

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146972.8 + phase: 0
(198 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
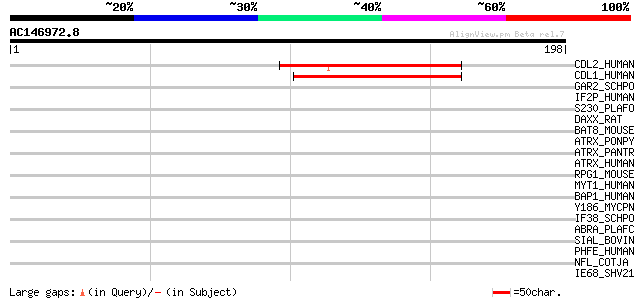
Sequences producing significant alignments: (bits) Value
CDL2_HUMAN (Q9UQ88) PITSLRE serine/threonine-protein kinase CDC2... 52 9e-07
CDL1_HUMAN (P21127) PITSLRE serine/threonine-protein kinase CDC2... 52 9e-07
GAR2_SCHPO (P41891) Protein gar2 42 0.001
IF2P_HUMAN (O60841) Eukaryotic translation initiation factor 5B ... 40 0.003
S230_PLAFO (Q08372) Transmission-blocking target antigen S230 pr... 40 0.003
DAXX_RAT (Q8VIB2) Death domain-associated protein 6 (Daxx) 40 0.003
BAT8_MOUSE (Q9Z148) Histone-lysine N-methyltransferase, H3 lysin... 40 0.003
ATRX_PONPY (Q7YQM3) Transcriptional regulator ATRX (X-linked hel... 40 0.003
ATRX_PANTR (Q7YQM4) Transcriptional regulator ATRX (X-linked hel... 40 0.003
ATRX_HUMAN (P46100) Transcriptional regulator ATRX (X-linked hel... 40 0.003
RPG1_MOUSE (Q9EPQ2) X-linked retinitis pigmentosa GTPase regulat... 40 0.005
MYT1_HUMAN (Q01538) Myelin transcription factor 1 (MYT1) (MYTI) ... 40 0.005
BAP1_HUMAN (Q96QZ7) BAI1-associated protein 1 (BAP-1) (Membrane ... 39 0.006
Y186_MYCPN (P75265) Lipoprotein MG186 homolog precursor (E07_orf... 39 0.008
IF38_SCHPO (O14164) Probable eukaryotic translation initiation f... 39 0.008
ABRA_PLAFC (P22620) 101 kDa malaria antigen (P101) (Acidic basic... 39 0.008
SIAL_BOVIN (Q28862) Bone sialoprotein II precursor (BSP II) (Cel... 39 0.010
PHFE_HUMAN (O94880) PHD finger protein 14 39 0.010
NFL_COTJA (Q02916) Neurofilament triplet L protein (Neurofilamen... 39 0.010
IE68_SHV21 (Q01042) Immediate-early protein 39 0.010
>CDL2_HUMAN (Q9UQ88) PITSLRE serine/threonine-protein kinase CDC2L2
(EC 2.7.1.37) (Galactosyltransferase associated protein
kinase p58/GTA) (Cell division cycle 2-like 2) (CDK11)
Length = 780
Score = 52.0 bits (123), Expect = 9e-07
Identities = 29/68 (42%), Positives = 44/68 (64%), Gaps = 3/68 (4%)
Query: 97 NGMKAKEDPRRAAERE---CEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVI 153
+G + +E+ E E EESEEEEEEEEEE+ +SEE+S +S+ +S+E+ + E
Sbjct: 289 SGSEEEEEEEEEEEEEGSTSEESEEEEEEEEEETGSNSEEASEQSAEEVSEEEMSEDEER 348
Query: 154 EKQNVLVV 161
E +N L+V
Sbjct: 349 ENENHLLV 356
>CDL1_HUMAN (P21127) PITSLRE serine/threonine-protein kinase CDC2L1
(EC 2.7.1.37) (Galactosyltransferase associated protein
kinase p58/GTA) (Cell division cycle 2-like 1) (CLK-1)
(CDK11) (p58 CLK-1)
Length = 795
Score = 52.0 bits (123), Expect = 9e-07
Identities = 26/60 (43%), Positives = 40/60 (66%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQNVLVV 161
+E+ + E EE EEEEEEEEEE+ +SEE+S +S+ +S+E+ + E E +N L+V
Sbjct: 312 EEEEEGSTSEESEEEEEEEEEEEEETGSNSEEASEQSAEEVSEEEMSEDEERENENHLLV 371
Score = 30.8 bits (68), Expect = 2.1
Identities = 18/50 (36%), Positives = 26/50 (52%), Gaps = 2/50 (4%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
K ++ E EEEEEEEEE + E S+SE S +E+ ++ E
Sbjct: 288 KTSSAESSSAESGSGSEEEEEEEEEE--EEEGSTSEESEEEEEEEEEEEE 335
>GAR2_SCHPO (P41891) Protein gar2
Length = 500
Score = 42.0 bits (97), Expect = 0.001
Identities = 38/113 (33%), Positives = 54/113 (47%), Gaps = 15/113 (13%)
Query: 48 SSSSSSSDENHISNNSDATLELNSNISLPYHWEQCLDLKTGEIYYINWRNGMKAKEDPRR 107
SSSS S + S +S + E +S+ S E+ + +KT E K +
Sbjct: 91 SSSSESESSSSESESSSSESESSSSESSSSESEEEVIVKTEE----------KKESSSES 140
Query: 108 AAERECEESEEE----EEEEEEESWYDSEESSSES-STIISKEQYDQREVIEK 155
++ E EE EE EE++E S SE SSSES S S E ++ EV+EK
Sbjct: 141 SSSSESEEEEEAVVKIEEKKESSSDSSSESSSSESESESSSSESEEEEEVVEK 193
Score = 36.2 bits (82), Expect = 0.051
Identities = 34/132 (25%), Positives = 58/132 (43%), Gaps = 7/132 (5%)
Query: 9 ERSLQNCSLNNNNQNEEGSATIDVVAGEGRGGGGGGIGISSSSSSSDENHISNNSDATLE 68
E S ++ S ++++++E S+ + + E S SSSS E + ++ E
Sbjct: 82 ESSSESESESSSSESESSSSESESSSSESESSS------SESSSSESEEEVIVKTEEKKE 135
Query: 69 LNSNISLPYHWEQCLDLKTGEIYYINWRNGMKAKEDPRRAAERECEESEEEEEEEEEESW 128
+S S E+ + +I + + E +E E SE EEEEE E
Sbjct: 136 SSSESSSSSESEEEEEAVV-KIEEKKESSSDSSSESSSSESESESSSSESEEEEEVVEKT 194
Query: 129 YDSEESSSESST 140
+ +E SSESS+
Sbjct: 195 EEKKEGSSESSS 206
