

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146972.11 + phase: 0
(362 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
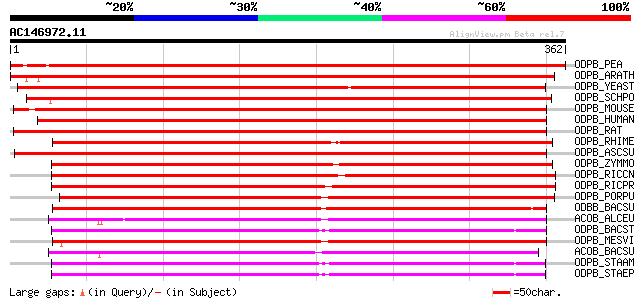
ODPB_PEA (P52904) Pyruvate dehydrogenase E1 component beta subun... 646 0.0
ODPB_ARATH (Q38799) Pyruvate dehydrogenase E1 component beta sub... 608 e-174
ODPB_YEAST (P32473) Pyruvate dehydrogenase E1 component beta sub... 430 e-120
ODPB_SCHPO (Q09171) Pyruvate dehydrogenase E1 component beta sub... 427 e-119
ODPB_MOUSE (Q9D051) Pyruvate dehydrogenase E1 component beta sub... 426 e-119
ODPB_HUMAN (P11177) Pyruvate dehydrogenase E1 component beta sub... 420 e-117
ODPB_RAT (P49432) Pyruvate dehydrogenase E1 component beta subun... 419 e-117
ODPB_RHIME (Q9R9N4) Pyruvate dehydrogenase E1 component, beta su... 417 e-116
ODPB_ASCSU (P26269) Pyruvate dehydrogenase E1 component beta sub... 397 e-110
ODPB_ZYMMO (O66113) Pyruvate dehydrogenase E1 component, beta su... 395 e-110
ODPB_RICCN (Q92IS2) Pyruvate dehydrogenase E1 component, beta su... 393 e-109
ODPB_RICPR (Q9ZDR3) Pyruvate dehydrogenase E1 component, beta su... 386 e-107
ODPB_PORPU (P51266) Pyruvate dehydrogenase E1 component beta sub... 266 8e-71
ODBB_BACSU (P37941) 2-oxoisovalerate dehydrogenase beta subunit ... 258 1e-68
ACOB_ALCEU (P27746) Acetoin:2,6-dichlorophenolindophenol oxidore... 251 3e-66
ODPB_BACST (P21874) Pyruvate dehydrogenase E1 component, beta su... 248 2e-65
ODPB_MESVI (Q9MUR4) Pyruvate dehydrogenase E1 component beta sub... 240 4e-63
ACOB_BACSU (O34591) Acetoin:2,6-dichlorophenolindophenol oxidore... 235 1e-61
ODPB_STAAM (Q9L6H5) Pyruvate dehydrogenase E1 component, beta su... 228 1e-59
ODPB_STAEP (Q8CPN2) Pyruvate dehydrogenase E1 component, beta su... 228 2e-59
>ODPB_PEA (P52904) Pyruvate dehydrogenase E1 component beta subunit,
mitochondrial precursor (EC 1.2.4.1) (PDHE1-B)
Length = 359
Score = 646 bits (1666), Expect = 0.0
Identities = 333/362 (91%), Positives = 341/362 (93%), Gaps = 3/362 (0%)
Query: 1 MLGVIRNKVNLLRPSFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQ 60
MLGVIRNK +RPSFSAFR SS AKQMTVRDALNSALD EMSAD KVFLMGEEVGEYQ
Sbjct: 1 MLGVIRNKT--IRPSFSAFRFFSS-AKQMTVRDALNSALDVEMSADSKVFLMGEEVGEYQ 57
Query: 61 GAYKISKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHII 120
GAYK++KGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHII
Sbjct: 58 GAYKVTKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHII 117
Query: 121 NSAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDAR 180
NSAAKSNYMSAGQI+VPIVFRG NG AAGVGAQHSHCYASWYGSCPGLKVL P+S+EDAR
Sbjct: 118 NSAAKSNYMSAGQISVPIVFRGLNGDAAGVGAQHSHCYASWYGSCPGLKVLVPHSAEDAR 177
Query: 181 GLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSK 240
GLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSF LPIGKAKIEREGKDVTITAFSK
Sbjct: 178 GLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFWLPIGKAKIEREGKDVTITAFSK 237
Query: 241 MVGFALKAAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAE 300
MVGFALKAAE LEKEGISAEVINLRSIRPLDR TINASVRKTNRLVTVEEGFPQHGVGAE
Sbjct: 238 MVGFALKAAEILEKEGISAEVINLRSIRPLDRPTINASVRKTNRLVTVEEGFPQHGVGAE 297
Query: 301 ICASVIEESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKRACHRSVPMAA 360
IC SVIEESFGYLDA VERI GADVPMPYA NLERL VP +EDIVRAAKRACHRSVP+AA
Sbjct: 298 ICTSVIEESFGYLDATVERIGGADVPMPYAGNLERLVVPHVEDIVRAAKRACHRSVPLAA 357
Query: 361 TA 362
A
Sbjct: 358 AA 359
>ODPB_ARATH (Q38799) Pyruvate dehydrogenase E1 component beta
subunit, mitochondrial precursor (EC 1.2.4.1) (PDHE1-B)
Length = 363
Score = 608 bits (1569), Expect = e-174
Identities = 311/362 (85%), Positives = 332/362 (90%), Gaps = 7/362 (1%)
Query: 1 MLGVIRNKV-----NLLRPSFS--AFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMG 53
MLG++R + L R F+ + R ++ AK+MTVRDALNSA+DEEMSADPKVF+MG
