

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147000.4 + phase: 1 /pseudo
(187 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
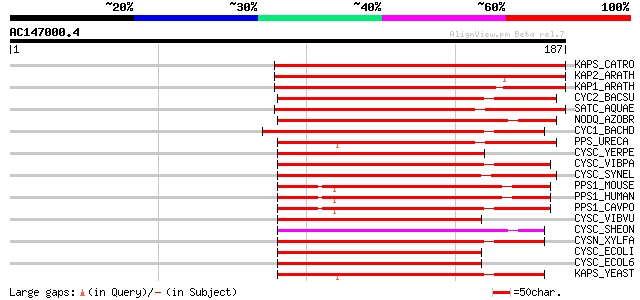
Score E
Sequences producing significant alignments: (bits) Value
KAPS_CATRO (O49204) Adenylyl-sulfate kinase, chloroplast precurs... 156 3e-38
KAP2_ARATH (O49196) Adenylyl-sulfate kinase 2, chloroplast precu... 156 3e-38
KAP1_ARATH (Q43295) Adenylyl-sulfate kinase 1, chloroplast precu... 152 6e-37
CYC2_BACSU (O06735) Probable adenylyl-sulfate kinase (EC 2.7.1.2... 103 2e-22
SATC_AQUAE (O67174) Probable bifunctional SAT/APS kinase [Includ... 99 6e-21
NODQ_AZOBR (P28604) NodQ bifunctional enzyme (Nodulation protein... 94 1e-19
CYC1_BACHD (Q9KCT0) Probable adenylyl-sulfate kinase (EC 2.7.1.2... 90 3e-18
PPS_URECA (Q27128) Bifunctional 3'-phosphoadenosine 5'-phosphosu... 88 1e-17
CYSC_YERPE (Q8ZBP3) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS k... 88 1e-17
CYSC_VIBPA (Q87SX6) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS k... 88 1e-17
CYSC_SYNEL (Q8DJ87) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS k... 87 2e-17
PPS1_MOUSE (Q60967) Bifunctional 3'-phosphoadenosine 5'-phosphos... 87 2e-17
PPS1_HUMAN (O43252) Bifunctional 3'-phosphoadenosine 5'-phosphos... 87 2e-17
PPS1_CAVPO (O54820) Bifunctional 3'-phosphoadenosine 5'-phosphos... 87 2e-17
CYSC_VIBVU (Q8DE75) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS k... 87 2e-17
CYSC_SHEON (Q8EB13) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS k... 86 4e-17
CYSN_XYLFA (Q9PD78) CysN/cysC bifunctional enzyme [Includes: Sul... 86 5e-17
CYSC_ECOLI (P23846) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS k... 86 5e-17
CYSC_ECOL6 (Q8FEJ2) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS k... 86 5e-17
KAPS_YEAST (Q02196) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS k... 85 1e-16
>KAPS_CATRO (O49204) Adenylyl-sulfate kinase, chloroplast precursor
(EC 2.7.1.25) (APS kinase) (Adenosine-5'phosphosulfate
kinase) (ATP adenosine-5'-phosphosulfate
3'-phosphotransferase)
Length = 312
Score = 156 bits (394), Expect = 3e-38
Identities = 75/98 (76%), Positives = 85/98 (86%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+K DACR+LLPEGDFIEVF+DVPL VCEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 214 LISPYRKPPDACRSLLPEGDFIEVFMDVPLKVCEARDPKGLYKLARAGKIKGFTGIDDPY 273
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP EI+L QK C SP D+A+ VISYLE++G+L+
Sbjct: 274 EPPLKSEIVLHQKLGMCDSPCDLADIVISYLEENGYLK 311
>KAP2_ARATH (O49196) Adenylyl-sulfate kinase 2, chloroplast
precursor (EC 2.7.1.25) (APS kinase)
(Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 293
Score = 156 bits (394), Expect = 3e-38
Identities = 71/99 (71%), Positives = 86/99 (86%), Gaps = 1/99 (1%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY++DRDACR+LLP+GDF+EVF+DVPLHVCE+RDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 194 LISPYRRDRDACRSLLPDGDFVEVFMDVPLHVCESRDPKGLYKLARAGKIKGFTGIDDPY 253
