKKQLLSQMEEKEEQYKKGTE 1671
Score = 37.7 bits (86), Expect = 0.27
Identities = 72/387 (18%), Positives = 154/387 (39%), Gaps = 52/387 (13%)
Query: 106 YELFRMKDDESIEEMYSRFQTLVS---------------GLQILKKSYVSSDHVSKILRS 150
+E MK++E I ++ SR + + + + L+K+ ++ + R
Sbjct: 374 HETLEMKEEE-IAQLRSRIKQMTTQGEELREQKEKSERAAFEELEKALSTAQKTEEARRK 432
Query: 151 LPSRWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKS 210
L + ++ IE+ + +S++ +S +K + + KKS + K +
Sbjct: 433 LKAEMDEQIKTIEKTSEEERISLQQELSRVKQEVV-----DVMKKSSEEQIAKLQKLHEK 487
Query: 211 SKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKG 270
A K E ++ + + + +K+A+ ++ EYL Q+K ++ S + E QK
Sbjct: 488 ELARKEQELTKKLQTREREFQEQMKVALEKSQSEYLKISQEK--EQQESLALEELELQKK 545
Query: 271 CFNCKKPGHFIADCPDLQKE---------KFKGKSKKSSFNSSKFRKQIKKSLMATWEDL 321
+ DLQ+E + + +KS + K + L A
Sbjct: 546 AILTESENKL----RDLQQEAETYRTRILELESSLEKSLQENKNQSKDLAVHLEAEKNKH 601
Query: 322 DSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPRQELVDSLKELL 381
+ E E+ + ++ + +E + + + E E + QE LK+
Sbjct: 602 NKEITVMVEKHKTELESLKHQQDALWTEKLQVLKQQYQTEMEKLREKCEQEKETLLKDKE 661
Query: 382 SLFE-------HRTNELTDLKEKYVDLMKQQKSTLLELKAS-EEELKGFNLISATYEDRL 433
+F+ +T E D+K+ ++ + + S +L+ + EEEL + +D+
Sbjct: 662 IIFQAHIEEMNEKTLEKLDVKQTELESLSSELSEVLKARHKLEEEL-------SVLKDQT 714
Query: 434 KSLCQKLQEKCDKGSGNKHEIALDDFI 460
+ Q+L+ K D+ N H+ +D I
Sbjct: 715 DKMKQELEAKMDE-QKNHHQQQVDSII 740
>CYL1_BOVIN (P35662) Cylicin I (Multiple-band polypeptide I)
Length = 667
Score = 52.4 bits (124), Expect = 1e-05
Identities = 76/317 (23%), Positives = 120/317 (36%), Gaps = 28/317 (8%)
Query: 193 SKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKK 252
S SK KG T + K K ++ GDS + + K +K + A+K
Sbjct: 300 SVDSKDAKKDKKGATKDTKKGAKKDTESTDAESGDSKDAKKGKKESKKDKKKD-AKKDAA 358
Query: 253 FLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQK-EKFKGKSKKSSFNSSKFRKQIK 311
++ G K+ KK+ +KG + KK D + + G SK + +S K +K K
Sbjct: 359 SDAESGDSKDAKKDSKKGKKDSKKDNKKKDAKKDAESTDAESGDSKDAKKDSKKGKKDSK 418
Query: 312 KS-----LMATWEDLDSESGSDKEEADD------DAKAAVGLVATVSSEAVSEAESDSED 360
K E D+ESG K D D K VS++A SE+E D++
Sbjct: 419 KDDKKKDAKKDAESTDAESGDSKNAKKDSKKGKKDDKKKDAKKDAVSTDADSESEGDAKK 478
Query: 361 ENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASEEELK 420
+ K + D K+ + T +D + K V ++ + KA+E
Sbjct: 479 SKKDSKKDKKDLKKDDQKKPAMKSKESTETESDWESKKVKRDSKKDTKKTAKKATESSGA 538
Query: 421 GFNLISATY---EDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGIDRSKVA------- 470
++ S Y + KS + +E K K +D+ D K A
Sbjct: 539 ESDVSSKRYLKKTEMFKSSDAESEESLFKPGSKKR---VDESDATSTDSKKDAVEPKRGI 595
Query: 471 --SMIYSTYKNKGKGIG 485
+T+K KGK IG
Sbjct: 596 KMPSRRTTFKEKGKKIG 612
Score = 47.0 bits (110), Expect = 4e-04
Identities = 58/209 (27%), Positives = 88/209 (41%), Gaps = 17/209 (8%)
Query: 163 EEAKDLNTLSVEDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEE 222
++AK S +D K S E SK +K + K K SK K ++ E
Sbjct: 336 KDAKKGKKESKKDKKKDAKKDAASDAESGDSKDAKKDSKKGK-KDSKKDNKKKDAKKDAE 394
Query: 223 SPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSK--------RGSYKNFKKEDQKG-CFN 273
S D +S + + K K + +KK K G KN KK+ +KG +
Sbjct: 395 STDAESGDSKDAKKDSKKGKKDSKKDDKKKDAKKDAESTDAESGDSKNAKKDSKKGKKDD 454
Query: 274 CKKPGHFIADCPDLQKEKFKGKSKKSSFNSSKFRKQIKKS-----LMATWEDLDSESGSD 328
KK A D E +G +KKS +S K +K +KK M + E ++ES +
Sbjct: 455 KKKDAKKDAVSTDADSES-EGDAKKSKKDSKKDKKDLKKDDQKKPAMKSKESTETESDWE 513
Query: 329 KEEADDDAKAAVGLVATVSSEAVSEAESD 357
++ D+K A ++E+ S AESD
Sbjct: 514 SKKVKRDSKKDTKKTAKKATES-SGAESD 541
Score = 38.1 bits (87), Expect = 0.20
Identities = 96/476 (20%), Positives = 166/476 (34%), Gaps = 53/476 (11%)
Query: 134 LKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVED---LVSSLKVHEMSLNEH 190
L+K + S ++ I+R S +R T I +K + +D + K+ + H
Sbjct: 99 LRKKF-QSPSINLIVRRQAS-FRHPYTHITHSKKAESKKYKDDKKETALKKISKKDTGPH 156
Query: 191 ETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQ 250
E +K K K SKSS + S+ + + + + S+ +++ K E K
Sbjct: 157 EVDEKPKRRNKADK-TPSKSSHGSQLSKKSKSKSETNPESKDSISVSIKHQKKEKRYSKD 215
Query: 251 KKFL-----SKRGSYKNFKKEDQKGCFNCKKPGHFIADC-----PDLQKEKFKGKSKKSS 300
K + S + K+ K C K + D + +F K S
Sbjct: 216 SKEMDFESTSTKKYSKSSKNNSDAVSETCSKNSSNVGLMVHLGESDAESMEFDMWLKNYS 275
Query: 301 FNSSK--FRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDS 358
N+SK +K KK D +S D ++ A A +E+ DS
Sbjct: 276 QNNSKKPTKKDAKKDAKGKGSDAESVDSKDAKKDKKGATKDTKKGAKKDTESTDAESGDS 335
Query: 359 EDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASEEE 418
+D + + + + D+ K+ S E D K+ D K +K + + K + +
Sbjct: 336 KDAKKGKKESKKDKKKDAKKDAAS-----DAESGDSKDAKKDSKKGKKDSKKDNKKKDAK 390
Query: 419 LKGFNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGIDRSKVASMIYSTYK 478
+ + + + + K +K K K + D S A S
Sbjct: 391 KDAESTDAESGDSKDAKKDSKKGKKDSKKDDKKKDAKKDA-------ESTDAESGDSKNA 443
Query: 479 NKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKPEASGSQAKITSK 538
K G ++K K D KD + T S+ E ++K SK
Sbjct: 444 KKDSKKGKKDDKKK--------DAKKDAVS-----------TDADSESEGDAKKSKKDSK 484
Query: 539 PENLKIKVMTKSDPKSQKIKILKRSEPVHQNLIKPESKIPKQKDQKNKAATASEKT 594
+ +K K D K +K K S + + K +KD K A A+E +
Sbjct: 485 KDKKDLK---KDDQKKPAMKS-KESTETESDWESKKVKRDSKKDTKKTAKKATESS 536
Score = 33.1 bits (74), Expect = 6.6
Identities = 63/278 (22%), Positives = 101/278 (35%), Gaps = 35/278 (12%)
Query: 323 SESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPRQELVDSLKELLS 382
S+ K+ DD + A+ ++ + E + + N+ K P + S S
Sbjct: 128 SKKAESKKYKDDKKETALKKISKKDT-GPHEVDEKPKRRNKA-DKTPSKSSHGSQLSKKS 185
Query: 383 LFEHRTNELTDLKEKYVDLMKQQKSTLLELKASEEELKGFNLISATYEDRLKSLCQKLQE 442
+ TN + K+ +K QK K S+E + + + Y K+ + E
Sbjct: 186 KSKSETNP--ESKDSISVSIKHQKKEKRYSKDSKE-MDFESTSTKKYSKSSKNNSDAVSE 242
Query: 443 KCDKGSGNKHEIALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDC 502
C K S N +M + S SM + + YS+ SK+ + K D
Sbjct: 243 TCSKNSSNVG-------LMVHLGESDAESMEFDMWLKN-----YSQNNSKKPTKK---DA 287
Query: 503 IKDGLKSTFVPEGTNAKTAVQSKPEASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKR 562
KD E ++K A + K A+ K K +D +S K K+