Score = 31.6 bits (70), Expect = 1.2
Identities = 20/62 (32%), Positives = 29/62 (46%), Gaps = 2/62 (3%)
Query: 96 RNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESS--SESSTIISKEQYDQREVI 153
+ +K ++ ++ E E E E E S +SE SS SESS+ S + EVI
Sbjct: 68 KKSVKKQKKSKKKEESSSESESESSSSESESSSSESESSSSESESSSSESSSSESEEEVI 127
Query: 154 EK 155
K
Sbjct: 128 VK 129
Score = 30.0 bits (66), Expect = 3.6
Identities = 17/60 (28%), Positives = 28/60 (46%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQNVLVV 161
K ++ ++ EES E E E S +S S SESS+ S+ + E + ++V
Sbjct: 69 KSVKKQKKSKKKEESSSESESESSSSESESSSSESESSSSESESSSSESSSSESEEEVIV 128
Score = 29.6 bits (65), Expect = 4.7
Identities = 17/57 (29%), Positives = 33/57 (57%), Gaps = 5/57 (8%)
Query: 100 KAKEDPRRAAERE-----CEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
K+K++ +RA+ E ++ ++ +++EE S +SE SSSES + S+ + E
Sbjct: 53 KSKKEAKRASSPEPSKKSVKKQKKSKKKEESSSESESESSSSESESSSSESESSSSE 109
>IF2P_HUMAN (O60841) Eukaryotic translation initiation factor 5B
(eIF-5B) (Translation initiation factor IF-2)
Length = 1220
Score = 40.4 bits (93), Expect = 0.003
Identities = 20/58 (34%), Positives = 32/58 (54%)
Query: 99 MKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
++ KE+P E E EE E+EE EEEEE +SE S + ++ D + ++K+
Sbjct: 525 IEVKENPEEEEEEEEEEEEDEESEEEEEEEGESEGSEGDEEDEKVSDEKDSGKTLDKK 582
Score = 32.7 bits (73), Expect = 0.56
Identities = 17/47 (36%), Positives = 26/47 (55%)
Query: 111 RECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQN 157
+E E EEEEEEEEEE EE E + S+ + +V ++++
Sbjct: 528 KENPEEEEEEEEEEEEDEESEEEEEEEGESEGSEGDEEDEKVSDEKD 574
Score = 30.4 bits (67), Expect = 2.8
Identities = 18/63 (28%), Positives = 27/63 (42%)
Query: 88 GEIYYINWRNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQY 147
G +I + + +E+ E + E EEEEEE E E EE S S +
Sbjct: 520 GNTVHIEVKENPEEEEEEEEEEEEDEESEEEEEEEGESEGSEGDEEDEKVSDEKDSGKTL 579
Query: 148 DQR 150
D++
Sbjct: 580 DKK 582
>S230_PLAFO (Q08372) Transmission-blocking target antigen S230
precursor
Length = 3135
Score = 40.0 bits (92), Expect = 0.003
Identities = 22/57 (38%), Positives = 30/57 (52%), Gaps = 3/57 (5%)
Query: 96 RNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDS---EESSSESSTIISKEQYDQ 149
R+ ++ E E EE EEEEEEEEEE YD EES E+ + +E ++
Sbjct: 272 RDNFVIDDEEEEEEEEEEEEEEEEEEEEEEEEEYDDYVYEESGDETEEQLQEEHQEE 328
Score = 37.0 bits (84), Expect = 0.030
Identities = 18/38 (47%), Positives = 22/38 (57%)
Query: 114 EESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
EE EEEEEEEEEE + EE E + +E D+ E
Sbjct: 281 EEEEEEEEEEEEEEEEEEEEEEEEYDDYVYEESGDETE 318
>DAXX_RAT (Q8VIB2) Death domain-associated protein 6 (Daxx)
Length = 731
Score = 40.0 bits (92), Expect = 0.003
Identities = 21/61 (34%), Positives = 37/61 (60%), Gaps = 1/61 (1%)
Query: 97 NGMKAKEDPRRA-AERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEK 155
+G+ ++EDP AE E EE +EE ++EEEE + EE++ + + + Q DQ + E+
Sbjct: 417 SGVASQEDPTTPKAETEDEEDDEESDDEEEEEEEEEEEATEDEDEDLEQLQEDQDDEEEE 476
Query: 156 Q 156
+
Sbjct: 477 E 477
>BAT8_MOUSE (Q9Z148) Histone-lysine N-methyltransferase, H3 lysine-9
specific 3 (EC 2.1.1.43) (Histone H3-K9
methyltransferase 3) (H3-K9-HMTase 3) (HLA-B associated
transcript 8) (G9a) (NG36)
Length = 1263
Score = 40.0 bits (92), Expect = 0.003
Identities = 30/94 (31%), Positives = 41/94 (42%), Gaps = 4/94 (4%)
Query: 47 ISSSSSSSDENHISNNSDATLELNSNISLPYHWEQCLDLKTG---EIYYINWRNGMKAKE 103
++S S S D + D LE L WE + + Y ++ R +K
Sbjct: 283 LNSGSLSEDLGSAGGSGDIILEKGEPRPLE-EWETVVGDDFSLYYDAYSVDERVDSDSKS 341
Query: 104 DPRRAAERECEESEEEEEEEEEESWYDSEESSSE 137
+ AE+ EE EEEEEEEEEE + EE E
Sbjct: 342 EVEALAEQLSEEEEEEEEEEEEEEEEEEEEEEEE 375
Score = 37.0 bits (84), Expect = 0.030
Identities = 18/29 (62%), Positives = 21/29 (72%)
Query: 110 ERECEESEEEEEEEEEESWYDSEESSSES 138
E E EE EEEEEEEEEE + EES ++S
Sbjct: 356 EEEEEEEEEEEEEEEEEEEEEDEESGNQS 384
Score = 36.2 bits (82), Expect = 0.051
Identities = 18/35 (51%), Positives = 22/35 (62%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSS 136
+E+ E E EE EEEEEEE+EES S+ S S
Sbjct: 355 EEEEEEEEEEEEEEEEEEEEEEDEESGNQSDRSGS 389
Score = 33.5 bits (75), Expect = 0.33