Query: 150 EPPCCCEIILQQKGSD-CKSPKDMAETVISYLEKSGHLQ 187
E P CE++L+ G D SP+ MAE +ISYL+ G+L+
Sbjct: 254 EAPVNCEVVLKHTGDDESCSPRQMAENIISYLQNKGYLE 292
>KAP1_ARATH (Q43295) Adenylyl-sulfate kinase 1, chloroplast
precursor (EC 2.7.1.25) (APS kinase)
(Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 276
Score = 152 bits (383), Expect = 6e-37
Identities = 73/98 (74%), Positives = 83/98 (84%), Gaps = 2/98 (2%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+ DRDACR+LLPEGDF+EVF+DVPL VCEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 180 LISPYRTDRDACRSLLPEGDFVEVFMDVPLSVCEARDPKGLYKLARAGKIKGFTGIDDPY 239
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP CEI L ++G SP +MAE V+ YL+ G+LQ
Sbjct: 240 EPPLNCEISLGREGG--TSPIEMAEKVVGYLDNKGYLQ 275
>CYC2_BACSU (O06735) Probable adenylyl-sulfate kinase (EC 2.7.1.25)
(APS kinase) (Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 199
Score = 103 bits (257), Expect = 2e-22
Identities = 50/94 (53%), Positives = 66/94 (70%), Gaps = 3/94 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP+++DRD RAL P+G+F E+++ PLHVCE RDPKGLYK AR G+IK FTGID PYE
Sbjct: 107 ISPFREDRDMVRALFPKGEFFEIYVKCPLHVCEQRDPKGLYKKARNGEIKHFTGIDSPYE 166
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSG 184
P + I++ SD S D A+ +I+ L+ G
Sbjct: 167 APLSPDFIIE---SDQTSISDGADLIINALQNRG 197
>SATC_AQUAE (O67174) Probable bifunctional SAT/APS kinase [Includes:
Sulfate adenylyltransferase (EC 2.7.7.4) (Sulfate
adenylate transferase) (SAT) (ATP-sulfurylase);
Adenylyl-sulfate kinase (EC 2.7.1.25) (APS kinase)
(Adenosine-5'phosphosulfate kinase) (
Length = 546
Score = 99.0 bits (245), Expect = 6e-21
Identities = 46/98 (46%), Positives = 67/98 (67%), Gaps = 3/98 (3%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
L+SPY+ R+ R ++ EG FIEVF+D P+ VCE RD KGLYK A+ G IKGFTG+DDPY
Sbjct: 451 LVSPYRSARNQVRNMMEEGKFIEVFVDAPVEVCEERDVKGLYKKAKEGLIKGFTGVDDPY 510
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP E+ + + +P++ A ++ +L+K G ++
Sbjct: 511 EPPVAPEV---RVDTTKLTPEESALKILEFLKKEGFIK 545
>NODQ_AZOBR (P28604) NodQ bifunctional enzyme (Nodulation protein Q)
[Includes: Sulfate adenylyltransferase subunit 1 (EC
2.7.7.4) (Sulfate adenylate transferase) (SAT)
(ATP-sulfurylase large subunit); Adenylylsulfate kinase
(EC 2.7.1.25) (APS kinase) (AT
Length = 620
Score = 94.4 bits (233), Expect = 1e-19
Identities = 45/94 (47%), Positives = 61/94 (64%), Gaps = 3/94 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
I+P++ +R+A RALLP+G F+EVF+D PL C RDPKGLY ARAG ++ FTG+D PYE
Sbjct: 526 IAPFRAEREAVRALLPDGAFLEVFVDTPLDECMRRDPKGLYAKARAGTLRNFTGVDSPYE 585
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSG 184
P E+ L D + +AE V+ L + G
Sbjct: 586 APDAPELRLDTTAEDADA---LAERVVELLHRKG 616
>CYC1_BACHD (Q9KCT0) Probable adenylyl-sulfate kinase (EC 2.7.1.25)
(APS kinase) (Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 202
Score = 90.1 bits (222), Expect = 3e-18