Sbjct: 288 KKDAKGKGSDAESVDSKDAKKDKKGATKDTKKGAKKDTE-------STDAESGDSKDAKK 340
Query: 563 SEPVHQNLIKPESKIPKQKDQKNKAATASEKTIPKGVK 600
K ESK K+KD K AA+ +E K K
Sbjct: 341 G--------KKESKKDKKKDAKKDAASDAESGDSKDAK 370
>SCP1_MOUSE (Q62209) Synaptonemal complex protein 1 (SCP-1 protein)
Length = 993
Score = 52.0 bits (123), Expect = 1e-05
Identities = 124/703 (17%), Positives = 274/703 (38%), Gaps = 102/703 (14%)
Query: 2 KTNMYSFIMGLDEELWDILEDGVDDLDLDEEGAAIDRK-IHTPAQKKLYKKHHKIRGIIV 60
KTN Y + +++ L ++ + L E + + KL + H KI+ +
Sbjct: 193 KTNKYEYEREETRQVYVDLNSNIEKMILAFEELRVQAENARLEMHFKLKEDHEKIQHL-- 250
Query: 61 ASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALM-----LVHQYELFRMKDDE 115
EY K + + L + E K+K+ L+ +Q E DE
Sbjct: 251 ----EEEYQKEVNNKENQVS-ELLIQSAEKENKMKDLTFLLEESRDKANQLEEKTKLQDE 305
Query: 116 SIEEMYSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVED 175
+++E+ + L S L+ +K S S K A+EE + T ++
Sbjct: 306 NLKELSEKKDHLTSELEDIKMSMQRSMSTQK--------------ALEEDLQIATKTISQ 351
Query: 176 LVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVK 235
L +V E + E +K + S + T+ + + +E + D + +V+
Sbjct: 352 LT---EVKEAQMEELNKAKTTHSFVVTELKATTCTLEELLRTEQQRLEKNEDQLKLITVE 408
Query: 236 MAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKE-KFKG 294
+ SN+LE + + + + KN EDQK K+ + + ++E F
Sbjct: 409 LQKKSNELEEMTKFKNNKEVELEELKNILAEDQKLLDEKKQVEKLAEELQEKEQELTFLL 468
Query: 295 KSKKSSFNSSKFRKQIKKS----LMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEA 350
++++ + + + + K+ + E++ +E +K + + + A+ ++ + +
Sbjct: 469 ETREKEVHDLQEQVTVTKTSEQHYLKQVEEMKTELEKEKLK-NTELTASCDMLLLENKKF 527
Query: 351 VSEA-----ESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQ 405
V EA E E+ + K + L+ ++ L H +EL ++++++ +Q
Sbjct: 528 VQEASDMALELKKHQEDIINCKKQEERLLKQIENLEEKEMHLRDELESVRKEFI---QQG 584
Query: 406 KSTLLELKASEEELKGFNLISATYEDRLKSL---CQKLQEKCDKGSGNKHEIALDDFIMA 462
+L SEE + E ++K L C L+++ + S N E+ ++
Sbjct: 585 DEVKCKLDKSEENARSIECEVLKKEKQMKILESKCNNLKKQVENKSKNIEELHQEN---- 640
Query: 463 GIDRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLK-SYCDCIKDGLKSTFVPEGTNAKTA 521
T K K ++ Y +K S + + K F N +
Sbjct: 641 ------------KTLKKKSSA---EIKQLNAYEIKVSKLELELESTKQRFEEMTNNYQKE 685
Query: 522 VQSKPEASGSQAKITSKPENLK------IKVMTKSDPKSQK-----IKILKRSEPVHQNL 570
+++K + G K+ + E K +K+ + D + Q + ++++ + + +
Sbjct: 686 IENKKISEG---KLLGEVEKAKATVDEAVKLQKEIDLRCQHKIAEMVALMEKHKHQYDKI 742
Query: 571 IKP---ESKIPKQKDQKNKAATASEKTIPKGVKPKVLNDQKPLSIHPKVQGRKSKTSKAN 627
++ E + K ++Q+ +A + +T ++ ++++ +K L I K + K K +K
Sbjct: 743 VEERDSELGLYKNREQEQSSAKIALETELSNIRNELVSLKKQLEIE-KEEKEKLKMAK-- 799
Query: 628 PKGPMKIWVPKSELAKNAGVLKGKRETKVMVPRQRMFKAHDWR 670
+N +LK K++ K+ +A W+
Sbjct: 800 ---------------ENTAILKDKKDKKIQASLLESPEATSWK 827
Score = 44.3 bits (103), Expect = 0.003
Identities = 91/484 (18%), Positives = 184/484 (37%), Gaps = 86/484 (17%)
Query: 173 VEDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQ 232
VE L L+ E L +++ + L + +K+S+ + + E + E +
Sbjct: 450 VEKLAEELQEKEQELTFLLETREKEVHDLQEQVTVTKTSEQHYLKQVEEMKTEL---EKE 506
Query: 233 SVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKF 292
+K L+ + L + KKF+ + K+ Q+ NCKK Q+E+
Sbjct: 507 KLKNTELTASCDMLLLENKKFVQEASDMALELKKHQEDIINCKK-----------QEER- 554
Query: 293 KGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVS 352
L+ E+L+ + ++E + K + V +
Sbjct: 555 ---------------------LLKQIENLEEKEMHLRDELESVRKEFIQQGDEVKCKLDK 593
Query: 353 EAESDSEDENEVYSKIPRQELVDS-LKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLE 411
E+ E EV K + ++++S L E+++ + +L ++ L K+ + + +
Sbjct: 594 SEENARSIECEVLKKEKQMKILESKCNNLKKQVENKSKNIEELHQENKTLKKKSSAEIKQ 653
Query: 412 LKASEEELKGFNLISATYEDRLKSLCQKLQEKCDK---------GSGNKHEIALDDFI-- 460
L A E ++ L + + R + + Q++ + G K + +D+ +
Sbjct: 654 LNAYEIKVSKLELELESTKQRFEEMTNNYQKEIENKKISEGKLLGEVEKAKATVDEAVKL 713
Query: 461 MAGID---RSKVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTN 517
ID + K+A M+ K+K + EE+ E GL E ++
Sbjct: 714 QKEIDLRCQHKIAEMVALMEKHKHQYDKIVEERDSEL-----------GLYKNREQEQSS 762
Query: 518 AKTAVQSKPEASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLIKPESKI 577
AK A+++ E S + ++ S + L+I+ K +K+K+ K + +
Sbjct: 763 AKIALET--ELSNIRNELVSLKKQLEIEKEEK-----EKLKMAKENTAI----------- 804
Query: 578 PKQKDQKNKAATASEKTIPKGVK----PKVLNDQKPLSIHPKVQGRKSKTSKANPKGPMK 633
KD+K+K AS P+ K Q + + KSK ++ N + K
Sbjct: 805 --LKDKKDKKIQASLLESPEATSWKFDSKTTPSQNISRLSSSMDSGKSKDNRDNLRASAK 862
Query: 634 IWVP 637
+P
Sbjct: 863 SILP 866
>UN89_CAEEL (O01761) Muscle M-line assembly protein unc-89
(Uncoordinated protein 89)
Length = 6632
Score = 50.8 bits (120), Expect = 3e-05
Identities = 115/548 (20%), Positives = 208/548 (36%), Gaps = 55/548 (10%)
Query: 133 ILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKD------LNTLSVEDLVSSLKVHEMS 186
+++K +SS V K + P+ T + T+S ++ S++ +
Sbjct: 1245 VVEKQDLSSSEVQKEIAQQVKEASPEATTTITMETSLTSTKTTTMSTTEVTSTVGGVTVE 1304
Query: 187 LNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEYL 246
E E+ + I S G T S K V D +D + +S E L
Sbjct: 1305 TKESESESATTVIGGGSGGVTEGSISVSKIE--VVSKTDSQTDVREGTPKRRVSFAEEEL 1362
Query: 247 A-------RKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFI------ADCPDLQKEK-- 291
RK+KK S K+ +K ++K KK G + + +KEK
Sbjct: 1363 PKEVIDSDRKKKKSPSPDKKEKSPEKTEEKPASPTKKTGEEVKSPKEKSPASPTKKEKSP 1422
Query: 292 ----FKGKSKKSSFNSSKFRKQ------IKKSLMATWEDLDSESGSDKE---EADDDAKA 338
K +KK SS +K+ KK+ E +S + KE E +D K+
Sbjct: 1423 AAEEVKSPTKKEKSPSSPTKKEKSPSSPTKKTGDEVKEKSPPKSPTKKEKSPEKPEDVKS 1482
Query: 339 AVGLVATVSSEAVSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKY 398
V + + + E S++ E + + + + +S + S+ + +T E D K K
Sbjct: 1483 PVKKEKSPDATNIVEVSSETTIE-KTETTMTTEMTHESEESRTSVKKEKTPEKVDEKPKS 1541
Query: 399 VDLMKQ--QKSTLLELKASEEELKGFNLI-----SATYEDRL--KSLCQKLQEKCDKGSG 449
+ +KS E+K+ ++ K + S T +++ K + + + S
Sbjct: 1542 PTKKDKSPEKSITEEIKSPVKKEKSPEKVEEKPASPTKKEKSPEKPASPTKKSENEVKSP 1601
Query: 450 NKHEIALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKS 509
K E + + ++ + K S + K K +EKS E KS + +K K
Sbjct: 1602 TKKEKSPEKSVVEELKSPKEKSPEKADDKPKSPT---KKEKSPE---KSATEDVKSPTKK 1655