Sbjct: 1 MLGILRQRAIDGASTLRRTRFALVSARSYAAGAKEMTVRDALNSAIDEEMSADPKVFVMG 60
Query: 54 EEVGEYQGAYKISKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSM 113
EEVG+YQGAYKI+KGLLEKYGPERV DTPITEAGFTGIGVGAAY GLKPVVEFMTFNFSM
Sbjct: 61 EEVGQYQGAYKITKGLLEKYGPERVYDTPITEAGFTGIGVGAAYAGLKPVVEFMTFNFSM 120
Query: 114 QAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAP 173
QAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHS CYA+WY S PGLKVLAP
Sbjct: 121 QAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHSQCYAAWYASVPGLKVLAP 180
Query: 174 YSSEDARGLLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDV 233
YS+EDARGLLKAAIRDPDPVVFLENELLYGESFP+S E LDSSFCLPIGKAKIEREGKDV
Sbjct: 181 YSAEDARGLLKAAIRDPDPVVFLENELLYGESFPISEEALDSSFCLPIGKAKIEREGKDV 240
Query: 234 TITAFSKMVGFALKAAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFP 293
TI FSKMVGFALKAAE L +EGISAEVINLRSIRPLDRATINASVRKT+RLVTVEEGFP
Sbjct: 241 TIVTFSKMVGFALKAAEKLAEEGISAEVINLRSIRPLDRATINASVRKTSRLVTVEEGFP 300
Query: 294 QHGVGAEICASVIEESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKRACH 353
QHGV AEICASV+EESF YLDAPVERIAGADVPMPYAANLERLA+PQIEDIVRA+KRAC+
Sbjct: 301 QHGVCAEICASVVEESFSYLDAPVERIAGADVPMPYAANLERLALPQIEDIVRASKRACY 360
Query: 354 RS 355
RS
Sbjct: 361 RS 362
>ODPB_YEAST (P32473) Pyruvate dehydrogenase E1 component beta
subunit, mitochondrial precursor (EC 1.2.4.1) (PDHE1-B)
Length = 366
Score = 430 bits (1106), Expect = e-120
Identities = 222/346 (64%), Positives = 267/346 (77%), Gaps = 3/346 (0%)
Query: 6 RNKVNLLRPSFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKI 65
R + +RPS +A SS K MTVR+ALNSA+ EE+ D VFL+GEEV +Y GAYK+
Sbjct: 16 RAPTSFVRPSAAAAALRFSSTKTMTVREALNSAMAEELDRDDDVFLIGEEVAQYNGAYKV 75
Query: 66 SKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAK 125
SKGLL+++G RV+DTPITE GFTG+ VGAA GLKP+VEFM+FNFSMQAIDH++NSAAK
Sbjct: 76 SKGLLDRFGERRVVDTPITEYGFTGLAVGAALKGLKPIVEFMSFNFSMQAIDHVVNSAAK 135
Query: 126 SNYMSAGQINVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKA 185
++YMS G +VFRGPNGAA GVGAQHS ++ WYGS PGLKVL PYS+EDARGLLKA
Sbjct: 136 THYMSGGTQKCQMVFRGPNGAAVGVGAQHSQDFSPWYGSIPGLKVLVPYSAEDARGLLKA 195
Query: 186 AIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFA 245
AIRDP+PVVFLENELLYGESF +S E L F LP KAKIEREG D++I +++ V F+
Sbjct: 196 AIRDPNPVVFLENELLYGESFEISEEALSPEFTLPY-KAKIEREGTDISIVTYTRNVQFS 254
Query: 246 LKAAETLEKE-GISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICAS 304
L+AAE L+K+ G+SAEVINLRSIRPLD I +V+KTN L+TVE FP GVGAEI A
Sbjct: 255 LEAAEILQKKYGVSAEVINLRSIRPLDTEAIIKTVKKTNHLITVESTFPSFGVGAEIVAQ 314
Query: 305 VIE-ESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAK 349
V+E E+F YLDAP++R+ GADVP PYA LE A P IV+A K
Sbjct: 315 VMESEAFDYLDAPIQRVTGADVPTPYAKELEDFAFPDTPTIVKAVK 360
>ODPB_SCHPO (Q09171) Pyruvate dehydrogenase E1 component beta
subunit, mitochondrial precursor (EC 1.2.4.1) (PDHE1-B)
Length = 366
Score = 427 bits (1098), Expect = e-119
Identities = 218/346 (63%), Positives = 273/346 (78%), Gaps = 4/346 (1%)
Query: 12 LRPSFSAFRHLSSS--AKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGL 69
L S A R+ SSS K+MTVRDALNSA++EEM D +VFL+GEEV +Y GAYKIS+GL
Sbjct: 19 LLSSTIAKRYSSSSNGVKEMTVRDALNSAMEEEMKRDDRVFLIGEEVAQYNGAYKISRGL 78
Query: 70 LEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYM 129
L+K+GP+RV+DTPITE GFTG+ GAA+ GL+P+ EFMTFNFSMQAIDHI+NSAA++ YM
Sbjct: 79 LDKFGPKRVIDTPITEMGFTGLATGAAFAGLRPICEFMTFNFSMQAIDHIVNSAARTLYM 138
Query: 130 SAGQINVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRD 189
S G PIVFRGPNG AA V AQHS +A WYGS PGLKV++PYS+EDARGLLKAAIRD
Sbjct: 139 SGGIQACPIVFRGPNGPAAAVAAQHSQHFAPWYGSIPGLKVVSPYSAEDARGLLKAAIRD 198
Query: 190 PDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAA 249
P+PVV LENE+LYG++FP+S E L F LP G AK+ER GKD+TI S V AL+AA