Identities = 17/46 (36%), Positives = 26/46 (55%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQY 147
+E+ E E EE EEEEEEEE+E + + S S +K+++
Sbjct: 354 EEEEEEEEEEEEEEEEEEEEEEEDEESGNQSDRSGSSGRRKAKKKW 399
Score = 33.5 bits (75), Expect = 0.33
Identities = 17/41 (41%), Positives = 23/41 (55%)
Query: 99 MKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESS 139
+ +E+ E E EE EEEEEEEEE+ ++ S SS
Sbjct: 350 LSEEEEEEEEEEEEEEEEEEEEEEEEEDEESGNQSDRSGSS 390
>ATRX_PONPY (Q7YQM3) Transcriptional regulator ATRX (X-linked helicase
II) (X-linked nuclear protein) (XNP)
Length = 2492
Score = 40.0 bits (92), Expect = 0.003
Identities = 36/115 (31%), Positives = 49/115 (42%), Gaps = 6/115 (5%)
Query: 46 GISSS-SSSSDENHISNNSDATLELNSNISLPYHWEQCLDLKTGEIYYINWRNGMKAKED 104
G+S S S DE S EL N ++ +K E ++ + +E+
Sbjct: 1388 GVSEEVSESEDEQRPRTRSAKKAELEENQRSYKQKKKRRRIKVQEDSSSENKSNSEEEEE 1447
Query: 105 PRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQNVL 159
E+E EE EEEEEEEEEE D +S + I K D + E QN L
Sbjct: 1448 -----EKEEEEEEEEEEEEEEEDENDDSKSPGKGRKKIRKILKDDKLRTETQNAL 1497
>ATRX_PANTR (Q7YQM4) Transcriptional regulator ATRX (X-linked helicase
II) (X-linked nuclear protein) (XNP)
Length = 2492
Score = 40.0 bits (92), Expect = 0.003
Identities = 36/115 (31%), Positives = 49/115 (42%), Gaps = 6/115 (5%)
Query: 46 GISSS-SSSSDENHISNNSDATLELNSNISLPYHWEQCLDLKTGEIYYINWRNGMKAKED 104
G+S S S DE S EL N ++ +K E ++ + +E+
Sbjct: 1388 GVSEEVSESEDEQRPRTRSAKKAELEENQRSYKQKKKRRRIKVQEDSSSENKSNSEEEEE 1447
Query: 105 PRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQNVL 159
E+E EE EEEEEEEEEE D +S + I K D + E QN L
Sbjct: 1448 -----EKEEEEEEEEEEEEEEEDENDDSKSPGKGRKKIRKILKDDKLRTETQNAL 1497
>ATRX_HUMAN (P46100) Transcriptional regulator ATRX (X-linked helicase
II) (X-linked nuclear protein) (XNP) (Znf-HX)
Length = 2492
Score = 40.0 bits (92), Expect = 0.003
Identities = 36/115 (31%), Positives = 49/115 (42%), Gaps = 6/115 (5%)
Query: 46 GISSS-SSSSDENHISNNSDATLELNSNISLPYHWEQCLDLKTGEIYYINWRNGMKAKED 104
G+S S S DE S EL N ++ +K E ++ + +E+
Sbjct: 1388 GVSEEVSESEDEQRPRTRSAKKAELEENQRSYKQKKKRRRIKVQEDSSSENKSNSEEEEE 1447
Query: 105 PRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQNVL 159
E+E EE EEEEEEEEEE D +S + I K D + E QN L
Sbjct: 1448 -----EKEEEEEEEEEEEEEEEDENDDSKSPGKGRKKIRKILKDDKLRTETQNAL 1497
>RPG1_MOUSE (Q9EPQ2) X-linked retinitis pigmentosa GTPase
regulator-interacting protein 1 (RPGR-interacting
protein 1)
Length = 1331
Score = 39.7 bits (91), Expect = 0.005
Identities = 21/56 (37%), Positives = 32/56 (56%), Gaps = 1/56 (1%)
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQ-REVIE 154
+ KE+ E E EE EE+EEE+EEE D +E E +E+ D+ ++V+E
Sbjct: 938 EVKEEKEEEEEEEREEEEEKEEEKEEEEEEDEKEEEEEEEEEEEEEEEDENKDVLE 993
Score = 38.5 bits (88), Expect = 0.010
Identities = 19/55 (34%), Positives = 29/55 (52%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
+E+ E E +E +EEEEEEE E + EE E KE+ ++ E E++
Sbjct: 928 EEEEEEEEEEEVKEEKEEEEEEEREEEEEKEEEKEEEEEEDEKEEEEEEEEEEEE 982
Score = 37.4 bits (85), Expect = 0.023
Identities = 19/54 (35%), Positives = 28/54 (51%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEK 155
+E+ + E EE EEEE EEEEE + EE E +E+ ++ E E+
Sbjct: 932 EEEEEEEVKEEKEEEEEEEREEEEEKEEEKEEEEEEDEKEEEEEEEEEEEEEEE 985
Score = 36.6 bits (83), Expect = 0.039
Identities = 25/98 (25%), Positives = 47/98 (47%), Gaps = 1/98 (1%)
Query: 61 NNSDATLELNSNISLPYHWEQCLDLKTG-EIYYINWRNGMKAKEDPRRAAERECEESEEE 119
N +D+ + N +I + W+ G ++ G + +E+ +E E EEE
Sbjct: 871 NLTDSGEKSNGSIKVQLDWKSHYLAPEGFQMSEAEKPEGEEKEEEGGEEEVKEEEVEEEE 930
Query: 120 EEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQN 157
EEEEEEE + +E E +E+ +++E E+++
Sbjct: 931 EEEEEEEEVKEEKEEEEEEEREEEEEKEEEKEEEEEED 968
Score = 36.6 bits (83), Expect = 0.039
Identities = 19/58 (32%), Positives = 32/58 (54%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQNVL 159
+E+ + + E EE E EEEEE+EE + EE + +E+ ++ E E ++VL
Sbjct: 935 EEEEVKEEKEEEEEEEREEEEEKEEEKEEEEEEDEKEEEEEEEEEEEEEEEDENKDVL 992
Score = 36.2 bits (82), Expect = 0.051
Identities = 19/57 (33%), Positives = 31/57 (54%)
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
+ KE+ E E EE EE +EE+EEE + EE + +E+ D++E E++