Query: 510 TFVPEGTNAKTAVQSKPEASGSQA---KITSKPENLKIKVMTKSDPKSQKIKILKRSEPV 566
PE K +K E+S ++ ++ S + K + P S K + V
Sbjct: 1656 EKSPEKVEEKPTSPTKKESSPTKKTDDEVKSPTKKEKSPQTVEEKPASPTKKEKSPEKSV 1715
Query: 567 HQNLIKPESKIPKQKDQKNKAATASEKTIPKGVKPKVLNDQKPLSIHPKVQGRKSKTSKA 626
+ + P+ K P++ ++K K+ T EK+ K +V + K K K K+
Sbjct: 1716 VEEVKSPKEKSPEKAEEKPKSPTKKEKSPEKSAAEEVKSPTKKEKSPEKSAEEKPKSPTK 1775
Query: 627 NPKGPMKI 634
P+K+
Sbjct: 1776 KESSPVKM 1783
>MYSB_CAEEL (P02566) Myosin heavy chain B (MHC B)
Length = 1966
Score = 50.8 bits (120), Expect = 3e-05
Identities = 77/388 (19%), Positives = 158/388 (39%), Gaps = 55/388 (14%)
Query: 49 YKKHHKIRGIIVASIPRTEYMKMSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYEL 108
+K + K++ ++ A E K++DK KA+ SL + K+++E+ A ++ + L
Sbjct: 842 FKLYGKVKPMLKAGKEAEELEKINDK--VKALEDSLAKEEKLRKELEESSAKLVEEKTSL 899
Query: 109 FRMKDDESIEEMYSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKDL 168
F + ES + S + ++ L+ +K +S +S++ L + ++ AK
Sbjct: 900 FT--NLESTKTQLSDAEERLAKLEAQQKD--ASKQLSELNDQLADN-EDRTADVQRAKKK 954
Query: 169 NTLSVEDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDS 228
VE L ++ EMSL + E+ K+SK + S D
Sbjct: 955 IEAEVEALKKQIQDLEMSLRKAESEKQSKDHQIRSLQ---------------------DE 993
Query: 229 DEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQ 288
+ Q +A L+ + ++ +K + ++ + E+ KG K DL+
Sbjct: 994 MQQQDEAIAKLNKEKKHQEEINRKLM------EDLQSEEDKGNHQNKVKAKLEQTLDDLE 1047
Query: 289 KEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSS 348
+ K ++ + K ++++ L E++D ESG + + +++ K + +VSS
Sbjct: 1048 DSLEREKRARADLDKQK--RKVEGELKIAQENID-ESGRQRHDLENNLKKKESELHSVSS 1104
Query: 349 EAVSE------------------AESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNE 390
E +E + E ENE S+ L+ L + +E
Sbjct: 1105 RLEDEQALVSKLQRQIKDGQSRISELEEELENERQSRSKADRAKSDLQRELEELGEKLDE 1164
Query: 391 LTDLKEKYVDLMKQQKSTLLELKASEEE 418
V++ K++++ L +L+ EE
Sbjct: 1165 QGGATAAQVEVNKKREAELAKLRRDLEE 1192
>FUTS_DROME (Q9W596) Microtubule-associated protein futsch
Length = 5412
Score = 50.8 bits (120), Expect = 3e-05
Identities = 100/527 (18%), Positives = 198/527 (36%), Gaps = 54/527 (10%)
Query: 158 KVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNE-----HETSKKSKSIALPSKGKTSKSSK 212
+V+ + KD +L V S + SL + ETS+ GK+ +SK
Sbjct: 3197 RVSVADSIKDEKSLLVSQEASRPESEAESLKDAAAPSQETSRPESVTESVKDGKSPVASK 3256
Query: 213 AYKASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCF 272
SV E+ +DE + + L + +K L+ + + K+E ++
Sbjct: 3257 EASRPASVAENAKDSADESKEQRPESLPQSKAGSIKDEKSPLASKDEAEKSKEESRRESV 3316
Query: 273 NCKKP-GHFIADCPDLQKEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEE 331
+ P P E K +++KS S K+ + S +GS K+E
Sbjct: 3317 AEQFPLVSKEVSRPASVAESVKDEAEKSKEESPLMSKEASRPA--------SVAGSVKDE 3368
Query: 332 A----DDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEHR 387
A ++ + +V + + S+ S S +E + K + +S E L
Sbjct: 3369 AEKSKEESRRESVAEKSPLPSKEASRPASVAESVKDEADKSKEESRRESGAEKSPLASKE 3428
Query: 388 TNELTDLKEKYVDLMKQQKSTLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDKG 447
+ + E D ++ K ++ E + + + + R S+ + ++++ +K
Sbjct: 3429 ASRPASVAESIKDEAEKSKE-----ESRRESVAEKSPLPSKEASRPTSVAESVKDEAEK- 3482
Query: 448 SGNKHEIALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLK---------- 497
+K E D ++S +AS S + + + EKSKE S +
Sbjct: 3483 --SKEESRRDSV----AEKSPLASKEASRPASVAESVQDEAEKSKEESRRESVAEKSPLA 3536
Query: 498 --------SYCDCIKDGLKSTFVPEGTNAKTAVQSKPEASGSQAKITSKPENLKIKVMTK 549
S + IKD + + E + ++ + P AS ++ TS E++K + K
Sbjct: 3537 SKEASRPASVAESIKDEAEKS--KEESRRESVAEKSPLASKEASRPTSVAESVKDEA-EK 3593
Query: 550 SDPKSQKIKILKRSEPVHQNLIKPESKIPKQKDQKNKAATASEKTIPKGVKPKVLND-QK 608
S +S + + ++S + +P S +D+ K+ S + P + +
Sbjct: 3594 SKEESSRDSVAEKSPLASKEASRPASVAESVQDEAEKSKEESRRESVAEKSPLASKEASR 3653
Query: 609 PLSIHPKVQ--GRKSKTSKANPKGPMKIWVPKSELAKNAGVLKGKRE 653
P S+ V+ KSK K + E ++ A V + ++
Sbjct: 3654 PASVAESVKDDAEKSKEESRRESVAEKSPLASKEASRPASVAESVKD 3700
Score = 47.0 bits (110), Expect = 4e-04
Identities = 96/430 (22%), Positives = 171/430 (39%), Gaps = 55/430 (12%)
Query: 163 EEAKDLNTLSVEDLVSSLKVHEMSLNEHETSKKS---KSIA----LPSKGKTSKSSKAYK 215
E + + L+ ++ V E +E E SK+ +S+A LPSK + +S A
Sbjct: 3675 ESVAEKSPLASKEASRPASVAESVKDEAEKSKEESRRESVAEKSPLPSKEASRPTSVAES 3734
Query: 216 ASESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCK 275
+ E+S + E + K ++ S K +R S + + K+E ++ K
Sbjct: 3735 VKDEAEKSKEESRRESVAEKSSLASKKA---SRPASVAESVKDEAEKSKEESRRESVAEK 3791
Query: 276 KP-GHFIADCPDLQKEKFKGKSKKSSFNSSKFRKQIKKSL--------MATWEDLDSESG 326
P A P E K +++KS S + K L + E + E+
Sbjct: 3792 SPLASKEASRPASVAESVKDEAEKSKEESRRESVAEKSPLPSKEASRPTSVAESVKDEAD 3851
Query: 327 SDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEH 386
KEE+ ++ A +A++ + + +DE E + R+E S+ E L
Sbjct: 3852 KSKEESRRESGAEKSPLASMEASRPTSVAESVKDETEKSKEESRRE---SVTEKSPLPSK 3908
Query: 387 RTNELTDLKEKYVDLMKQQKSTLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDK 446
+ T + E D ++ K ++ E + + +++ R S+ + ++ D+
Sbjct: 3909 EASRPTSVAESVKDEAEKSKE-----ESRRESVAEKSPLASKESSRPASVAESIK---DE 3960
Query: 447 GSGNKHEIALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSEEKSKEYSL-KSYCDCIKD 505
G K E S+ SM S KG S SKE S S + +KD
Sbjct: 3961 AEGTKQE-------------SRRESMPESGKAESIKG-DQSSLASKETSRPDSVVESVKD 4006
Query: 506 GLKSTFVPEGTNA-KTAVQSKPEASGSQAK-----ITSKPENLKIKVMTKSDPKSQKIKI 559
T PEG+ K+ V S+PE+ AK + S+PE++ K S S+ + +
Sbjct: 4007 ---ETEKPEGSAIDKSQVASRPESVAVSAKDEKSPLHSRPESVADKSPDASKEASRSLSV 4063
Query: 560 LK-RSEPVHQ 568
+ S P+ +
Sbjct: 4064 AETASSPIEE 4073
Score = 45.8 bits (107), Expect = 0.001
Identities = 88/499 (17%), Positives = 197/499 (38%), Gaps = 44/499 (8%)
Query: 163 EEAKDLNTLSVEDLVSSLKVHEMSLNEHETSKKS---KSIALPSKGKTSKSSKAYKASES 219
+ + + L+ ++ V E +E E SK+ +S+A S + ++S+ +ES
Sbjct: 3490 DSVAEKSPLASKEASRPASVAESVQDEAEKSKEESRRESVAEKSPLASKEASRPASVAES 3549
Query: 220 VEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFL-SKRGSYKNFKKEDQKGCFNCKKP- 277
+++ + +E + +A S A + S + + K+E + K P
Sbjct: 3550 IKDEAEKSKEESRRESVAEKSPLASKEASRPTSVAESVKDEAEKSKEESSRDSVAEKSPL 3609