Sbjct: 199 PNPVVVLENEILYGKTFPISKEALSEDFVLPFGLAKVERPGKDITIVGESISVVTALEAA 258
Query: 250 ETLEKE-GISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIE- 307
+ L+ + G+ AEVINLRSIRPLD TI ASV+KTNR+VTV++ + QHG+G+EI A ++E
Sbjct: 259 DKLKADYGVEAEVINLRSIRPLDINTIAASVKKTNRIVTVDQAYSQHGIGSEIAAQIMES 318
Query: 308 ESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKRACH 353
++F YLDAPVER++ ADVPMPY+ +E +VP + +V AAK+ +
Sbjct: 319 DAFDYLDAPVERVSMADVPMPYSHPVEAASVPNADVVVAAAKKCLY 364
>ODPB_MOUSE (Q9D051) Pyruvate dehydrogenase E1 component beta
subunit, mitochondrial precursor (EC 1.2.4.1) (PDHE1-B)
Length = 359
Score = 426 bits (1095), Expect = e-119
Identities = 212/349 (60%), Positives = 267/349 (75%), Gaps = 4/349 (1%)
Query: 3 GVIRNKVNLLRPSFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGA 62
G +R LL+ F + +A Q+TVR+A+N +DEE+ D KVFL+GEEV +Y GA
Sbjct: 10 GPLRQASGLLK---RRFHRSAPAAVQLTVREAINQGMDEELERDEKVFLLGEEVAQYDGA 66
Query: 63 YKISKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINS 122
YK+S+GL +KYG +R++DTPI+E GF GI VGAA GL+P+ EFMTFNFSMQAID +INS
Sbjct: 67 YKVSRGLWKKYGDKRIIDTPISEMGFAGIAVGAAMAGLRPICEFMTFNFSMQAIDQVINS 126
Query: 123 AAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGL 182
AAK+ YMSAG VPIVFRGPNGA+AGV AQHS C+A+WYG CPGLKV++P++SEDA+GL
Sbjct: 127 AAKTYYMSAGLQPVPIVFRGPNGASAGVAAQHSQCFAAWYGHCPGLKVVSPWNSEDAKGL 186
Query: 183 LKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMV 242
+K+AIRD +PVV LENEL+YG +F + AE F +PIGKAKIER+G +T+ A S+ V
Sbjct: 187 IKSAIRDNNPVVMLENELMYGVAFELPAEAQSKDFLIPIGKAKIERQGTHITVVAHSRPV 246
Query: 243 GFALKAAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEIC 302
G L+AA L KEGI EVINLR+IRP+D I ASV KTN LVTVE G+PQ GVGAEIC
Sbjct: 247 GHCLEAAAVLSKEGIECEVINLRTIRPMDIEAIEASVMKTNHLVTVEGGWPQFGVGAEIC 306
Query: 303 ASVIE-ESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKR 350
A ++E +F +LDAP R+ GADVPMPYA LE +VPQ++DI+ A K+
Sbjct: 307 ARIMEGPAFNFLDAPAVRVTGADVPMPYAKVLEDNSVPQVKDIIFAVKK 355
>ODPB_HUMAN (P11177) Pyruvate dehydrogenase E1 component beta
subunit, mitochondrial precursor (EC 1.2.4.1) (PDHE1-B)
Length = 359
Score = 420 bits (1080), Expect = e-117
Identities = 204/333 (61%), Positives = 258/333 (77%), Gaps = 1/333 (0%)
Query: 19 FRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERV 78
F + +A Q+TVRDA+N +DEE+ D KVFL+GEEV +Y GAYK+S+GL +KYG +R+
Sbjct: 23 FHWTAPAAVQVTVRDAINQGMDEELERDEKVFLLGEEVAQYDGAYKVSRGLWKKYGDKRI 82
Query: 79 LDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPI 138
+DTPI+E GF GI VGAA GL+P+ EFMTFNFSMQAID +INSAAK+ YMS G VPI
Sbjct: 83 IDTPISEMGFAGIAVGAAMAGLRPICEFMTFNFSMQAIDQVINSAAKTYYMSGGLQPVPI 142
Query: 139 VFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLEN 198
VFRGPNGA+AGV AQHS C+A+WYG CPGLKV++P++SEDA+GL+K+AIRD +PVV LEN
Sbjct: 143 VFRGPNGASAGVAAQHSQCFAAWYGHCPGLKVVSPWNSEDAKGLIKSAIRDNNPVVVLEN 202
Query: 199 ELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGIS 258
EL+YG F E F +PIGKAKIER+G +T+ + S+ VG L+AA L KEG+
Sbjct: 203 ELMYGVPFEFPPEAQSKDFLIPIGKAKIERQGTHITVVSHSRPVGHCLEAAAVLSKEGVE 262
Query: 259 AEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIE-ESFGYLDAPV 317
EVIN+R+IRP+D TI ASV KTN LVTVE G+PQ GVGAEICA ++E +F +LDAP
Sbjct: 263 CEVINMRTIRPMDMETIEASVMKTNHLVTVEGGWPQFGVGAEICARIMEGPAFNFLDAPA 322
Query: 318 ERIAGADVPMPYAANLERLAVPQIEDIVRAAKR 350
R+ GADVPMPYA LE ++PQ++DI+ A K+
Sbjct: 323 VRVTGADVPMPYAKILEDNSIPQVKDIIFAIKK 355
>ODPB_RAT (P49432) Pyruvate dehydrogenase E1 component beta subunit,
mitochondrial precursor (EC 1.2.4.1) (PDHE1-B)
Length = 359
Score = 419 bits (1077), Expect = e-117
Identities = 207/350 (59%), Positives = 264/350 (75%), Gaps = 2/350 (0%)
Query: 3 GVIRNKVNLLRPSFSAFRHLSS-SAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQG 61
G++R L H S+ +A Q+TVR+A+N +DEE+ D KVFL+GEEV +Y G
Sbjct: 6 GLVRGLCGRLSGLLKRLFHCSAPAAVQLTVREAINQGMDEELERDEKVFLLGEEVAQYDG 65