Identities = 44/95 (46%), Positives = 61/95 (63%), Gaps = 3/95 (3%)
Query: 86 TKDCLISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGI 145
T ISP+++DRD R +L +G+FIEV++ PL CE RDPKGLYK AR+G I FTGI
Sbjct: 104 TSTAFISPFREDRDNVRGILDDGEFIEVYVRCPLETCEKRDPKGLYKKARSGDIPEFTGI 163
Query: 146 DDPYEPPCCCEIILQQKGSDCKSPKDMAETVISYL 180
PYE P E+I+ +D + ++ E + +YL
Sbjct: 164 SSPYEEPVNPELIID---TDQLAVEEAVEKIYAYL 195
>PPS_URECA (Q27128) Bifunctional 3'-phosphoadenosine
5'-phosphosulfate synthethase (PAPS synthethase) (PAPSS)
(Sulfurylase kinase) (SK) [Includes: Sulfate
adenylyltransferase (EC 2.7.7.4) (Sulfate adenylate
transferase) (SAT) (ATP-sulfurylase); Adenylyl-s
Length = 610
Score = 88.2 bits (217), Expect = 1e-17
Identities = 49/96 (51%), Positives = 62/96 (64%), Gaps = 5/96 (5%)
Query: 91 ISPYQKDRDACRALLPEGD--FIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDP 148
ISP+++DRD R+L + F E F+D PL VCE RD KGLYK ARAG+IKGFTGID
Sbjct: 117 ISPFKRDRDLARSLHEQAGLPFFECFVDTPLDVCEQRDVKGLYKKARAGQIKGFTGIDQQ 176
Query: 149 YEPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSG 184
YE P EI Q + KS + + V+S L+K+G
Sbjct: 177 YESPDAPEI---QLYAGNKSIDECVQEVVSLLQKNG 209
>CYSC_YERPE (Q8ZBP3) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS
kinase) (Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 213
Score = 87.8 bits (216), Expect = 1e-17
Identities = 41/70 (58%), Positives = 51/70 (72%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP++ +R + +L G FIEVF+D PL +CEARDPKGLYK ARAG++K FTGID YE
Sbjct: 120 ISPHRAERKMVQDMLASGQFIEVFVDTPLAICEARDPKGLYKKARAGELKNFTGIDSVYE 179
Query: 151 PPCCCEIILQ 160
P +I LQ
Sbjct: 180 SPASPDIHLQ 189
>CYSC_VIBPA (Q87SX6) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS
kinase) (Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 205
Score = 87.8 bits (216), Expect = 1e-17
Identities = 46/92 (50%), Positives = 62/92 (67%), Gaps = 3/92 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP++ +R R LLPEG+FIEVF++ L VCE RDPKGLYK ARAG+I FTGID Y+
Sbjct: 112 ISPHRAERQLVRDLLPEGEFIEVFVNASLEVCEGRDPKGLYKKARAGEIPNFTGIDSEYQ 171
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEK 182
P EI L + KS +++ E ++ L++
Sbjct: 172 APINPEIDLP---AGEKSVEELVELCLNELKQ 200
>CYSC_SYNEL (Q8DJ87) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS
kinase) (Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 211
Score = 87.4 bits (215), Expect = 2e-17
Identities = 42/94 (44%), Positives = 61/94 (64%), Gaps = 3/94 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP++ DR+A R +P GDF+E++ V L CEARD KGLYK ARAG+IK +TGID YE
Sbjct: 117 ISPFRADREAVRNRMPYGDFLEIYCQVSLEACEARDVKGLYKKARAGQIKNYTGIDSAYE 176
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSG 184
P E+++ ++C + ++ V+ L + G
Sbjct: 177 APLKPELVV---NTECLTLEESVSAVLRMLHERG 207
>PPS1_MOUSE (Q60967) Bifunctional 3'-phosphoadenosine
5'-phosphosulfate synthethase 1 (PAPS synthethase 1)
(PAPSS 1) (Sulfurylase kinase 1) (SK1) (SK 1) [Includes:
Sulfate adenylyltransferase (EC 2.7.7.4) (Sulfate