Sbjct: 920 EVKEEEVEEEEEEEEEEEEVKEEKEEEEEEEREEEEEKEEEKEEEEEEDEKEEEEEE 976
Score = 35.8 bits (81), Expect = 0.066
Identities = 19/58 (32%), Positives = 29/58 (49%)
Query: 99 MKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
++ +E+ E EE EEEEEEE EE EE E +E+ ++ E E++
Sbjct: 926 VEEEEEEEEEEEEVKEEKEEEEEEEREEEEEKEEEKEEEEEEDEKEEEEEEEEEEEEE 983
Score = 35.4 bits (80), Expect = 0.086
Identities = 20/54 (37%), Positives = 29/54 (53%)
Query: 103 EDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
E+ + E E EE EEEEEEE +E + EE E +E+ ++ E EK+
Sbjct: 918 EEEVKEEEVEEEEEEEEEEEEVKEEKEEEEEEEREEEEEKEEEKEEEEEEDEKE 971
Score = 35.4 bits (80), Expect = 0.086
Identities = 18/56 (32%), Positives = 28/56 (49%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQN 157
+E+ E + EE EEE EEEEE+ EE + +E+ ++ E E +N
Sbjct: 933 EEEEEEVKEEKEEEEEEEREEEEEKEEEKEEEEEEDEKEEEEEEEEEEEEEEEDEN 988
Score = 35.0 bits (79), Expect = 0.11
Identities = 19/47 (40%), Positives = 26/47 (54%), Gaps = 5/47 (10%)
Query: 96 RNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTII 142
R + KE+ + E E E+ EEEEEEEEEE EE E+ ++
Sbjct: 951 REEEEEKEEEKEEEEEEDEKEEEEEEEEEEE-----EEEEDENKDVL 992
Score = 34.3 bits (77), Expect = 0.19
Identities = 17/56 (30%), Positives = 30/56 (53%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQN 157
+E+ +E +E EEEEE EEEE + +E E +E+ ++ E E+++
Sbjct: 931 EEEEEEEEVKEEKEEEEEEEREEEEEKEEEKEEEEEEDEKEEEEEEEEEEEEEEED 986
Score = 33.5 bits (75), Expect = 0.33
Identities = 18/55 (32%), Positives = 28/55 (50%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
+E E E EE EEEEE +EE+ + EE E KE+ ++ + E++
Sbjct: 919 EEVKEEEVEEEEEEEEEEEEVKEEKEEEEEEEREEEEEKEEEKEEEEEEDEKEEE 973
Score = 32.3 bits (72), Expect = 0.73
Identities = 18/55 (32%), Positives = 28/55 (50%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
+E+ E E EE E +EE+EEEE EE E +E+ ++ E E++
Sbjct: 923 EEEVEEEEEEEEEEEEVKEEKEEEEEEEREEEEEKEEEKEEEEEEDEKEEEEEEE 977
>MYT1_HUMAN (Q01538) Myelin transcription factor 1 (MYT1) (MYTI)
(Proteolipid protein binding protein) (PLPB1)
Length = 1121
Score = 39.7 bits (91), Expect = 0.005
Identities = 20/56 (35%), Positives = 30/56 (52%)
Query: 96 RNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
+ G+ + E+ E E EE EE+EEEEEEE + EE E +E+ ++ E
Sbjct: 251 QKGILSHEEEDEEEEEEEEEEEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 306
Score = 39.7 bits (91), Expect = 0.005
Identities = 20/44 (45%), Positives = 25/44 (56%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKE 145
+ED E E EE EEEEEEEEEE + EE + +I +E
Sbjct: 272 EEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEAAPDVIFQE 315
Score = 38.1 bits (87), Expect = 0.013
Identities = 19/47 (40%), Positives = 27/47 (57%)
Query: 110 ERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
E E EE EEEEEEEEE+ + EE E +E+ ++ E E++
Sbjct: 258 EEEDEEEEEEEEEEEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 304
Score = 37.4 bits (85), Expect = 0.023
Identities = 18/38 (47%), Positives = 23/38 (60%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESS 139
+E+ E E EE EEEEEEEEEE + EE E++
Sbjct: 271 EEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEAA 308
Score = 37.0 bits (84), Expect = 0.030
Identities = 19/47 (40%), Positives = 27/47 (57%)
Query: 110 ERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
E E EE EEEEEEE+EE + EE E +E+ ++ E E++
Sbjct: 260 EDEEEEEEEEEEEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 306
Score = 37.0 bits (84), Expect = 0.030
Identities = 20/55 (36%), Positives = 29/55 (52%)
Query: 103 EDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQN 157
E+ E E E+ EEEEEEEEEE + EE E +E+ +VI +++
Sbjct: 262 EEEEEEEEEEEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEAAPDVIFQED 316
Score = 36.6 bits (83), Expect = 0.039
Identities = 18/47 (38%), Positives = 27/47 (57%)
Query: 110 ERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
E + EE EEEEEEEE+E + EE E +E+ ++ E E++
Sbjct: 259 EEDEEEEEEEEEEEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 305
Score = 36.2 bits (82), Expect = 0.051
Identities = 18/48 (37%), Positives = 27/48 (55%)