Query: 62 AYKISKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIIN 121
AYK+S+GL +KYG +R++DTPI+E GF GI VGAA GL+P+ EFMTFNFSMQAID +IN
Sbjct: 66 AYKVSRGLWKKYGDKRIIDTPISEMGFAGIAVGAAMAGLRPICEFMTFNFSMQAIDQVIN 125
Query: 122 SAAKSNYMSAGQINVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARG 181
SAAK+ YMSAG VPIVFRGPNGA+AGV AQHS C+A+WYG CPGLKV++P++SEDA+G
Sbjct: 126 SAAKTYYMSAGLQPVPIVFRGPNGASAGVAAQHSQCFAAWYGHCPGLKVVSPWNSEDAKG 185
Query: 182 LLKAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKM 241
L+K+AIRD +PVV LENEL+YG +F + E F +PIGKAKIER+G + + +S+
Sbjct: 186 LIKSAIRDDNPVVMLENELMYGVAFELPTEAQSKDFLIPIGKAKIERQGTHINVVCYSRP 245
Query: 242 VGFALKAAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEI 301
VG L+AA L K GI EVINLR+IRP+D I ASV KTN LVTVE G+PQ GVGAEI
Sbjct: 246 VGHCLEAAAVLSKGGIECEVINLRTIRPMDIEAIEASVMKTNHLVTVEGGWPQFGVGAEI 305
Query: 302 CASVIE-ESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKR 350
CA ++E +F +LDAP R+ GADVPMPYA LE ++PQ++DI+ A K+
Sbjct: 306 CARIMEGPAFNFLDAPAVRVTGADVPMPYAKILEDNSIPQVKDIIFAIKK 355
>ODPB_RHIME (Q9R9N4) Pyruvate dehydrogenase E1 component, beta
subunit (EC 1.2.4.1)
Length = 460
Score = 417 bits (1071), Expect = e-116
Identities = 205/326 (62%), Positives = 258/326 (78%), Gaps = 3/326 (0%)
Query: 29 MTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTPITEAGF 88
MTVR+AL A+ EEM A+ VF+MGEEV EYQGAYK+++GLL+++G RV+DTPITE GF
Sbjct: 138 MTVREALRDAMAEEMRANEDVFVMGEEVAEYQGAYKVTQGLLQEFGARRVVDTPITEHGF 197
Query: 89 TGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAAA 148
G+GVGAA GL+P+VEFMTFNF+MQAID IINSAAK+ YMS GQ+ PIVFRGP+GAAA
Sbjct: 198 AGVGVGAAMTGLRPIVEFMTFNFAMQAIDQIINSAAKTLYMSGGQMGAPIVFRGPSGAAA 257
Query: 149 GVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLYGESFPV 208
V AQHS CYA+WY PGLKV+ PY++ DA+GLLKAAIRDP+PV+FLENE+LYG+SF V
Sbjct: 258 RVAAQHSQCYAAWYSHIPGLKVVMPYTAADAKGLLKAAIRDPNPVIFLENEILYGQSFEV 317
Query: 209 SAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVINLRSIR 268
LD F LPIGKA+I R GKD T+ +F + +A+KAA LE +GI E+I+LR+IR
Sbjct: 318 PK--LD-DFVLPIGKARIHRTGKDATLVSFGIGMTYAIKAAAELEAQGIDVEIIDLRTIR 374
Query: 269 PLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAGADVPMP 328
P+D T+ SV+KT RLVTVEEG+PQ VG EI V++++F YLDAP+ IAG DVPMP
Sbjct: 375 PMDLPTVIESVKKTGRLVTVEEGYPQSSVGTEIATRVMQQAFDYLDAPILTIAGKDVPMP 434
Query: 329 YAANLERLAVPQIEDIVRAAKRACHR 354
YAANLE+LA+P + ++V A K C++
Sbjct: 435 YAANLEKLALPNVAEVVDAVKAVCYK 460
>ODPB_ASCSU (P26269) Pyruvate dehydrogenase E1 component beta
subunit, mitochondrial precursor (EC 1.2.4.1) (PDHE1-B)
Length = 361
Score = 397 bits (1019), Expect = e-110
Identities = 202/348 (58%), Positives = 249/348 (71%), Gaps = 1/348 (0%)
Query: 4 VIRNKVNLLRPSFSAFRHLSSSAKQMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAY 63
++RN + + R L+S +TVRDALN+ALDEE+ D +VFL+GEEV +Y GAY
Sbjct: 9 LLRNGLTSACALEQSVRRLASGTLNVTVRDALNAALDEEIKRDDRVFLIGEEVAQYDGAY 68
Query: 64 KISKGLLEKYGPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSA 123
KISKGL +KYG R+ DTPITE G+ VGAA GL+P+ EFM+ NFSMQ IDHIINSA
Sbjct: 69 KISKGLWKKYGDGRIWDTPITEMAIAGLSVGAAMNGLRPICEFMSMNFSMQGIDHIINSA 128
Query: 124 AKSNYMSAGQINVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLL 183
AK++YMSAG+ +VPIVFRG NGAA GV QHS + +W+ CPG+KV+ PY EDARGLL
Sbjct: 129 AKAHYMSAGRFHVPIVFRGANGAAVGVAQQHSQDFTAWFMHCPGVKVVVPYDCEDARGLL 188
Query: 184 KAAIRDPDPVVFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVG 243
KAA+RD +PV+ LENE+LYG FPVS E F LP G+AKI+R GKD+TI + S V
Sbjct: 189 KAAVRDDNPVICLENEILYGMKFPVSPEAQSPDFVLPFGQAKIQRPGKDITIVSLSIGVD 248
Query: 244 FALKAAETLEKEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICA 303
+L AA+ L K GI EVINLR +RPLD T+ SV KT LVTVE G+P GVGAEI A
Sbjct: 249 VSLHAADELAKSGIDCEVINLRCVRPLDFQTVKDSVIKTKHLVTVESGWPNCGVGAEISA 308