Query: 278 -------GHFIADCPDLQKEKFKGKSKKSSF-NSSKFRKQIKKSLMATWEDLDSESGSDK 329
+A+ + EK K +S++ S S + + E + ++ K
Sbjct: 3610 ASKEASRPASVAESVQDEAEKSKEESRRESVAEKSPLASKEASRPASVAESVKDDAEKSK 3669
Query: 330 EEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTN 389
EE+ ++ A +A+ + + +DE E + R+E S+ E L +
Sbjct: 3670 EESRRESVAEKSPLASKEASRPASVAESVKDEAEKSKEESRRE---SVAEKSPLPSKEAS 3726
Query: 390 ELTDLKEKYVDLMKQQKSTLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDKG-S 448
T + E D ++ K ++ E + + +++ R S+ + ++++ +K
Sbjct: 3727 RPTSVAESVKDEAEKSKE-----ESRRESVAEKSSLASKKASRPASVAESVKDEAEKSKE 3781
Query: 449 GNKHEIALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSEEKSKEYSL------------ 496
++ E + +A + S+ AS+ S K S+E+S+ S+
Sbjct: 3782 ESRRESVAEKSPLASKEASRPASVAESVKDEAEK----SKEESRRESVAEKSPLPSKEAS 3837
Query: 497 --KSYCDCIKDGLKSTFVPEGTNAKTAVQSKPEASGSQAKITSKPENLKIKVMTKSDPKS 554
S + +KD + E + ++ + P AS ++ TS E++K + KS +S
Sbjct: 3838 RPTSVAESVKDEADKS--KEESRRESGAEKSPLASMEASRPTSVAESVKDET-EKSKEES 3894
Query: 555 QKIKILKRSEPVHQNLIKPESKIPKQKDQKNKAATASEKTIPKGVKPKVLND-QKPLSIH 613
++ + ++S + +P S KD+ K+ S + P + +P S+
Sbjct: 3895 RRESVTEKSPLPSKEASRPTSVAESVKDEAEKSKEESRRESVAEKSPLASKESSRPASVA 3954
Query: 614 PKVQGRKSKTSKANPKGPM 632
++ T + + + M
Sbjct: 3955 ESIKDEAEGTKQESRRESM 3973
Score = 43.1 bits (100), Expect = 0.006
Identities = 112/576 (19%), Positives = 222/576 (38%), Gaps = 75/576 (13%)
Query: 114 DESIEEMYSRFQTLVSGL----------QILKKSYVSSDHVSKILRSLPSRWRPKVTAIE 163
DES+ + SR +++ G + + + +D + ++ R + + +I+
Sbjct: 1879 DESMSKEPSRRESVKDGAAQSRETSRPASVAESAKDGADDLKELSRPESTTQSKEAGSIK 1938
Query: 164 EAKDLNTLSVEDLVSSLKVHEMSLNEHETSKKS---KSIA----LPSKGKTSKSSKAYKA 216
+ K + L+ E+ V E +E E SK+ +S+A LPSK + +S A
Sbjct: 1939 DEK--SPLASEEASRPASVAESVKDEAEKSKEESRRESVAEKSPLPSKEASRPASVAESI 1996
Query: 217 SESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKK 276
+ E+S + E + K + S + +R S + + K+E ++ K
Sbjct: 1997 KDEAEKSKEESRRESVAEKSPLPSKEA---SRPASVAESIKDEAEKSKEESRRESVAEKS 2053
Query: 277 P-GHFIADCPDLQKEKFKGKSKKSSFNSSKFRKQIKKSL--------MATWEDLDSESGS 327
P A P E K +++KS S + K L + E + E+
Sbjct: 2054 PLPSKEASRPASVAESIKDEAEKSKEESRRESVAEKSPLPSKEASRPASVAESIKDEAEK 2113
Query: 328 DKEEADDDAKAAVG-LVATVSSEAVSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEH 386
KEE+ ++ A L + +S S AES +DE E + R+E S+ E L
Sbjct: 2114 SKEESRRESVAEKSPLPSKEASRPASVAES-IKDEAEKSKEESRRE---SVAEKSPLPSK 2169
Query: 387 RTNELTDLKEKYVDLMKQQKSTLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDK 446
+ + E D ++ K ++ E + + + + R S+ + ++++ +K
Sbjct: 2170 EASRPASVAESIKDEAEKSKE-----ESRRESVAEKSPLPSKEASRPASVAESIKDEAEK 2224
Query: 447 GSGN-KHEIALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKD 505
+ E + + + S+ AS+ S K EE +E + + K+
Sbjct: 2225 SKEETRRESVAEKSPLPSKEASRPASVAESIKDEAEKS---KEESRRESAAEKSPLPSKE 2281
Query: 506 GLKSTFVPEGTNA---KTAVQSKPEASGSQAKITS-KPENLKIKVMTKSDPKSQKIKI-- 559
+ V E K+ +S+ E+ K S K + +K +++ + ++ +K
Sbjct: 2282 ASRPASVAESVKDEADKSKEESRRESMAESGKAQSIKGDQSPLKEVSRPESVAESVKDDP 2341
Query: 560 LKRSEPVHQNLI----------------------KPESKIPKQKDQKNKAATASEKTIPK 597
+K EP + + +PES + KD+ K + E
Sbjct: 2342 VKSKEPSRRESVAGSVTADSARDDQSPLESKGASRPESVVDSVKDEAEKQESRRESKTES 2401
Query: 598 GVKPKVLNDQKPLSIHPKVQGRKSKTSKANPKGPMK 633
+ PK +D+ P + V ++T + + PMK
Sbjct: 2402 VIPPKAKDDKSPKEVLQPVS--MTETIREDADQPMK 2435
Score = 38.5 bits (88), Expect = 0.16
Identities = 76/367 (20%), Positives = 132/367 (35%), Gaps = 57/367 (15%)
Query: 299 SSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDS 358
+S+ ++KF +L + A A AA AT++ ++D+
Sbjct: 523 ASYKTTKFSPVASAALAVQHPQQQDNKAKEAAAAAAAAAAAAASAATIARAKADSMDTDA 582
Query: 359 EDENEVYSK--------IPRQELVDSLKELLSLFEHRTNELTDL-KEKYVD--LMKQQKS 407
E E+E + P ++ ++ E E + D+ +EK V+ +MK Q++
Sbjct: 583 EPEHEADPEPADTGDEAAPTEQEPEAETEPEPEHEPEAEQDKDVGEEKKVEVLIMKPQQA 642
Query: 408 TLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGIDRS 467
T + AS ++ G + SA + KL + KG +K + + + ID
Sbjct: 643 TPAVIAASGKD--GVDAASAD-----ATPTGKLSKASAKGKADKPRAEVKPVVRSRIDTK 695
Query: 468 KVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKP- 526
SM K E+KS + + P NAK V S+P
Sbjct: 696 PPKSMDRKLAKR-------DEKKSSPTT-----------TPAARAPVAQNAKPKVLSRPA 737
Query: 527 ---EASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLIKPESKIPKQKDQ 583
S + AK + N K+ + + Q + R E ++ +Q +
Sbjct: 738 TKSSPSSTPAKSAKEANNRKVLESKQQAARVQATSTVSR----RVTSTASERRVQQQAEA 793
Query: 584 KNKAATASEKTIPKGVKPKVLNDQKPLSIHPKVQGRKSKTSKANPKGPMKIWVPKSELAK 643
K A A++ T +KP+S P+ G P P+K PK+ K
Sbjct: 794 KTAATGATQAT-----------QRKPISRRPR--GVSPSKRAPAPGSPVKQAKPKAADLK 840
Query: 644 NAGVLKG 650
+ KG
Sbjct: 841 KTRLDKG 847
Score = 36.6 bits (83), Expect = 0.59
Identities = 56/302 (18%), Positives = 112/302 (36%), Gaps = 23/302 (7%)
Query: 320 DLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPRQ-ELVDSLK 378
D++ + S +EA K+ +G A E+ +E+ D+ + E R +V+S K
Sbjct: 2785 DIEKTASSPIDEAP---KSLIGCPAEERPESPAESAKDAAESVEKSKDASRPPSVVESTK 2841
Query: 379 ELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASEEELKGFNLISATYEDRLKSLCQ 438
+ +++ E ++ K E S R S+ +
Sbjct: 2842 A-----DSTKGDISPSPESVLEGPKDDVEKSKESSRPPSVSASITGDSTKDVSRPASVVE 2896
Query: 439 KLQEKCDKGSGNKHEIALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLKS 498
++++ DK + IA + ++ +S S + K++ + E +E ++S
Sbjct: 2897 SVKDEHDKAESRRESIAKVESVIDEAGKSDSKSSSQDSQKDEKSTLASKEASRRESVVES 2956
Query: 499 YCDCIKDGLKSTFVPEGTNAKTAVQSKPEASGSQAKITSKPENLKIKVMTKSDPKSQK-- 556
D D KS PE A + S +K TS+P ++ ++ +T D KS++
Sbjct: 2957 SKD---DAEKSESRPESVIASGEPVPRESKSPLDSKDTSRPGSM-VESVTAEDEKSEQQS 3012
Query: 557 --------IKILKRSEPVHQNLIKPESKIPKQKDQKNKAATASEKTIPKGVKPKVLNDQK 608
+K + + Q +P S KD K + + + D+
Sbjct: 3013 RRESVAESVKADTKKDGKSQEASRPSSVDELLKDDDEKQESRRQSITGSHKAMSTMGDES 3072
Query: 609 PL 610
P+
Sbjct: 3073 PM 3074
Score = 36.2 bits (82), Expect = 0.78
Identities = 101/514 (19%), Positives = 196/514 (37%), Gaps = 67/514 (13%)
Query: 163 EEAKDLNTLSVEDLVSSLKVHEMSLNEHETSKKS---KSIALPSKGKTSKSSKAYKASES 219
E + + L ++ V E +E E SK+ +S+A S + +SS+ +ES
Sbjct: 3897 ESVTEKSPLPSKEASRPTSVAESVKDEAEKSKEESRRESVAEKSPLASKESSRPASVAES 3956