adenylate transferase) (SAT) (ATP-sulfury
Length = 624
Score = 87.0 bits (214), Expect = 2e-17
Identities = 48/95 (50%), Positives = 60/95 (62%), Gaps = 7/95 (7%)
Query: 91 ISPYQKDRDACRALLPEG---DFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDD 147
ISPY +DR+ R + EG F EVF+D PLHVCE RD KGLYK ARAG+IKGFTGID
Sbjct: 132 ISPYTQDRNNARQI-HEGASLPFFEVFVDAPLHVCEQRDVKGLYKKARAGEIKGFTGIDS 190
Query: 148 PYEPPCCCEIILQQKGSDCKSPKDMAETVISYLEK 182
YE P E++L+ D D + V+ L++
Sbjct: 191 EYEKPEAPELVLKTDSCDV---NDCVQQVVELLQE 222
>PPS1_HUMAN (O43252) Bifunctional 3'-phosphoadenosine
5'-phosphosulfate synthethase 1 (PAPS synthethase 1)
(PAPSS 1) (Sulfurylase kinase 1) (SK1) (SK 1) [Includes:
Sulfate adenylyltransferase (EC 2.7.7.4) (Sulfate
adenylate transferase) (SAT) (ATP-sulfury
Length = 624
Score = 87.0 bits (214), Expect = 2e-17
Identities = 48/95 (50%), Positives = 60/95 (62%), Gaps = 7/95 (7%)
Query: 91 ISPYQKDRDACRALLPEG---DFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDD 147
ISPY +DR+ R + EG F EVF+D PLHVCE RD KGLYK ARAG+IKGFTGID
Sbjct: 132 ISPYTQDRNNARQI-HEGASLPFFEVFVDAPLHVCEQRDVKGLYKKARAGEIKGFTGIDS 190
Query: 148 PYEPPCCCEIILQQKGSDCKSPKDMAETVISYLEK 182
YE P E++L+ D D + V+ L++
Sbjct: 191 EYEKPEAPELVLKTDSCDV---NDCVQQVVELLQE 222
>PPS1_CAVPO (O54820) Bifunctional 3'-phosphoadenosine
5'-phosphosulfate synthethase 1 (PAPS synthethase 1)
(PAPSS 1) (Sulfurylase kinase 1) (SK1) (SK 1) [Includes:
Sulfate adenylyltransferase (EC 2.7.7.4) (Sulfate
adenylate transferase) (SAT) (ATP-sulfury
Length = 624
Score = 87.0 bits (214), Expect = 2e-17
Identities = 48/95 (50%), Positives = 61/95 (63%), Gaps = 7/95 (7%)
Query: 91 ISPYQKDRDACRALLPEG---DFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDD 147
ISPY +DR+ R + EG F EVF+D PLHVCE RD KGLYK ARAG+IKGFTGID
Sbjct: 132 ISPYTQDRNNARQI-HEGASLPFFEVFVDAPLHVCEQRDVKGLYKKARAGEIKGFTGIDS 190
Query: 148 PYEPPCCCEIILQQKGSDCKSPKDMAETVISYLEK 182
YE P E++L+ +D D + V+ L++
Sbjct: 191 EYEKPEAPELVLK---TDACDVNDCVQQVVELLQE 222
>CYSC_VIBVU (Q8DE75) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS
kinase) (Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 207
Score = 87.0 bits (214), Expect = 2e-17
Identities = 43/69 (62%), Positives = 50/69 (72%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP++ +R R LLPEG+FIEVF++ L VCE RDPKGLYK ARAG+I FTGID YE
Sbjct: 112 ISPHRAERQLVRDLLPEGEFIEVFVNTSLEVCEQRDPKGLYKKARAGEIANFTGIDSEYE 171
Query: 151 PPCCCEIIL 159
P EI L
Sbjct: 172 VPLNPEIDL 180
>CYSC_SHEON (Q8EB13) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS
kinase) (Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 205
Score = 86.3 bits (212), Expect = 4e-17
Identities = 44/90 (48%), Positives = 53/90 (58%), Gaps = 3/90 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP +++RD+ RA PEG FIEV + PL VCE RDPKGLY AR G+I FTGI PYE
Sbjct: 104 ISPTREERDSIRARFPEGQFIEVHVSTPLSVCELRDPKGLYVKARKGEIAHFTGISSPYE 163
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYL 180