Query: 109 AERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
+ E +E EEEEEEEEEE + EE E +E+ ++ E E++
Sbjct: 256 SHEEEDEEEEEEEEEEEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 303
Score = 36.2 bits (82), Expect = 0.051
Identities = 18/38 (47%), Positives = 22/38 (57%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESS 139
+E+ E E EE EEEEEEEEEE + EE E +
Sbjct: 270 EEEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEA 307
Score = 35.8 bits (81), Expect = 0.066
Identities = 17/44 (38%), Positives = 24/44 (53%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKE 145
+++ E E EE EEEEEEEEEE + EE + I ++
Sbjct: 273 EDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEAAPDVIFQED 316
Score = 33.1 bits (74), Expect = 0.43
Identities = 19/40 (47%), Positives = 25/40 (62%), Gaps = 2/40 (5%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYD--SEESSSESS 139
+E+ E E EE EEEEEEEEEE+ D +E +S +S
Sbjct: 282 EEEEEEEEEEEEEEEEEEEEEEEEEAAPDVIFQEDTSHTS 321
>BAP1_HUMAN (Q96QZ7) BAI1-associated protein 1 (BAP-1) (Membrane
associated guanylate kinase inverted-1) (MAGI-1)
(Atrophin-1 interacting protein 3) (AIP3) (WW
domain-containing protein 3) (WWP3) (Trinucleotide
repeat-containing gene 19 protein)
Length = 1491
Score = 39.3 bits (90), Expect = 0.006
Identities = 20/83 (24%), Positives = 47/83 (56%), Gaps = 7/83 (8%)
Query: 66 TLELNSNISLPYHWEQCLDLKTGEIYYINWRNGMKAKEDP------RRAAERECEESEEE 119
T EL+S + LP WE+ D G IYY++ N E+P ++ E++ ++ +++
Sbjct: 352 TEELDSELELPAGWEKIEDPVYG-IYYVDHINRKTQYENPVLEAKRKKQLEQQQQQQQQQ 410
Query: 120 EEEEEEESWYDSEESSSESSTII 142
+++++++ +EE + + S ++
Sbjct: 411 QQQQQQQQQQQTEEWTEDHSALV 433
Score = 29.6 bits (65), Expect = 4.7
Identities = 20/60 (33%), Positives = 30/60 (49%), Gaps = 5/60 (8%)
Query: 75 LPYHWEQCLDLKTGEIYYINWRNGMKAKEDPR--RAAERECEESEEEEEEEEEESWYDSE 132
LP +WE + GE+Y+I+ + DPR ++ EE E++E EE DSE
Sbjct: 302 LPENWEMAYT-ENGEVYFIDHNTKTTSWLDPRCLNKQQKPLEECEDDEGVHTEE--LDSE 358
>Y186_MYCPN (P75265) Lipoprotein MG186 homolog precursor
(E07_orf301)
Length = 301
Score = 38.9 bits (89), Expect = 0.008
Identities = 23/66 (34%), Positives = 41/66 (61%), Gaps = 7/66 (10%)
Query: 85 LKTGEIYYINWRNGMK-----AKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESS 139
LK I Y WR+G A E+ R+ AE++ +E+ ++E+++EE+ DS++SSS S+
Sbjct: 47 LKPATIEY--WRDGDTPEINYASEERRKEAEQKSKENAKKEDKKEEKKTEDSQDSSSAST 104
Query: 140 TIISKE 145
+ S +
Sbjct: 105 QVRSSK 110
>IF38_SCHPO (O14164) Probable eukaryotic translation initiation
factor 3 93 kDa subunit (eIF3 p93)
Length = 918
Score = 38.9 bits (89), Expect = 0.008
Identities = 34/122 (27%), Positives = 57/122 (45%), Gaps = 16/122 (13%)
Query: 41 GGGGIGISSSSSSSDENHI-----------SNNSDATLELNSNISLPYHWEQCLDLKTGE 89
GG + S SS+EN + S+ +++ E +++ S E+ + + E
Sbjct: 7 GGSSDSDAESVDSSEENRLTSSRLKKQDDSSSEEESSEEESASSSESESSEEESESEESE 66
Query: 90 IYYINWRNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQ 149
+ + + + A ED +E + E SEEEEE E EE S+ES SES + E+ +
Sbjct: 67 VE-VPKKKAVAASED----SESDSESSEEEEETESEEDSEVSDESESESESESESEEESE 121
Query: 150 RE 151
E
Sbjct: 122 SE 123
Score = 34.3 bits (77), Expect = 0.19
Identities = 28/103 (27%), Positives = 45/103 (43%), Gaps = 8/103 (7%)
Query: 48 SSSSSSSDENHISNNSDATLELNSNISLPYHWEQCLDLKTGEIYYINWRNGMKAKEDPRR 107
SS S SS+E S S+ + ++ E + E +++ED
Sbjct: 50 SSESESSEEESESEESEVEVPKKKAVAASEDSESDSESSEEE-------EETESEEDSEV 102
Query: 108 AAERECEESEEEEEEEEEESWYDSEESS-SESSTIISKEQYDQ 149
+ E E E E E EEE ES +S+ES S S+ + K + ++
Sbjct: 103 SDESESESESESESEEESESEEESDESERSGPSSFLKKPEKEE 145
>ABRA_PLAFC (P22620) 101 kDa malaria antigen (P101) (Acidic basic
repeat antigen)
Length = 743
Score = 38.9 bits (89), Expect = 0.008
Identities = 20/59 (33%), Positives = 32/59 (53%)
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQNV 158
K KE + E+E EE E+EE+E+E+E +E E +EQ ++ E I +N+
Sbjct: 679 KEKEKEKEKEEKEKEEKEKEEKEKEKEEKEKEKEEKEEEKKEKEEEQEEEEEEIVPENL 737
Score = 35.4 bits (80), Expect = 0.086
Identities = 20/60 (33%), Positives = 35/60 (58%), Gaps = 1/60 (1%)
Query: 100 KAKEDPRRAAERECEESEEEE-EEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQNV 158
K +E+ + E+E EE E+EE E+EE+E + +E E KE+ +++E E++ V