Query: 304 SVIE-ESFGYLDAPVERIAGADVPMPYAANLERLAVPQIEDIVRAAKR 350
V E ++FGYLD P+ R+ G DVPMPYA LE A+PQ D+V+ K+
Sbjct: 309 RVTESDAFGYLDGPILRVTGVDVPMPYAQPLETAALPQPADVVKMVKK 356
>ODPB_ZYMMO (O66113) Pyruvate dehydrogenase E1 component, beta
subunit (EC 1.2.4.1)
Length = 462
Score = 395 bits (1015), Expect = e-110
Identities = 201/327 (61%), Positives = 248/327 (75%), Gaps = 3/327 (0%)
Query: 28 QMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTPITEAG 87
Q T+R+AL A+ EEM D +VF+MGEEV EYQGAYK+++GLL+++G RV+DTPI+E G
Sbjct: 138 QQTLREALRDAMAEEMRRDDRVFVMGEEVAEYQGAYKVTQGLLQEFGARRVVDTPISEYG 197
Query: 88 FTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAA 147
F+GIGVGAA GL+PV+EFMT NFSMQAIDHIINSAAK++YMS GQ+ PIVFRGPNGAA
Sbjct: 198 FSGIGVGAAMEGLRPVIEFMTMNFSMQAIDHIINSAAKTHYMSGGQVRCPIVFRGPNGAA 257
Query: 148 AGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLYGESFP 207
VGAQH+ + WY + PGL VLAPY + DA+GLLKAAIR DPVVFLE ELLYG++F
Sbjct: 258 PRVGAQHTQNFGPWYAAVPGLVVLAPYDAIDAKGLLKAAIRSDDPVVFLECELLYGKTFD 317
Query: 208 VSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVINLRSI 267
V F LPIGKA+I REGKDVTI ++S V FAL AAE L KEGI AEVI+LR++
Sbjct: 318 VPKM---DDFVLPIGKARIIREGKDVTIVSYSIGVSFALTAAEALAKEGIDAEVIDLRTL 374
Query: 268 RPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAGADVPM 327
RPLD+ TI S+ KTNR+VTVE+G+P + +EI A +EE F LDAPV R+ AD P
Sbjct: 375 RPLDKETILQSLAKTNRIVTVEDGWPVCSISSEIAAIAMEEGFDNLDAPVLRVTNADTPT 434
Query: 328 PYAANLERLAVPQIEDIVRAAKRACHR 354
PYA NLE+ + E I+ A ++ C+R
Sbjct: 435 PYAENLEKKGLVNPEAIIEAVRKVCYR 461
>ODPB_RICCN (Q92IS2) Pyruvate dehydrogenase E1 component, beta
subunit (EC 1.2.4.1)
Length = 326
Score = 393 bits (1010), Expect = e-109
Identities = 195/329 (59%), Positives = 250/329 (75%), Gaps = 4/329 (1%)
Query: 28 QMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTPITEAG 87
Q+TVR+AL A+ EEM D KVF++GEEV EYQGAYK+++GLLE++GP+RV+DTPITE G
Sbjct: 2 QITVREALRDAMQEEMIRDDKVFVIGEEVAEYQGAYKVTQGLLEQFGPKRVIDTPITEYG 61
Query: 88 FTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAA 147
F G+ VGAA+ GL+P+VEFMTFNF+MQA DHI+NSAAK++YMS GQ+ PIVFRGPNGAA
Sbjct: 62 FAGLAVGAAFAGLRPIVEFMTFNFAMQAFDHIVNSAAKTHYMSGGQVKCPIVFRGPNGAA 121
Query: 148 AGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLYGESFP 207
+ V AQHS Y + Y PGLKV+APYS+ED +GL+ AIRD +PVVFLENE+LYG SF
Sbjct: 122 SRVAAQHSQNYTACYSHIPGLKVVAPYSAEDHKGLMLTAIRDDNPVVFLENEILYGHSFD 181
Query: 208 VSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVINLRSI 267
V + +P G+AKI REG VTI FS V AL AA ++ + I EVI+LR+I
Sbjct: 182 VPKTIEP----IPFGQAKILREGSSVTIVTFSIQVKLALDAANFVQNDNIDCEVIDLRTI 237
Query: 268 RPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAGADVPM 327
+PLD TI SV+KTNRLV VEEG+ GVGA I + V++E+F YLDAP+E ++G D+P+
Sbjct: 238 KPLDTETIIESVKKTNRLVVVEEGWFFAGVGASIASIVMKEAFDYLDAPIEIVSGKDLPL 297
Query: 328 PYAANLERLAVPQIEDIVRAAKRACHRSV 356
PYA NLE LA+P D++ A K+ C+ S+
Sbjct: 298 PYAVNLETLALPSESDVIEAVKKVCYYSI 326
>ODPB_RICPR (Q9ZDR3) Pyruvate dehydrogenase E1 component, beta
subunit (EC 1.2.4.1)
Length = 326
Score = 386 bits (991), Expect = e-107
Identities = 191/329 (58%), Positives = 249/329 (75%), Gaps = 4/329 (1%)
Query: 28 QMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTPITEAG 87
Q+TVR+AL A+ EEM D KVF++GEEV EYQGAYK+++GLLE++G +RV+DTPITE G
Sbjct: 2 QITVREALRDAMQEEMLRDEKVFVIGEEVAEYQGAYKVTQGLLEQFGSKRVIDTPITEYG 61
Query: 88 FTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAA 147
F G+ VGAA+ GL+P+VEFMTFNF+MQA DHI+NSAAK++YMS GQ+ PIVFRGPNGAA
Sbjct: 62 FAGLAVGAAFAGLRPIVEFMTFNFAMQAFDHIVNSAAKTHYMSGGQVKCPIVFRGPNGAA 121
Query: 148 AGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLYGESFP 207
+ V AQHS Y + Y PGLKV+APYS+ED +GL+ AIRD +PV+FLENE+LYG SF
Sbjct: 122 SRVAAQHSQNYTACYSHIPGLKVVAPYSAEDHKGLMLTAIRDDNPVIFLENEILYGHSF- 180