P E+ + D S +I YL
Sbjct: 164 APLSAELTIDTSKGDLAS---QVHALIDYL 190
>CYSN_XYLFA (Q9PD78) CysN/cysC bifunctional enzyme [Includes:
Sulfate adenylyltransferase subunit 1 (EC 2.7.7.4)
(Sulfate adenylate transferase) (SAT) (ATP-sulfurylase
large subunit); Adenylylsulfate kinase (EC 2.7.1.25)
(APS kinase) (ATP adenosine-5'-pho
Length = 623
Score = 85.9 bits (211), Expect = 5e-17
Identities = 45/90 (50%), Positives = 59/90 (65%), Gaps = 3/90 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP++ +R R +F+EVF+DVPL V EARD KGLY ARAG I FTGID PYE
Sbjct: 532 ISPFRDERQLARERFAANEFVEVFVDVPLAVAEARDVKGLYAKARAGLITDFTGIDSPYE 591
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYL 180
PP E+ L+ +D +P+ +A V+S+L
Sbjct: 592 PPPHPELHLR---ADQGTPEQLASQVLSFL 618
>CYSC_ECOLI (P23846) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS
kinase) (Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 200
Score = 85.9 bits (211), Expect = 5e-17
Identities = 41/69 (59%), Positives = 50/69 (72%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP++ +R R + EG FIEVF+D PL +CEARDPKGLYK ARAG+++ FTGID YE
Sbjct: 107 ISPHRAERQMVRERVGEGRFIEVFVDTPLAICEARDPKGLYKKARAGELRNFTGIDSVYE 166
Query: 151 PPCCCEIIL 159
P EI L
Sbjct: 167 APESAEIHL 175
>CYSC_ECOL6 (Q8FEJ2) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS
kinase) (Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 200
Score = 85.9 bits (211), Expect = 5e-17
Identities = 41/69 (59%), Positives = 50/69 (72%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP++ +R R + EG FIEVF+D PL +CEARDPKGLYK ARAG+++ FTGID YE
Sbjct: 107 ISPHRAERQMVRERVGEGRFIEVFVDTPLAICEARDPKGLYKKARAGELRNFTGIDSVYE 166
Query: 151 PPCCCEIIL 159
P EI L
Sbjct: 167 APESAEIHL 175
>KAPS_YEAST (Q02196) Adenylyl-sulfate kinase (EC 2.7.1.25) (APS
kinase) (Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 202
Score = 84.7 bits (208), Expect = 1e-16
Identities = 48/92 (52%), Positives = 58/92 (62%), Gaps = 5/92 (5%)
Query: 91 ISPYQKDRDACRALLPEGD--FIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDP 148
ISPY+ DRD R L E FIE+F+DVPL V E RDPKGLYK AR G IK FTGI P
Sbjct: 104 ISPYRVDRDRARELHKEAGLKFIEIFVDVPLEVAEQRDPKGLYKKAREGVIKEFTGISAP 163
Query: 149 YEPPCCCEIILQQKGSDCKSPKDMAETVISYL 180
YE P E+ L+ +D K+ ++ A + YL
Sbjct: 164 YEAPKAPELHLR---TDQKTVEECATIIYEYL 192
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.340 0.150 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,332,842
Number of Sequences: 164201
Number of extensions: 830234
Number of successful extensions: 2115
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 53
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 2051
Number of HSP's gapped (non-prelim): 55
length of query: 187
length of database: 59,974,054
effective HSP length: 104
effective length of query: 83
effective length of database: 42,897,150
effective search space: 3560463450
effective search space used: 3560463450
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC147000.4