Query: 220 VEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGH 279
+++ +G E + M S K E + Q SK S
Sbjct: 3957 IKDEAEGTKQESRRESMPE-SGKAESIKGDQSSLASKETSR------------------- 3996
Query: 280 FIADCPDLQKEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAA 339
PD E K +++K ++ I KS +A+ + + S D++ +
Sbjct: 3997 -----PDSVVESVKDETEKPEGSA------IDKSQVASRPESVAVSAKDEKSPLHSRPES 4045
Query: 340 VGLVATVSSEAVSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYV 399
V + +S+ S + S +E + + PR + L L+L +L L +
Sbjct: 4046 VADKSPDASKEASRSLSVAETASSPIEEGPRS--IADLSLPLNLTGEAKGKLPTLSSP-I 4102
Query: 400 DLMKQQKSTLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDKGSGNKHEI-ALDD 458
D+ + LE+KA ++S E D+ S EI ++
Sbjct: 4103 DVAE---GDFLEVKAESSPRPA--VLSKPAEFSQPDTGHTASTPVDEASPVLEEIEVVEQ 4157
Query: 459 FIMAGIDRSKVASM--IYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGT 516
+G+ + + + + K + + E + +L S + + L+S+
Sbjct: 4158 HTTSGVGATGATAETDLLDLTETKSETVTKQSETTLFETLTSKVESKVEVLESSVKQVEE 4217
Query: 517 NAKTAV-QSKPEASGSQAKITSKPENLKIKVMTKSDPKSQKI--------KILKRSEPVH 567
+T+V Q++ + S ++T K ++ + D +++ K+LK + +
Sbjct: 4218 KVQTSVKQAETTVTDSLEQLTKKSSEQLTEIKSVLDTNFEEVAKIVADVAKVLKSDKDIT 4277
Query: 568 QNLIKPESKIPKQKDQKNKAATASEKTIPKGVKPKVLNDQKPLSIHPKVQGRKSKTSKAN 627
I P+ +Q ++K K+ +E+ K + D+K L I KV+ K+S
Sbjct: 4278 D--IIPDFD-ERQLEEKLKSTADTEEESDKSTR-----DEKSLEISVKVEIESEKSSPDQ 4329
Query: 628 PKGPMKI----WVPKSELAK-NAGVLKGKRETKV 656
GP+ I + +SE A+ G+L R V
Sbjct: 4330 KSGPISIEEKDKIEQSEKAQLRQGILTSSRPESV 4363
Score = 35.8 bits (81), Expect = 1.0
Identities = 82/438 (18%), Positives = 163/438 (36%), Gaps = 39/438 (8%)
Query: 20 LEDGVDDLDLDEEGAAIDRKIHTPAQKKLYKKHHKIR-----GIIVASIPRTEYMKMSDK 74
L G+ E A+ + +P+Q +H ++ +S P + ++S+K
Sbjct: 4350 LRQGILTSSRPESVASQPESVPSPSQSAASHEHKEVELSESHKAEKSSRPESVASQVSEK 4409
Query: 75 STAKAMFASLCANF---EGSKKVKEA--KALMLVHQYELFRMKDDESIEEM---YSRFQT 126
+ AS + F EG ++ E+ +L E +M++ S E + ++
Sbjct: 4410 DMKTSRPASSTSQFSTKEGDEETTESLLHSLTTTETVETKQMEEKSSFESVSTSVTKSTV 4469
Query: 127 LVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKD---LNTLSVEDLVSSLKVH 183
L S + + +S+ +S L+ S R ++++ K NT ED +S
Sbjct: 4470 LSSQSTVQLREESTSESLSSSLKVEDSSRRESLSSLLAEKGGIATNTSLKEDTSASASQL 4529
Query: 184 EMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKL 243
E L + E ++ KS+K K + + + + + K L +
Sbjct: 4530 EELLVQSEECSSESIVSEIQTSIAQKSNKEIKDARETKVTSQFTTTTSSATKDDSLKETV 4589
Query: 244 -EYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKG--KSKKSS 300
E+LA + K +S + ++ + C K ++ Q+ F G +S++ S
Sbjct: 4590 AEFLATE--KIVSAKEAFSTEATKSADDCLK-KTTASAVSSTSASQRALFVGTDESRRES 4646
Query: 301 FNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSED 360
S Q +S + + D E D +E ++ +AT+ ++ + D E
Sbjct: 4647 LLS-----QASESRLTHSDPEDEEPADDVDERSSVKESRSKSIATIMMTSIYKPSEDME- 4700
Query: 361 ENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASEEELK 420
P +LV+ E + E E+T + L ++S+ S+
Sbjct: 4701 --------PISKLVEEEHEHV---EELAQEVTSTSKTTTLLQSSEQSSTTTSSTSKTGAS 4749
Query: 421 GFNLISATYEDRLKSLCQ 438
I+ T D+ S Q
Sbjct: 4750 RVESITLTQMDQQTSQSQ 4767
Score = 34.7 bits (78), Expect = 2.3
Identities = 95/475 (20%), Positives = 182/475 (38%), Gaps = 58/475 (12%)
Query: 191 ETSKKSKSIALPSKGKTS--KSSKAYKASESVEESPDGDSDEDQSVKMAML-SNKLEYLA 247
+ + +SIA K ++S K+ + ES+ ES +S D+ +A +++ E +
Sbjct: 1495 KAESRRESIAKTHKDESSLDKAKEQESRRESLAESIKPESGIDEKSALASKEASRPESVT 1554
Query: 248 RKQKKFLSKRGSYKNFKKE---DQKGCFNCK---KPGHFIADCPDLQKEKFKGKSKKSSF 301
K K+ + ++ K E D+K K +PG + D + EK K S++ S
Sbjct: 1555 DKSKEPSRRESIAESLKAESTKDEKSAPPSKEASRPGSVVESVKD-ETEKSKEPSRRESI 1613
Query: 302 NSS-----KFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAES 356
S +FR+ + + G E A +++ L + +S S ES
Sbjct: 1614 AESAKPPIEFREVSRPESVI--------DGIKDESAKPESRRDSPLASKEASRPESVLES 1665
Query: 357 DSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASE 416
++ + K R+ + +S K + +E + L K + + +++ S
Sbjct: 1666 VKDEPIKSTEKSRRESVAESFKA-----DSTKDEKSPLTSKDISRPESAVENVMDAVGSA 1720
Query: 417 EELKGFNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGIDRSKVASMIYST 476
E + ++ ++ R +S+ + ++ DK + + + I S V I
Sbjct: 1721 ERSQPESVTASRDVSRPESVAESEKDDTDKP---------ESVVESVIPASDVVE-IEKG 1770
Query: 477 YKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEG---TNAKTAVQSKPEASGSQA 533
+K KG+ S E K S ++S PE ++ + + E S +
Sbjct: 1771 AADKEKGVFVSLEIGKPDSPSEVISRPGPVVESV-KPESRRESSTEIVLPCHAEDSKEPS 1829
Query: 534 KITSKPENLKIKV-MTKSDPKSQKIKILKRSEPVHQNLIKPESKIPKQKDQ--------- 583
+ SK E LK + + K + + + +S + +PES + KD+
Sbjct: 1830 RPESKVECLKDESEVLKGSTRRESVAESDKSSQPFKETSRPESAVGSMKDESMSKEPSRR 1889
Query: 584 ---KNKAATASEKTIPKGVKPKV---LNDQKPLSIHPKVQGRKSKTSKANPKGPM 632
K+ AA + E + P V +D K LS K S + K P+
Sbjct: 1890 ESVKDGAAQSRETSRPASVAESAKDGADDLKELSRPESTTQSKEAGSIKDEKSPL 1944
Score = 33.5 bits (75), Expect = 5.0
Identities = 73/324 (22%), Positives = 131/324 (39%), Gaps = 38/324 (11%)
Query: 295 KSKKSSFNSS-KFRKQIKKSLMATWEDLDSESGSD--KEEADDDAKAAVGLVATVSSEAV 351
KSKKS S + + KKS A+ E ES ++ K+EA + T E+
Sbjct: 1453 KSKKSPVPSRPESEAKDKKSPFASGEASRPESVAESVKDEAGKAESRRESIAKTHKDESS 1512
Query: 352 SEAESDSEDENEVYSKIPRQEL-VDSLKELLSLFEHRTNELTDL------KEKYVDLMKQ 404
+ + E E ++ + E +D L S R +TD +E + +K
Sbjct: 1513 LDKAKEQESRRESLAESIKPESGIDEKSALASKEASRPESVTDKSKEPSRRESIAESLKA 1572
Query: 405 QKSTLLELKA--SEEELKGFNLISATYEDRLKSLCQKLQEKCDKGSGNK---HEIALDDF 459
+ ST E A S+E + +++ + ++ KS +E + + E++ +
Sbjct: 1573 E-STKDEKSAPPSKEASRPGSVVESVKDETEKSKEPSRRESIAESAKPPIEFREVSRPES 1631
Query: 460 IMAGI--DRSKVASMIYSTYKNKGKGIGYSE-EKSKEYSLKSYCDCIKDGLKSTFVPEGT 516
++ GI + +K S S +K S E K+ +KS ++ + +F + T
Sbjct: 1632 VIDGIKDESAKPESRRDSPLASKEASRPESVLESVKDEPIKSTEKSRRESVAESFKADST 1691
Query: 517 NAKTAVQSKPEASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEP----VHQNLIK 572
K E S +K S+PE+ VM + +RS+P +++ +
Sbjct: 1692 --------KDEKSPLTSKDISRPESAVENVM-------DAVGSAERSQPESVTASRDVSR 1736
Query: 573 PESKIPKQKDQKNKAATASEKTIP 596
PES +KD +K + E IP
Sbjct: 1737 PESVAESEKDDTDKPESVVESVIP 1760
>MFP1_ARATH (Q9LW85) MAR binding filament-like protein 1