Query: 208 VSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVINLRSI 267
+V D +P KAKI +EG +VTI FS V AL L+ + I E+I+LR+I
Sbjct: 181 ---DVPDIIEPIPFSKAKILKEGSNVTIVTFSIQVKLALDVVNILQNDNIDCELIDLRTI 237
Query: 268 RPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAGADVPM 327
+PLD +I SV+KTNRLV VEEG+ GVGA I + V++E+F YLDAP+E ++G DVP+
Sbjct: 238 KPLDTDSIIESVKKTNRLVIVEEGWFFAGVGASIASIVMKEAFDYLDAPIEIVSGKDVPL 297
Query: 328 PYAANLERLAVPQIEDIVRAAKRACHRSV 356
PYA NLE+LA+P D++ A K+ C+ S+
Sbjct: 298 PYAVNLEKLAMPSANDLIEAVKKVCYYSI 326
>ODPB_PORPU (P51266) Pyruvate dehydrogenase E1 component beta
subunit (EC 1.2.4.1)
Length = 331
Score = 266 bits (679), Expect = 8e-71
Identities = 139/323 (43%), Positives = 204/323 (63%), Gaps = 4/323 (1%)
Query: 33 DALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTPITEAGFTGIG 92
DAL +A DEEM D V ++GE+VG Y G+YK++K L KYG RVLDTPI E FTG+
Sbjct: 8 DALRAATDEEMEKDLTVCVIGEDVGHYGGSYKVTKDLHSKYGDLRVLDTPIAENSFTGMA 67
Query: 93 VGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAAAGVGA 152
+GAA GL+P+VE M +F + A + I N+A Y S G +P+V RGP G +GA
Sbjct: 68 IGAAITGLRPIVEGMNMSFLLLAFNQISNNAGMLRYTSGGNFTLPLVIRGPGGVGRQLGA 127
Query: 153 QHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLYGESFPVSAEV 212
+HS +++ + PGLK++A + +A+GLLK+AIRD +PVVF E+ LLY + E+
Sbjct: 128 EHSQRLEAYFQAIPGLKIVACSTPYNAKGLLKSAIRDNNPVVFFEHVLLYN----LQEEI 183
Query: 213 LDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVINLRSIRPLDR 272
+ + +P+ KA++ R+GKD+TI +S+M +A L +G EV++L S++PLD
Sbjct: 184 PEDEYLIPLDKAEVVRKGKDITILTYSRMRHHVTEALPLLLNDGYDPEVLDLISLKPLDI 243
Query: 273 ATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAGADVPMPYAAN 332
+I+ SV+KT+R++ VEE G+GAE+ A + E F LDAPV R++ D+P PY +
Sbjct: 244 DSISVSVKKTHRVLIVEECMKTAGIGAELIAQINEHLFDELDAPVVRLSSQDIPTPYNGS 303
Query: 333 LERLAVPQIEDIVRAAKRACHRS 355
LE+ V Q I+ A K + S
Sbjct: 304 LEQATVIQPHQIIDAVKNIVNSS 326
>ODBB_BACSU (P37941) 2-oxoisovalerate dehydrogenase beta subunit (EC
1.2.4.4) (Branched-chain alpha-keto acid dehydrogenase
E1 component beta chain) (BCKDH E1-beta)
Length = 327
Score = 258 bits (660), Expect = 1e-68
Identities = 136/323 (42%), Positives = 201/323 (62%), Gaps = 5/323 (1%)
Query: 29 MTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTPITEAGF 88
M+ DA+N A+ EEM D +VF++GE+VG G +K + GL E++G ERV+DTP+ E+
Sbjct: 4 MSYIDAINLAMKEEMERDSRVFVLGEDVGRKGGVFKATAGLYEQFGEERVMDTPLAESAI 63
Query: 89 TGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAAA 148
G+G+GAA YG++P+ E +F M A++ II+ AAK Y S + PIV R P G
Sbjct: 64 AGVGIGAAMYGMRPIAEMQFADFIMPAVNQIISEAAKIRYRSNNDWSCPIVVRAPYGGGV 123
Query: 149 GVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLYGESFPV 208
HS + + + PGLK++ P + DA+GLLKAA+RD DPV+F E++ Y +
Sbjct: 124 HGALYHSQSVEAIFANQPGLKIVMPSTPYDAKGLLKAAVRDEDPVLFFEHKRAYR---LI 180
Query: 209 SAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVINLRSIR 268
EV + LPIGKA ++REG D+T+ + V FAL+AAE LEK+GISA V++LR++
Sbjct: 181 KGEVPADDYVLPIGKADVKREGDDITVITYGLCVHFALQAAERLEKDGISAHVVDLRTVY 240
Query: 269 PLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAGADVP-M 327
PLD+ I + KT +++ V E + + +E+ A + E LDAP++R+AG D+P M
Sbjct: 241 PLDKEAIIEAASKTGKVLLVTEDTKEGSIMSEVAAIISEHCLFDLDAPIKRLAGPDIPAM 300
Query: 328 PYAANLERLAVPQIEDIVRAAKR 350
PYA +E+ + D V AA R
Sbjct: 301 PYAPTMEKYFMVN-PDKVEAAMR 322
>ACOB_ALCEU (P27746) Acetoin:2,6-dichlorophenolindophenol
oxidoreductase beta subunit (EC 1.1.1.-) (Acetoin:DCPIP
oxidoreductase-beta) (AO:DCPIP OR) (TPP-dependent
acetoin dehydrogenase E1 beta-subunit)
Length = 337
Score = 251 bits (640), Expect = 3e-66
Identities = 133/335 (39%), Positives = 202/335 (59%), Gaps = 15/335 (4%)
Query: 26 AKQMTVRDALNSALDEEMSADPKVFLMGEEV-------GE---YQGAYKISKGLLEKYGP 75
A++++++ A+N A+D+EM+ DP V ++GE++ GE + G ++KGL K+G