Sbjct: 674 KEEEEKEKEKEKEKEEKEKEEKEKEEKEKEKEEKEKEKEEKEEEKKEKEEEQEEEEEEIV 733
Score = 33.9 bits (76), Expect = 0.25
Identities = 17/39 (43%), Positives = 24/39 (60%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESST 140
KE + E+E EE EEE++E+EEE + EE E+ T
Sbjct: 700 KEKEKEEKEKEKEEKEEEKKEKEEEQEEEEEEIVPENLT 738
Score = 33.1 bits (74), Expect = 0.43
Identities = 19/62 (30%), Positives = 34/62 (54%), Gaps = 1/62 (1%)
Query: 100 KAKEDPRRAAERECEESEEE-EEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQNV 158
K KE+ + E+E E+ E+E EE+E+EE + EE E +++ + E E++
Sbjct: 672 KNKEEEEKEKEKEKEKEEKEKEEKEKEEKEKEKEEKEKEKEEKEEEKKEKEEEQEEEEEE 731
Query: 159 LV 160
+V
Sbjct: 732 IV 733
Score = 31.6 bits (70), Expect = 1.2
Identities = 15/52 (28%), Positives = 28/52 (53%)
Query: 105 PRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
P+ E E E+ +E+E+EE+E+ + EE E +++ + E EK+
Sbjct: 671 PKNKEEEEKEKEKEKEKEEKEKEEKEKEEKEKEKEEKEKEKEEKEEEKKEKE 722
Score = 31.2 bits (69), Expect = 1.6
Identities = 16/52 (30%), Positives = 26/52 (49%)
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
K KE+ + + + +E +E+E+EE+EE + EE E I E E
Sbjct: 690 KEKEEKEKEEKEKEKEEKEKEKEEKEEEKKEKEEEQEEEEEEIVPENLTTEE 741
Score = 30.4 bits (67), Expect = 2.8
Identities = 15/38 (39%), Positives = 23/38 (60%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESS 139
+E + E+E E+ E+EEE+EEEE E ++E S
Sbjct: 705 EEKEKEKEEKEEEKKEKEEEQEEEEEEIVPENLTTEES 742
Score = 30.0 bits (66), Expect = 3.6
Identities = 16/58 (27%), Positives = 31/58 (52%)
Query: 99 MKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
++A+ D +E EE E+E+E+E+EE + +E + KE+ + + EK+
Sbjct: 662 IEAEVDALAPKNKEEEEKEKEKEKEKEEKEKEEKEKEEKEKEKEEKEKEKEEKEEEKK 719
>SIAL_BOVIN (Q28862) Bone sialoprotein II precursor (BSP II)
(Cell-binding sialoprotein) (Integrin-binding
sialoprotein)
Length = 310
Score = 38.5 bits (88), Expect = 0.010
Identities = 41/151 (27%), Positives = 63/151 (41%), Gaps = 31/151 (20%)
Query: 3 AITASLERSLQNCSLNNNNQNEEGSATIDVVAGEGRGGGGGGIGISSSSSSSDENHISNN 62
A+ +S + S +N + +++ + EE T + EG GG S + +E+ N
Sbjct: 59 AVQSSSDSSEENGNGDSSEEEEEEEETSNE---EGNNGGNE----DSDENEDEESEAENT 111
Query: 63 SDATLELNSNISLPYHWEQCLDLKTGEIYYINWRNGMKAKEDPRRAA--------ERECE 114
+ +T L P TG+I G+ A PR+A E E +
Sbjct: 112 TLSTTTLGYGEITP---------GTGDI-------GLAAIWLPRKAGATGKKATKEDESD 155
Query: 115 ESEEEEEEEEEESWYDSEESSSESSTIISKE 145
E EEEEEEEE E+ D E ++ S E
Sbjct: 156 EEEEEEEEEENEAEVDDNEQGINGTSSNSTE 186
>PHFE_HUMAN (O94880) PHD finger protein 14
Length = 888
Score = 38.5 bits (88), Expect = 0.010
Identities = 27/117 (23%), Positives = 48/117 (40%)
Query: 25 EGSATIDVVAGEGRGGGGGGIGISSSSSSSDENHISNNSDATLELNSNISLPYHWEQCLD 84
+ S D G+ G G G +S S E S++ + LE N + EQ +
Sbjct: 24 DSSDDSDFKVGDASDSEGSGNGSEDASKDSGEGSCSDSEENILEEELNEDIKVKEEQLKN 83
Query: 85 LKTGEIYYINWRNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTI 141
E+ + K++ ER ++ E+E+E+E+E+ E E +T+
Sbjct: 84 SAEEEVLSSEKQLIKMEKKEEEENGERPRKKREKEKEKEKEKEKEKEREKEKEKATV 140
>NFL_COTJA (Q02916) Neurofilament triplet L protein (Neurofilament
light polypeptide) (NF-L)
Length = 555
Score = 38.5 bits (88), Expect = 0.010
Identities = 20/63 (31%), Positives = 34/63 (53%)
Query: 98 GMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQN 157
G + E+ A+ E EE +E EEEE EE+ + E S E++ +E+ +++E K+
Sbjct: 489 GGEEAEEEEEGAKEESEEGKEGEEEEGEETAAEDGEESQETAEETGEEEKEEKEAAGKEE 548
Query: 158 VLV 160
V
Sbjct: 549 TEV 551
Score = 30.8 bits (68), Expect = 2.1
Identities = 19/51 (37%), Positives = 26/51 (50%)
Query: 101 AKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
AK +AA E E EE+EE EEEE+ + E E + S+E + E
Sbjct: 462 AKAGEAKAAPAEEGEEEEKEEGEEEEAGGEEAEEEEEGAKEESEEGKEGEE 512
Score = 30.8 bits (68), Expect = 2.1
Identities = 19/55 (34%), Positives = 26/55 (46%)
Query: 101 AKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEK 155
AKE+ E E EE EE E+ EES +EE+ E ++ EV +K
Sbjct: 500 AKEESEEGKEGEEEEGEETAAEDGEESQETAEETGEEEKEEKEAAGKEETEVKKK 554
Score = 30.4 bits (67), Expect = 2.8
Identities = 19/70 (27%), Positives = 31/70 (44%)
Query: 86 KTGEIYYINWRNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKE 145
K GE G + +++ E EE+EEEEE +EES E E +++