Length = 727
Score = 50.4 bits (119), Expect = 4e-05
Identities = 110/547 (20%), Positives = 202/547 (36%), Gaps = 40/547 (7%)
Query: 114 DESIEEMYSRFQTLVSGLQILKKSYVSS-----DHVSKILRSLPSRWRPKVTAIEEAKDL 168
+E+IE + ++ + L + +K + + + K + + + AKDL
Sbjct: 156 EETIESLKNQLKDRERALVLKEKDFEAKLQHEQEERKKEVEKAKEEQLSLINQLNSAKDL 215
Query: 169 NTLSVEDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDS 228
T +L S K+ E ++ E+ + S S A + K + +K + + VE D
Sbjct: 216 VTELGRELSSEKKLCEKLKDQIESLENSLSKA--GEDKEALETKLREKLDLVEGLQD--- 270
Query: 229 DEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQ 288
++ +LS +L+ K ++F + KKE + N + DL
Sbjct: 271 ------RINLLSLELKDSEEKAQRFNASLA-----KKEAELKELN----SIYTQTSRDLA 315
Query: 289 KEKFKGKSKKSSFNSSKFRKQIKKS----LMATWEDLDSESGSDKEEADDDAKAAVGLVA 344
+ K + K +K ++ K S L L +E S ++ D +K L
Sbjct: 316 EAKLEIKQQKEELIRTQSELDSKNSAIEELNTRITTLVAEKESYIQKLDSISKDYSALKL 375
Query: 345 TVSSEAVSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQ 404
T ++A ++AE S E E+ Q+L ++L L +++ DL EKY D +
Sbjct: 376 TSETQAAADAELISRKEQEI------QQLNENLDRALDDVNKSKDKVADLTEKYEDSKRM 429
Query: 405 QKSTLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGI 464
L +K EL+G DR+ L L E S + E+A+
Sbjct: 430 LDIELTTVKNLRHELEGTKKTLQASRDRVSDLETMLDESRALCSKLESELAIVHEEWKEA 489
Query: 465 DRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQS 524
++ KN+ + EK +K + + LK + V + K V+
Sbjct: 490 KERYERNLDAEKQKNEISASELALEKDLRRRVKDELEGVTHELKESSVKNQSLQKELVEI 549
Query: 525 KPEASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLIKPESKIPKQKDQK 584
+ S ++ E K + + K + +IL E ++L + K D+
Sbjct: 550 YKKVETSNKEL---EEEKKTVLSLNKEVKGMEKQILMERE-ARKSLETDLEEAVKSLDEM 605
Query: 585 NKAATASEKTIPKGVKPKVLNDQKPLSIHPKVQGRKSKTSKANPKGPMKIWVPKSELAKN 644
NK + + + K V N + + + G SK + + L K
Sbjct: 606 NKNTSILSRELEK-VNTHASNLEDEKEVLQRSLGEAKNASKEAKENVEDAHILVMSLGKE 664
Query: 645 AGVLKGK 651
VL+ K
Sbjct: 665 REVLEKK 671
>GAR2_SCHPO (P41891) Protein gar2
Length = 500
Score = 50.1 bits (118), Expect = 5e-05
Identities = 44/186 (23%), Positives = 87/186 (46%), Gaps = 9/186 (4%)
Query: 178 SSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMA 237
SS + + S+ + + SKK + + S+ ++S S +SES ES +S+ S +
Sbjct: 62 SSPEPSKKSVKKQKKSKKKEESSSESESESSSSESESSSSES--ESSSSESESSSSESSS 119
Query: 238 MLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSK 297
S + + ++KK S S + +E+++ ++ +D + + +S+
Sbjct: 120 SESEEEVIVKTEEKKESSSESSSSSESEEEEEAVVKIEEKKESSSDSSS-ESSSSESESE 178
Query: 298 KSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESD 357
SS S + + ++K+ ++ SES SD E + D + + S + SE+ES
Sbjct: 179 SSSSESEEEEEVVEKT--EEKKEGSSESSSDSESSSDSSSES----GDSDSSSDSESESS 232
Query: 358 SEDENE 363
SEDE +
Sbjct: 233 SEDEKK 238
Score = 44.7 bits (104), Expect = 0.002
Identities = 47/200 (23%), Positives = 86/200 (42%), Gaps = 9/200 (4%)
Query: 185 MSLNEHETSKKSKSIALPSKGKTSKSSKAYK-ASESVEESPDGDSDEDQSVKMAMLSNKL 243
M+ + + KKS KG K SK+ K E+ +E S D S K + K
Sbjct: 1 MAKKDKTSVKKSVKETASKKGAIEKPSKSKKITKEAAKEIAKQSSKTDVSPKKSKKEAKR 60
Query: 244 EYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSKKSSFNS 303
KK + K+ K KK+++ + + ++ + E +S+ SS S
Sbjct: 61 ASSPEPSKKSVKKQ---KKSKKKEESSSESESESSSSESESSSSESESSSSESESSSSES 117
Query: 304 SKFRKQIKKSLMATWEDLDSES-GSDKEEADDDAKAAVGLVATVSSEAVSEAE---SDSE 359
S + ++ ++ T E +S S S E++++ +A V + S + S +E S+SE
Sbjct: 118 SSSESE-EEVIVKTEEKKESSSESSSSSESEEEEEAVVKIEEKKESSSDSSSESSSSESE 176
Query: 360 DENEVYSKIPRQELVDSLKE 379
E+ +E+V+ +E
Sbjct: 177 SESSSSESEEEEEVVEKTEE 196
Score = 42.4 bits (98), Expect = 0.011
Identities = 42/199 (21%), Positives = 77/199 (38%), Gaps = 7/199 (3%)
Query: 172 SVEDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDED 231
S E S+K + S + E+S +S+S + S+ ++S S +SES S + S E
Sbjct: 63 SPEPSKKSVKKQKKSKKKEESSSESESESSSSESESSSSESESSSSESESSSSESSSSES 122
Query: 232 QSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDL---- 287
+ + K E + S+ K E++K + ++
Sbjct: 123 EEEVIVKTEEKKESSSESSSSSESEEEEEAVVKIEEKKESSSDSSSESSSSESESESSSS 182
Query: 288 ---QKEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVA 344
++E+ K+++ SS+ + S ++ E DS+S SD E
Sbjct: 183 ESEEEEEVVEKTEEKKEGSSESSSDSESSSDSSSESGDSDSSSDSESESSSEDEKKRKAE 242
Query: 345 TVSSEAVSEAESDSEDENE 363
S E ++ S+D NE
Sbjct: 243 PASEERPAKITKPSQDSNE 261
Score = 37.7 bits (86), Expect = 0.27
Identities = 39/161 (24%), Positives = 67/161 (41%), Gaps = 10/161 (6%)
Query: 288 QKEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVS 347
+K K + K S S K K+ KKS +SES S E++ + + ++
Sbjct: 52 KKSKKEAKRASSPEPSKKSVKKQKKSKKKEESSSESESESSSSESESSSSESES--SSSE 109
Query: 348 SEAVSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKS 407
SE+ S S SE E EV K ++ S S E + ++EK +
Sbjct: 110 SESSSSESSSSESEEEVIVKTEEKKESSSESSSSSESEEEEEAVVKIEEK----KESSSD 165
Query: 408 TLLELKASEEELKGFNLISATYEDRLKSLCQKLQEKCDKGS 448
+ E +SE E + S++ + + + +K +EK + S
Sbjct: 166 SSSESSSSESESES----SSSESEEEEEVVEKTEEKKEGSS 202
>GCC2_HUMAN (Q8IWJ2) GRIP and coiled-coil domain-containing protein 2
(Golgi coiled coil protein GCC185) (CTCL tumor antigen
se1-1) (CLL-associated antigen KW-11)
Length = 1583
Score = 49.3 bits (116), Expect = 9e-05
Identities = 124/649 (19%), Positives = 258/649 (39%), Gaps = 78/649 (12%)
Query: 2 KTNMYSFIMGLDEELWDILE--DGVDDLDLDEEGAAI--DRKIHTPAQK--KLYKKHHKI 55
KT +Y F+ + E+ + E D V+ L E A + K + Q+ K+ + +I
Sbjct: 655 KTQLYGFLKEMGSEVSEDSEEKDVVNVLQAVGESLAKINEEKCNLAFQRDEKVLELEKEI 714
Query: 56 RGIIVASIPRTEYMK--MSDKSTAKAMFASLCANFEGSKKVKEAKALMLVHQYELFRMKD 113
+ + S+ + E +K + D K + + K+ ++ L + + E R+++
Sbjct: 715 KCLQEESVVQCEELKSLLRDYEQEKVLLRKELEEIQSEKEALQSDLLEMKNANEKTRLEN 774
Query: 114 DESIEEMYSRFQTLV-SGLQILKKSYVSSDH--VSKILRSLPSR-WRPKVTAIEEAKDLN 169
+ ++ QT S + K+ +H + +L R R ++ ++++ +
Sbjct: 775 QNLLIQVEEVSQTCSKSEIHNEKEKCFIKEHENLKPLLEQKELRDRRAELILLKDSLAKS 834
Query: 170 TLSVEDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSD 229
D +SS+K E +K +++ K K K +K + ++ D
Sbjct: 835 PSVKNDPLSSVK---------ELEEKIENLEKECKEKEEKINKIKLVAVKAKKELDSSRK 885
Query: 230 EDQSVK--MAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDL 287