Sbjct: 1 ARKLSIKLAINEAIDQEMTRDPSVIMLGEDIVGGAGADGEKDAWGGVLGVTKGLYAKHG- 59
Query: 76 ERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQIN 135
+R+LDTP++E+ + G +GAA G++P+ E M +F D I N AAK YM G+
Sbjct: 60 DRLLDTPLSESAYVGAAIGAAACGMRPIAELMFIDFMGVCFDQIFNQAAKFRYMFGGKAE 119
Query: 136 VPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVF 195
P+V R GA AQHS + PGLKV+ P + D +GLL AIRD DPV+F
Sbjct: 120 TPVVIRAMVGAGFRAAAQHSQMLTPLFTHIPGLKVVCPSTPYDTKGLLIQAIRDNDPVIF 179
Query: 196 LENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKE 255
E++ LYG + EV + ++ +P G+A I R+GKDV+I + MV AL+AA TL KE
Sbjct: 180 CEHKNLYG----LEGEVPEGAYAIPFGEANIVRDGKDVSIVTYGLMVHRALEAAATLAKE 235
Query: 256 GISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDA 315
GI AE+++LR++ PLD T+ SV T RLV V+E P+ + +I A V +++FG L A
Sbjct: 236 GIEAEIVDLRTLSPLDMDTVLESVENTGRLVVVDEASPRCNIATDISAQVAQQAFGALKA 295
Query: 316 PVERIAGADVPMPYAANLERLAVPQIEDIVRAAKR 350
+E + P+P++ LE L +P I AA++
Sbjct: 296 GIEMVCPPHTPVPFSPTLEDLYIPSAAQIAAAARK 330
>ODPB_BACST (P21874) Pyruvate dehydrogenase E1 component, beta
subunit (EC 1.2.4.1)
Length = 324
Score = 248 bits (633), Expect = 2e-65
Identities = 136/323 (42%), Positives = 193/323 (59%), Gaps = 4/323 (1%)
Query: 28 QMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTPITEAG 87
QMT+ A+ AL E+ DP V + GE+VG G ++ ++GL ++G +RV DTP+ E+G
Sbjct: 2 QMTMVQAITDALRIELKNDPNVLIFGEDVGVNGGVFRATEGLQAEFGEDRVFDTPLAESG 61
Query: 88 FTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAA 147
G+ +G A G +PV E F F + +D I A+ Y + G+ ++PI R P G
Sbjct: 62 IGGLAIGLALQGFRPVPEIQFFGFVYEVMDSICGQMARIRYRTGGRYHMPITIRSPFGGG 121
Query: 148 AGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLYGESFP 207
HS PGLKV+ P + DA+GLL +AIRD DPV+FLE+ LY SF
Sbjct: 122 VHTPELHSDSLEGLVAQQPGLKVVIPSTPYDAKGLLISAIRDNDPVIFLEHLKLY-RSF- 179
Query: 208 VSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVINLRSI 267
EV + + +PIGKA I+REGKD+TI A+ MV +LKAA LEKEGISAEV++LR++
Sbjct: 180 -RQEVPEGEYTIPIGKADIKREGKDITIIAYGAMVHESLKAAAELEKEGISAEVVDLRTV 238
Query: 268 RPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAGADVPM 327
+PLD TI SV KT R + V+E Q G+ A + A + E + L+APV R+A D
Sbjct: 239 QPLDIETIIGSVEKTGRAIVVQEAQRQAGIAANVVAEINERAILSLEAPVLRVAAPDTVY 298
Query: 328 PYAANLERLAVPQIEDIVRAAKR 350
P+ A E + +P +D++ AK+
Sbjct: 299 PF-AQAESVWLPNFKDVIETAKK 320
>ODPB_MESVI (Q9MUR4) Pyruvate dehydrogenase E1 component beta
subunit (EC 1.2.4.1)
Length = 327
Score = 240 bits (613), Expect = 4e-63
Identities = 128/325 (39%), Positives = 200/325 (61%), Gaps = 7/325 (2%)
Query: 29 MTVR---DALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTPITE 85
MTVR +ALN A+DEEM+ + KV L+GE++G Y G+YK+++ L KYG RV+DTPI E
Sbjct: 1 MTVRFLFEALNMAIDEEMARNDKVALLGEDIGHYGGSYKVTQNLYAKYGEHRVIDTPIAE 60
Query: 86 AGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRGPNG 145
F G +GAA GL VVE M F + A I N+ + S G ++PIV RGP G
Sbjct: 61 NSFVGAAIGAAMTGLVTVVEGMNMGFILLAFSQISNNMGMLSATSGGHYHIPIVLRGPGG 120
Query: 146 AAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLYGES 205
+GA+HS ++ S PGL+++A + +A+GLLK+AIR +P+ FLE+ LLY
Sbjct: 121 VGKQLGAEHSQRLECYFQSVPGLQIVACSTPYNAKGLLKSAIRSKNPIFFLEHVLLYN-- 178
Query: 206 FPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVINLR 265
+ AEV D+ + LP+ KA+I R+G D+TI +S+M ++A + L ++G E+I+L
Sbjct: 179 --LKAEVPDNDYVLPLEKAEIVRQGNDITILTYSRMRYNVIQAVKVLVEKGYDPEIIDLI 236
Query: 266 SIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAGADV 325
S++P D TI S++KT++++ VEE G+ + + ++E F LD ++ +V
Sbjct: 237 SLKPFDIETIGKSIQKTHKVLIVEESMMTGGISNVLQSLILENFFDDLDNRPMCLSSPNV 296
Query: 326 PMPYAANLERLAVPQIEDIVRAAKR 350
P PY+ LE +++ Q DI+ + ++
Sbjct: 297 PTPYSGPLEEVSIVQTADIIESVEQ 321
>ACOB_BACSU (O34591) Acetoin:2,6-dichlorophenolindophenol
oxidoreductase beta subunit (EC 1.1.1.-) (Acetoin:DCPIP