Sbjct: 463 KAGEAKAAPAEEGEEEEKEEGEEEEAGGEEAEEEEEGAKEESEEGKEGEEEEGEETAAED 522
Query: 146 QYDQREVIEK 155
+ +E E+
Sbjct: 523 GEESQETAEE 532
>IE68_SHV21 (Q01042) Immediate-early protein
Length = 407
Score = 38.5 bits (88), Expect = 0.010
Identities = 19/54 (35%), Positives = 31/54 (57%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEK 155
+E+ A E E EE+EEE EEEE E + E +E + ++E+ ++ E E+
Sbjct: 140 EEEAEEAEEEEAEEAEEEAEEEEAEEEAEEEAEEAEEAEEEAEEEAEEAEEAEE 193
Score = 37.7 bits (86), Expect = 0.017
Identities = 20/57 (35%), Positives = 29/57 (50%)
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
+A+E A E E EE+E EEEE EEE + E +E E+ + E E++
Sbjct: 110 EAEEAEEEAEEEEAEEAEAEEEEAEEEEAEEEEAEEAEEEEAEEAEEEAEEEEAEEE 166
Score = 37.0 bits (84), Expect = 0.030
Identities = 19/55 (34%), Positives = 31/55 (55%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
+E+ AE E EE+EE EEE EEE+ E +E + ++E ++ E E++
Sbjct: 160 EEEAEEEAEEEAEEAEEAEEEAEEEAEEAEEAEEAEEAEEEAEEAEEEAEEAEEE 214
Score = 37.0 bits (84), Expect = 0.030
Identities = 19/52 (36%), Positives = 30/52 (57%)
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
+A+E+ E E EE+EE EEEE EE+ ++EE +E E+ ++ E
Sbjct: 127 EAEEEEAEEEEAEEEEAEEAEEEEAEEAEEEAEEEEAEEEAEEEAEEAEEAE 178
Score = 37.0 bits (84), Expect = 0.030
Identities = 19/52 (36%), Positives = 29/52 (55%)
Query: 104 DPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEK 155
+ A E E EE E EEEE EEE ++EE +E + ++E+ + E E+
Sbjct: 119 EEEEAEEAEAEEEEAEEEEAEEEEAEEAEEEEAEEAEEEAEEEEAEEEAEEE 170
Score = 36.6 bits (83), Expect = 0.039
Identities = 20/54 (37%), Positives = 31/54 (57%), Gaps = 1/54 (1%)
Query: 104 DPRRAAERECEESEEE-EEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
+ + A E E EE+EEE EEEE EE+ + EE+ E + E+ ++ E E +
Sbjct: 102 EEKEAEEEEAEEAEEEAEEEEAEEAEAEEEEAEEEEAEEEEAEEAEEEEAEEAE 155
Score = 36.2 bits (82), Expect = 0.051
Identities = 18/50 (36%), Positives = 28/50 (56%)
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQ 149
+A+E A E E EE EEE EE EE+ ++EE + E+ E+ ++
Sbjct: 150 EAEEAEEEAEEEEAEEEAEEEAEEAEEAEEEAEEEAEEAEEAEEAEEAEE 199
Score = 35.4 bits (80), Expect = 0.086
Identities = 19/56 (33%), Positives = 28/56 (49%)
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEK 155
+A+E+ AE E EE E EEE EEE + E +E ++E + E E+
Sbjct: 145 EAEEEEAEEAEEEAEEEEAEEEAEEEAEEAEEAEEEAEEEAEEAEEAEEAEEAEEE 200
Score = 35.4 bits (80), Expect = 0.086
Identities = 22/64 (34%), Positives = 36/64 (55%), Gaps = 7/64 (10%)
Query: 100 KAKEDPRRAAERECEESEEEEEE-------EEEESWYDSEESSSESSTIISKEQYDQREV 152
+A+E+ AE E EE+EEEE E EEEE+ EE+ E + ++E+ ++ E
Sbjct: 117 EAEEEEAEEAEAEEEEAEEEEAEEEEAEEAEEEEAEEAEEEAEEEEAEEEAEEEAEEAEE 176
Query: 153 IEKQ 156
E++
Sbjct: 177 AEEE 180
Score = 33.9 bits (76), Expect = 0.25
Identities = 17/50 (34%), Positives = 27/50 (54%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
+E AE E EE EE EE EEE+ ++EE+ ++E+ ++ E
Sbjct: 156 EEAEEEEAEEEAEEEAEEAEEAEEEAEEEAEEAEEAEEAEEAEEEAEEAE 205
Score = 33.5 bits (75), Expect = 0.33
Identities = 19/54 (35%), Positives = 29/54 (53%), Gaps = 5/54 (9%)
Query: 103 EDPRRAAERECEESEEEEE-----EEEEESWYDSEESSSESSTIISKEQYDQRE 151
E+ AE E EE+EE EE EE EE+ ++EE+ E+ E+ ++ E
Sbjct: 175 EEAEEEAEEEAEEAEEAEEAEEAEEEAEEAEEEAEEAEEEAEEAEEAEEAEEAE 228
Score = 33.5 bits (75), Expect = 0.33
Identities = 18/52 (34%), Positives = 27/52 (51%)
Query: 104 DPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEK 155
+ A E E EE+EE EEE EEE + E +E + +E ++ E E+
Sbjct: 139 EEEEAEEAEEEEAEEAEEEAEEEEAEEEAEEEAEEAEEAEEEAEEEAEEAEE 190
Score = 33.5 bits (75), Expect = 0.33
Identities = 21/62 (33%), Positives = 35/62 (55%), Gaps = 2/62 (3%)
Query: 96 RNGMKAKE-DPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIE 154
R G +E + R AE E E +E EEEE EE+ ++EE +E + +E+ ++ E E
Sbjct: 82 REGEGGEEGEGREEAEEEEAEEKEAEEEEAEEAEEEAEEEEAEEAE-AEEEEAEEEEAEE 140
Query: 155 KQ 156
++
Sbjct: 141 EE 142
Score = 33.5 bits (75), Expect = 0.33
Identities = 19/54 (35%), Positives = 26/54 (47%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEK 155
KE AE EE+EEEE EE E ++EE +E E+ + E E+
Sbjct: 104 KEAEEEEAEEAEEEAEEEEAEEAEAEEEEAEEEEAEEEEAEEAEEEEAEEAEEE 157