E Q+VK + L ++ + L+ + + SYKN E +K ++ D+
Sbjct: 886 ETQTVKEELESLRSEKDQLSASMRDLIQGAESYKNLLLEYEKQ-----------SEQLDV 934
Query: 288 QKEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVS 347
+KE+ + +Q++ S + E + D + + + T+
Sbjct: 935 EKER----ANNFEHRIEDLTRQLRNSTLQC------------ETINSDNEDLLARIETLQ 978
Query: 348 SEAVSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEHRT--NELTDLKEKYVDLMKQQ 405
S A E + + + ++ + L++ + EH T NEL +L+ + KQ
Sbjct: 979 SNA-KLLEVQILEVQRAKAMVDKELEAEKLQKEQKIKEHATTVNELEELQVQLQKEKKQL 1037
Query: 406 KSTLLELKASEEELKGFNLIS---ATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMA 462
+ T+ EL+ +++ + L++ A YE +K L QKL K +K + EI
Sbjct: 1038 QKTMQELELVKKDAQQTTLMNMEIADYERLMKELNQKLTNKNNKIEDLEQEIK------- 1090
Query: 463 GIDRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAV 522
I + K ++ + Y E+ +K +K K L + E +
Sbjct: 1091 -IQKQKQETLQEEITSLQSSVQQYEEKNTK---IKQLLVKTKKELADSKQAETDHLILQA 1146
Query: 523 QSKPEASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQNLIKPESKIPKQKD 582
K E SQ ++ E KI++ + K + + LK S HQ + +
Sbjct: 1147 SLKGELEASQQQV----EVYKIQLAEITSEKHKIHEHLKTSAEQHQRTLSAYQQRVTALQ 1202
Query: 583 QKNKAATASEKTIPKGVKPKVLNDQKPLSIHPKVQGRKSKT-SKANPKG 630
++ +AA A + T+ + + +H ++ +K+K+ S+A +G
Sbjct: 1203 EECRAAKAEQATVTSEF------ESYKVRVHNVLKQQKNKSMSQAETEG 1245
>GCC2_MOUSE (Q8CHG3) GRIP and coiled-coil domain-containing protein 2
(Golgi coiled coil protein GCC185)
Length = 1679
Score = 48.5 bits (114), Expect = 2e-04
Identities = 107/506 (21%), Positives = 203/506 (39%), Gaps = 62/506 (12%)
Query: 105 QYELFRMKDDESIEEMYSRFQTLVSGLQILKKSYVSSD--HVSKIL----RSLPSRWRPK 158
Q +L MK+ E+ QTL + ++ L ++ S + H K+L +L + +
Sbjct: 849 QLDLLEMKNTN--EKASLENQTLSTQVEELSQTLHSRNEVHDEKVLVIEHENLRLLLKQR 906
Query: 159 VTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHETSKKSKSIALPSKGKTSKSSKAYKASE 218
+ +++ + L + L S V + E +K +S+ SK K K SK +
Sbjct: 907 ESELQDVRAELILLKDSLEKSPSVKDQLSLVKELEEKIESLEKESKDKDEKISKIKLVAV 966
Query: 219 SVEESPDGDSDEDQSVKMAMLSNKLEY--LARKQKKFLSKRGSYKNFKKEDQKGCFNCKK 276
++ D + E Q+++ + S + E L+ K+FL SYK+ E K
Sbjct: 967 KAKKELDSNRKEAQTLREELESVRSEKDRLSASMKEFLQGAESYKSLLLEYDKQ------ 1020
Query: 277 PGHFIADCPDLQKEKFKGKSKKSSFNSSKFRKQIKKSLMATWEDLDSESG---SDKEEAD 333
++ D++KE+ + + KQ++ S +E L S++ + E
Sbjct: 1021 -----SEQLDVEKERAHNFER----HIEDLTKQLRNST-CQYERLTSDNEDLLARIETLQ 1070
Query: 334 DDAKAAVGLVATVS-SEAVSEAESDSEDENEVYSKIPRQELVDSLKELLSLFEHRTNELT 392
+AK + V ++ V E E D+E+ + +KE +S NEL
Sbjct: 1071 ANAKLLEAQILEVQKAKGVVEKELDAEELQKE----------QKIKEHVST----VNELE 1116
Query: 393 DLKEKYVDLMKQQKSTLLELKASEEELKGFNLIS---ATYEDRLKSLCQKLQEKCDKGSG 449
+L+ ++ KQ + T+ EL+ +++ + L++ A YE +K L QKL K
Sbjct: 1117 ELQLQFQKEKKQLQKTMQELELVKKDAQQTTLMNMEIADYERLMKELNQKLTNKNSTIED 1176
Query: 450 NKHEIALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKS 509
+ E+ I + K ++ + Y E+ +K +K K L
Sbjct: 1177 LEQEMK--------IQKEKQETLQEEITSLQSSVQHYEEKNTK---IKQLLVKTKKELAD 1225
Query: 510 TFVPEGTNAKTAVQSKPEASGSQAKITSKPENLKIKVMTKSDPKSQKIKILKRSEPVHQN 569
E + K E SQ ++ E KI++ + K + + LK S HQ
Sbjct: 1226 AKQAETDHLLLQASLKGELEASQQQV----EVYKIQLAEMTSEKHKIHEHLKTSAEQHQR 1281
Query: 570 LIKPESKIPKQKDQKNKAATASEKTI 595
+ + ++++AA A + +
Sbjct: 1282 TLSAYQQRVVALQEESRAAKAEQAAV 1307
>MY5A_MOUSE (Q99104) Myosin Va (Myosin 5A) (Dilute myosin heavy chain,
non-muscle)
Length = 1853
Score = 48.1 bits (113), Expect = 2e-04
Identities = 110/477 (23%), Positives = 189/477 (39%), Gaps = 71/477 (14%)
Query: 142 DHVSKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHETSKKSKSIAL 201
+ ++K+ + L + R + +IEE D + LVS+LK E +L + E + I
Sbjct: 990 EEIAKLRKDL-EQTRSEKKSIEERADKYKQETDQLVSNLK-EENTLLKQEKETLNHRIVE 1047
Query: 202 PSKGKTSKSSKAYKASESVEESPDGDSD-EDQSVKMAMLSNKLEYLARKQKKFLSKRGSY 260
+K T + + VEE+ + D D+ ++ L N+ L +
Sbjct: 1048 QAKEMTETMER-----KLVEETKQLELDLNDERLRYQNLLNEFSRLEERYDDL------- 1095
Query: 261 KNFKKEDQKGCFNCKKPGHFIADCPDLQKEK---FKGKSKKSSFNSSKFRKQIKK----- 312
KE+ N KPGH D E F + ++ + + + I+K
Sbjct: 1096 ----KEEMTLMLNVPKPGHKRTDSTHSSNESEYTFSSEFAETEDIAPRTEEPIEKKVPLD 1151
Query: 313 -SLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPRQ 371
SL + +E +K+ D+ V S+A E Y + RQ
Sbjct: 1152 MSLFLKLQKRVTELEQEKQLMQDELDRKEEQV--FRSKAKEEERPQIRGAELEYESLKRQ 1209
Query: 372 ELVDSLKELLSLFEHRTNELTD-LKEK------------YVDLMKQQKSTLLELKASEEE 418
EL K+L ++ NEL L EK Y LM+Q S EL +EE
Sbjct: 1210 ELESENKKL----KNELNELRKALSEKSAPEVTAPGAPAYRVLMEQLTSVSEELDVRKEE 1265
Query: 419 LKGFNLISATYEDRLKSLCQKLQEKCDKGSGNKHEIALDDFIMAGIDRSKVASMIYSTYK 478
+ +L S + +Q K DK + I L+D + D+ ++A Y K
Sbjct: 1266 V-------LILRSQLVSQKEAIQPKDDKNTMTDSTILLED-VQKMKDKGEIA-QAYIGLK 1316
Query: 479 NKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTA-----VQSKPEA---SG 530
+ + S+ +S++ S ++ + ++ ++S + E N + +Q PEA +
Sbjct: 1317 ETNR-LLESQLQSQKRSHENEAEALRGEIQS--LKEENNRQQQLLAQNLQLPPEARIEAS 1373
Query: 531 SQAKITS-KPENLKIKVMTKSDPK---SQKIKILKRSEPVHQNLIKPESKIPKQKDQ 583
Q +IT ENL + + DPK S +I + KR + + L K + + K K Q
Sbjct: 1374 LQHEITRLTNENLYFEELYADDPKKYQSYRISLYKRMIDLMEQLEKQDKTVRKLKKQ 1430
>PR4B_HUMAN (Q13523) Serine/threonine-protein kinase PRP4 homolog
(EC 2.7.1.37) (PRP4 pre-mRNA processing factor 4
homolog) (PRP4 kinase)
Length = 1007
Score = 47.4 bits (111), Expect = 3e-04
Identities = 101/460 (21%), Positives = 179/460 (37%), Gaps = 80/460 (17%)
Query: 217 SESVEESPDGDSDEDQSVKMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKK 276
SE +G+ EDQS NK +K+ K SK +K+ +ED+ KK
Sbjct: 20 SEKSINEENGEVSEDQS------QNKHSRHKKKKHKHRSKHKKHKHSSEEDKD-----KK 68
Query: 277 PGHFIADCPDLQKEKFKGKSKKSSFNSSKFR-----KQIKKSLMATWEDLDSESGSDKEE 331
H K K K +K ++S K+ K +A EDL+ + K E
Sbjct: 69 HKH---------KHKHKKHKRKEVIDASDKEGMSPAKRTKLDDLALLEDLEKQRALIKAE 119
Query: 332 ADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKI--------------PRQELVD-- 375
D++ G V + + ES SE+E E++ K + ELVD
Sbjct: 120 LDNELME--GKVQSGMGLILQGYESGSEEEGEIHEKARNGNRSSTRSSSTKGKLELVDNK 177
Query: 376 --SLKELLSLFEHRTNELTDLKEKY--VDLMKQQ--KSTLLELKASEEELKGFNLISATY 429
+ K S + RT +D K+ ++++K++ +S E K S+ K