Score = 33.1 bits (74), Expect = 0.43
Identities = 19/57 (33%), Positives = 29/57 (50%)
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
+A+E AE EE+EEEE EEE E + E + E + ++E + E E +
Sbjct: 142 EAEEAEEEEAEEAEEEAEEEEAEEEAEEEAEEAEEAEEEAEEEAEEAEEAEEAEEAE 198
Score = 33.1 bits (74), Expect = 0.43
Identities = 19/57 (33%), Positives = 31/57 (54%), Gaps = 4/57 (7%)
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
+A+E+ E E EE+EE EEE EEE ++E E+ +E+ ++ E E +
Sbjct: 95 EAEEEEAEEKEAEEEEAEEAEEEAEEEEAEEAEAEEEEA----EEEEAEEEEAEEAE 147
Score = 33.1 bits (74), Expect = 0.43
Identities = 17/50 (34%), Positives = 27/50 (54%)
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQ 149
+A+E+ A E E E EEE EE EE ++EE + E+ E+ ++
Sbjct: 180 EAEEEAEEAEEAEEAEEAEEEAEEAEEEAEEAEEEAEEAEEAEEAEEAEE 229
Score = 33.1 bits (74), Expect = 0.43
Identities = 17/43 (39%), Positives = 25/43 (57%)
Query: 109 AERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
AE E EE+EEE EE EEE+ E +E + ++E ++ E
Sbjct: 197 AEEEAEEAEEEAEEAEEEAEEAEEAEEAEEAEEEAEEAEEEEE 239
Score = 33.1 bits (74), Expect = 0.43
Identities = 19/57 (33%), Positives = 31/57 (54%), Gaps = 1/57 (1%)
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
+A+E+ AE E E E EEE EEEE+ ++EE + E+ + + + E E +
Sbjct: 137 EAEEEEAEEAEEE-EAEEAEEEAEEEEAEEEAEEEAEEAEEAEEEAEEEAEEAEEAE 192
Score = 32.0 bits (71), Expect = 0.95
Identities = 19/56 (33%), Positives = 31/56 (54%), Gaps = 1/56 (1%)
Query: 100 KAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEK 155
+A+E+ A E E EE+EEE EE EE + E +E + ++E ++ E E+
Sbjct: 166 EAEEEAEEAEEAE-EEAEEEAEEAEEAEEAEEAEEEAEEAEEEAEEAEEEAEEAEE 220
Score = 31.6 bits (70), Expect = 1.2
Identities = 18/56 (32%), Positives = 26/56 (46%)
Query: 96 RNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQRE 151
R + +E + AE E E EEE EEEE ++EE +E +E + E
Sbjct: 93 REEAEEEEAEEKEAEEEEAEEAEEEAEEEEAEEAEAEEEEAEEEEAEEEEAEEAEE 148
Score = 31.2 bits (69), Expect = 1.6
Identities = 18/43 (41%), Positives = 24/43 (54%), Gaps = 2/43 (4%)
Query: 100 KAKEDPRRAAER--ECEESEEEEEEEEEESWYDSEESSSESST 140
+A+E+ A E E EE+EE EE EEE + EE + ST
Sbjct: 203 EAEEEAEEAEEEAEEAEEAEEAEEAEEEAEEAEEEEEEAGPST 245
Score = 31.2 bits (69), Expect = 1.6
Identities = 19/60 (31%), Positives = 29/60 (47%), Gaps = 4/60 (6%)
Query: 100 KAKEDPRRAAERECEESE----EEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEK 155
+A+ + A E E EE E EEEE EE E + EE+ E+ + + + E E+
Sbjct: 125 EAEAEEEEAEEEEAEEEEAEEAEEEEAEEAEEEAEEEEAEEEAEEEAEEAEEAEEEAEEE 184
Score = 31.2 bits (69), Expect = 1.6
Identities = 18/54 (33%), Positives = 28/54 (51%), Gaps = 2/54 (3%)
Query: 104 DPRRAAERECEESEEEE--EEEEEESWYDSEESSSESSTIISKEQYDQREVIEK 155
+ A E E EE+EEEE E EEE ++EE + E + + + + E E+
Sbjct: 134 EEEEAEEEEAEEAEEEEAEEAEEEAEEEEAEEEAEEEAEEAEEAEEEAEEEAEE 187
Score = 30.8 bits (68), Expect = 2.1
Identities = 17/55 (30%), Positives = 24/55 (42%)
Query: 102 KEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEKQ 156
+E+ E EE E EE EEE E EE+ E+ E+ + E E +
Sbjct: 135 EEEAEEEEAEEAEEEEAEEAEEEAEEEEAEEEAEEEAEEAEEAEEEAEEEAEEAE 189
Score = 30.8 bits (68), Expect = 2.1
Identities = 18/61 (29%), Positives = 31/61 (50%)
Query: 96 RNGMKAKEDPRRAAERECEESEEEEEEEEEESWYDSEESSSESSTIISKEQYDQREVIEK 155
R G + +E EE+EEEE EE+E ++EE+ E+ ++E + E E+
Sbjct: 76 RRGEEEREGEGGEEGEGREEAEEEEAEEKEAEEEEAEEAEEEAEEEEAEEAEAEEEEAEE 135
Query: 156 Q 156
+
Sbjct: 136 E 136
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.306 0.126 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,398,679
Number of Sequences: 164201
Number of extensions: 1102325
Number of successful extensions: 24044
Number of sequences better than 10.0: 718
Number of HSP's better than 10.0 without gapping: 483
Number of HSP's successfully gapped in prelim test: 258
Number of HSP's that attempted gapping in prelim test: 13775
Number of HSP's gapped (non-prelim): 4199
length of query: 198
length of database: 59,974,054
effective HSP length: 105
effective length of query: 93
effective length of database: 42,732,949
effective search space: 3974164257
effective search space used: 3974164257
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146972.8