Sbjct: 178 ITTKKRSKSRSKERTRHRSDKKKSKGGIEIVKEKTTRSKSKERKKSKSPSKRSKSQDQAR 237
Query: 430 EDRLKSLCQKLQEKCDKG-SGNKHEIALDDFIMAGIDRSKVASMIYSTYKNKGKGIGYSE 488
+ + +L ++ QEK K S ++ ++D + DR K + S +++GK
Sbjct: 238 KSKSPTLRRRSQEKIGKARSPTDDKVKIEDKSKSK-DRKKSPIINESRSRDRGK------ 290
Query: 489 EKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKPEASGSQAKITSKPENLKIKVMT 548
K S KD + + ++T + KP S S+ + K EN
Sbjct: 291 ---KSRSPVDLRGKSKDRRSRSKERKSKRSETDKEKKPIKSPSKDASSGK-ENRSPSRRP 346
Query: 549 KSDPKSQKIKILKRSEPVHQNLIKPESKIPKQKDQKNKAATASEKTIPKGVKPKVLN--- 605
PK + + R + S+ P D+++K + + +T+ G + K +
Sbjct: 347 GRSPKRRSLSPKPRDK-------SRRSRSPLLNDRRSKQSKSPSRTLSPGRRAKSRSLER 399
Query: 606 -----DQKPLSIHPKVQGRK---SKTSKANPKGPMKIWVP 637
+++ LS P+ + R S+ ++ P+ W P
Sbjct: 400 KRREPERRRLS-SPRTRPRDDILSRRERSKDASPINRWSP 438
>GOB1_HUMAN (Q14789) Golgi autoantigen, golgin subfamily B member 1
(Giantin) (Macrogolgin) (Golgi complex-associated
protein, 372-kDa) (GCP372)
Length = 3259
Score = 47.4 bits (111), Expect = 3e-04
Identities = 99/508 (19%), Positives = 210/508 (40%), Gaps = 64/508 (12%)
Query: 83 SLCANFEGSKKVKEAKALMLVHQYELFRMKDDESIEEMYSRFQTLVSGLQILKKSY--VS 140
SL EG + KE LV + E + E + + L +IL +SY VS
Sbjct: 1571 SLKLALEGLTEDKEK----LVKEIESLKSSKIAESTEWQEKHKELQKEYEILLQSYENVS 1626
Query: 141 SD-----HVSKILRSLPSRWRPKVTAIEEAKDLNTLSVEDLVSSLKVHEMSLNEHETSKK 195
++ HV + +R K+ + E K +++ ++ + + + SK+
Sbjct: 1627 NEAERIQHVVEAVRQEKQELYGKLRSTEANKKETEKQLQEAEQEMEEMKEKMRKFAKSKQ 1686
Query: 196 SKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSVKMAMLSNKLEY--LARKQKKF 253
K + L + ++ + + A ++ +E + + S+K + K+EY L++K +
Sbjct: 1687 QKILELEEENDRLRA-EVHPAGDTAKECMETLLSSNASMKEELERVKMEYETLSKKFQSL 1745
Query: 254 LSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKGKSKKSSFNSSKFRKQIKKS 313
+S++ S + +D K H I D ++ K+ ++K N + ++ +S
Sbjct: 1746 MSEKDSLSE-EVQDLK---------HQIED--NVSKQANLEATEKHD-NQTNVTEEGTQS 1792
Query: 314 LMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEAESDSEDENEVYSKIPRQEL 373
+ E+ DS S S + + +A ++ AVS+ S ++ N +I
Sbjct: 1793 IPGETEEQDSLSMSTRPTCSESVPSAKS-----ANPAVSKDFSSHDEINNYLQQI----- 1842
Query: 374 VDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLL--------ELKASEEELKGFNLI 425
D LKE ++ E + +++ ++ +K+TLL ELK +EE+ NL+
Sbjct: 1843 -DQLKERIAGLEEEKQK----NKEFSQTLENEKNTLLSQISTKDGELKMLQEEVTKMNLL 1897
Query: 426 SATYEDRLKSLCQKLQEKCDKGSGNKHEI----------ALDDFIMAGIDRSKVASMIYS 475
+ ++ L S KL+E ++ + E ++ ++ D ++ S
Sbjct: 1898 NQQIQEEL-SRVTKLKETAEEEKDDLEERLMNQLAELNGSIGNYCQDVTDAQIKNELLES 1956
Query: 476 TYKNKGKGIGYSEEKSKEYSLKSYCDCIKDGLKSTFVPEGTNAKTAVQSKPEASGSQAKI 535
KN K + EE+ ++ L ++ ++ ++ + A+ +K A Q +
Sbjct: 1957 EMKNLKKCVSELEEEKQQ--LVKEKTKVESEIRKEYLEKIQGAQKEPGNKSHAKELQELL 2014
Query: 536 TSKPENLKIKVMTKSDPKSQKIKILKRS 563
K + +K ++ +KI L+R+
Sbjct: 2015 KEKQQEVK-QLQKDCIRYQEKISALERT 2041
Score = 40.0 bits (92), Expect = 0.054
Identities = 72/347 (20%), Positives = 137/347 (38%), Gaps = 55/347 (15%)
Query: 115 ESIEEMYSRFQTLVSGLQILKKSYVSSDHVSKILRS-------LPSRWRPKVTAIEEAKD 167
ES+++ Q ++GL+ L++ D + K++ L + K A+ + +
Sbjct: 1368 ESLQQKLESSQLQIAGLEHLRELQPKLDELQKLISKKEEDVSYLSGQLSEKEAALTKIQT 1427
Query: 168 LNTLSVEDLVSSLKVH-EMSLNEHETSKKSKSIAL-PSKGKTSKSSKAYKASESVEESPD 225
+ EDL+ +L EM EH+ K + L K K + + +A + ++
Sbjct: 1428 -EIIEQEDLIKALHTQLEMQAKEHDERIKQLQVELCEMKQKPEEIGEESRAKQQIQ---- 1482
Query: 226 GDSDEDQSVKMAMLSNKLEYLARK--QKKFLSKRGSYKNFKK-----EDQKGCFNCKKPG 278
+ ++ A++S K K Q++ RG+ + K E Q N K+
Sbjct: 1483 ------RKLQAALISRKEALKENKSLQEELSLARGTIERLTKSLADVESQVSAQN-KEKD 1535
Query: 279 HFIADCPDLQKEKFK-----------GKSKKSSFNSSKFR----KQIKKSLMATWEDLDS 323
+ LQ+E+ K +S SS S K + K+ L+ E L S
Sbjct: 1536 TVLGRLALLQEERDKLITEMDRSLLENQSLSSSCESLKLALEGLTEDKEKLVKEIESLKS 1595
Query: 324 ESGSDKEEADDDAKAA-------VGLVATVSSEAVS---EAESDSEDENEVYSKIPRQEL 373
++ E + K + VS+EA E+ +++ E+Y K+ E
Sbjct: 1596 SKIAESTEWQEKHKELQKEYEILLQSYENVSNEAERIQHVVEAVRQEKQELYGKLRSTEA 1655
Query: 374 VDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELKASEEELK 420
+ KE + E+ ++KEK K ++ +LEL+ + L+
Sbjct: 1656 --NKKETEKQLQEAEQEMEEMKEKMRKFAKSKQQKILELEEENDRLR 1700
Score = 35.4 bits (80), Expect = 1.3
Identities = 61/307 (19%), Positives = 123/307 (39%), Gaps = 44/307 (14%)
Query: 120 MYSRFQTLVSGLQILKKSYVSSDHVSKILRSLPSRWRPKVTAIEEAKD--LNTLSVEDLV 177
+ SR + L + ++ ++ + ++ +SL +V+A + KD L L++
Sbjct: 1489 LISRKEALKENKSLQEELSLARGTIERLTKSLADV-ESQVSAQNKEKDTVLGRLALLQEE 1547
Query: 178 SSLKVHEMS---LNEHETSKKSKSIALPSKGKTSKSSKAYKASESVEESPDGDSDEDQSV 234
+ EM L S +S+ L +G T K K ES++ S +S E Q
Sbjct: 1548 RDKLITEMDRSLLENQSLSSSCESLKLALEGLTEDKEKLVKEIESLKSSKIAESTEWQE- 1606
Query: 235 KMAMLSNKLEYLARKQKKFLSKRGSYKNFKKEDQKGCFNCKKPGHFIADCPDLQKEKFKG 294
K L + E L + SY+N E ++ + + +K++ G
Sbjct: 1607 KHKELQKEYEILLQ----------SYENVSNEAERI--------QHVVEAVRQEKQELYG 1648
Query: 295 KSKKSSFNSSKFRKQIKKSLMATWEDLDSESGSDKEEADDDAKAAVGLVATVSSEAVSEA 354
K + + N + KQ++++ E+ ++ K + A + + E
Sbjct: 1649 KLRSTEANKKETEKQLQEA----------------EQEMEEMKEKMRKFAKSKQQKILEL 1692
Query: 355 ESDSED-ENEVYSKIPRQELVDSLKELLSLFEHRTNELTDLKEKYVDLMKQQKSTLLELK 413
E +++ EV+ + + ++ LLS EL +K +Y L K+ +S + E
Sbjct: 1693 EEENDRLRAEVHPAGDTAK--ECMETLLSSNASMKEELERVKMEYETLSKKFQSLMSEKD 1750
Query: 414 ASEEELK 420
+ EE++
Sbjct: 1751 SLSEEVQ 1757
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.357 0.156 0.559
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 153,503,090
Number of Sequences: 164201
Number of extensions: 5649447
Number of successful extensions: 47819
Number of sequences better than 10.0: 467
Number of HSP's better than 10.0 without gapping: 34
Number of HSP's successfully gapped in prelim test: 460
Number of HSP's that attempted gapping in prelim test: 45648
Number of HSP's gapped (non-prelim): 1978
length of query: 1664
length of database: 59,974,054
effective HSP length: 124
effective length of query: 1540
effective length of database: 39,613,130
effective search space: 61004220200
effective search space used: 61004220200
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 73 (32.7 bits)
Medicago: description of AC147002.8