oxidoreductase-beta) (AO:DCPIP OR) (TPP-dependent
acetoin dehydrogenase E1 beta-subunit)
Length = 341
Score = 235 bits (600), Expect = 1e-61
Identities = 132/332 (39%), Positives = 190/332 (56%), Gaps = 16/332 (4%)
Query: 26 AKQMTVRDALNSALDEEMSADPKVFLMGEEVG------------EYQGAYKISKGLLEKY 73
A+ +++ DA+N A+ M D V L+GE+V + G ++KGL++++
Sbjct: 1 ARVISMSDAINEAMKLAMRKDENVLLIGEDVAGGAAVDHLQDDEAWGGVLGVTKGLVQEF 60
Query: 74 GPERVLDTPITEAGFTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQ 133
G RVLDTPI+EAG+ G + AA GL+P+ E M +F D +IN AK YM G+
Sbjct: 61 GRTRVLDTPISEAGYMGAAMAAASTGLRPIAELMFNDFIGTCFDQVINQGAKFRYMFGGK 120
Query: 134 INVPIVFRGPNGAAAGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPV 193
VPI R GA AQHS + S PGLK + P + DA+GLL AAI D DPV
Sbjct: 121 AQVPITVRTTYGAGFRAAAQHSQSLYGLFTSIPGLKTVVPSNPYDAKGLLLAAIEDNDPV 180
Query: 194 VFLENELLYGESFPVSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLE 253
F E++ S+ + EV + + +P+GKA I+REG DVT+ A K V AL+AA L
Sbjct: 181 FFFEDK----TSYNMKGEVPEDYYTIPLGKADIKREGNDVTLFAVGKQVNTALEAAAQLS 236
Query: 254 KEGISAEVINLRSIRPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYL 313
+ GI AEV++ RS+ PLD I S+ KTNRL+ ++E P+ + +I A V ++ F L
Sbjct: 237 ERGIEAEVLDPRSLSPLDEDAIFTSLEKTNRLIIIDEANPRCSIATDIAALVADKGFDLL 296
Query: 314 DAPVERIAGADVPMPYAANLERLAVPQIEDIV 345
DAP++RI P+P++ LE +P + IV
Sbjct: 297 DAPIKRITAPHTPVPFSPVLEDQYLPTPDKIV 328
>ODPB_STAAM (Q9L6H5) Pyruvate dehydrogenase E1 component, beta
subunit (EC 1.2.4.1)
Length = 325
Score = 228 bits (582), Expect = 1e-59
Identities = 126/322 (39%), Positives = 186/322 (57%), Gaps = 4/322 (1%)
Query: 28 QMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTPITEAG 87
QMT+ A+N AL E+ D V + GE+VG G +++++GL +++G +RV DTP+ E+G
Sbjct: 3 QMTMVQAINDALKTELKNDQDVLIFGEDVGVNGGVFRVTEGLQKEFGEDRVFDTPLAESG 62
Query: 88 FTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAA 147
G+ +G A G +PV+E F + D I A++ + S G P+ R P G
Sbjct: 63 IGGLAMGLAVEGFRPVMEVQFLGFVFEVFDAIAGQIARTRFRSGGTKTAPVTIRSPFGGG 122
Query: 148 AGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLYGESFP 207
H+ PGLKV+ P DA+GLL ++IR DPVV+LE+ LY SF
Sbjct: 123 VHTPELHADNLEGILAQSPGLKVVIPSGPYDAKGLLISSIRSNDPVVYLEHMKLY-RSF- 180
Query: 208 VSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVINLRSI 267
EV + + + IGKA +++EG D++I + MV ++KAAE LEK+G S EVI+LR++
Sbjct: 181 -REEVPEEEYTIDIGKANVKKEGNDISIITYGAMVQESMKAAEELEKDGYSVEVIDLRTV 239
Query: 268 RPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAGADVPM 327
+P+D TI ASV KT R V V+E Q GVGA + A + E + L+AP+ R+A AD
Sbjct: 240 QPIDVDTIVASVEKTGRAVVVQEAQRQAGVGAAVVAELSERAILSLEAPIGRVAAADTIY 299
Query: 328 PYAANLERLAVPQIEDIVRAAK 349
P+ E + +P DI+ AK
Sbjct: 300 PF-TQAENVWLPNKNDIIEKAK 320
>ODPB_STAEP (Q8CPN2) Pyruvate dehydrogenase E1 component, beta
subunit (EC 1.2.4.1)
Length = 325
Score = 228 bits (581), Expect = 2e-59
Identities = 123/322 (38%), Positives = 189/322 (58%), Gaps = 4/322 (1%)
Query: 28 QMTVRDALNSALDEEMSADPKVFLMGEEVGEYQGAYKISKGLLEKYGPERVLDTPITEAG 87
QMT+ A+N AL E+ D V + GE+VG G +++++GL +++G +RV DTP+ E+G
Sbjct: 3 QMTMVQAINDALKSELKRDEDVLVFGEDVGVNGGVFRVTEGLQKEFGEDRVFDTPLAESG 62
Query: 88 FTGIGVGAAYYGLKPVVEFMTFNFSMQAIDHIINSAAKSNYMSAGQINVPIVFRGPNGAA 147
G+ +G A G +PV+E F + D + A++ + S G P+ R P G
Sbjct: 63 IGGLALGLAVTGFRPVMEIQFLGFVYEVFDEVAGQIARTRFRSGGTKPAPVTIRTPFGGG 122
Query: 148 AGVGAQHSHCYASWYGSCPGLKVLAPYSSEDARGLLKAAIRDPDPVVFLENELLYGESFP 207
H+ PGLKV+ P DA+GLL ++I+ DPVV+LE+ LY SF
Sbjct: 123 VHTPELHADNLEGILAQSPGLKVVIPSGPYDAKGLLISSIQSNDPVVYLEHMKLY-RSF- 180
Query: 208 VSAEVLDSSFCLPIGKAKIEREGKDVTITAFSKMVGFALKAAETLEKEGISAEVINLRSI 267
EV + + + IGKA +++EG D+T+ ++ MV +LKAAE LEK+G S EVI+LR++
Sbjct: 181 -REEVPEEEYKIDIGKANVKKEGNDITLISYGAMVQESLKAAEELEKDGYSVEVIDLRTV 239
Query: 268 RPLDRATINASVRKTNRLVTVEEGFPQHGVGAEICASVIEESFGYLDAPVERIAGADVPM 327
+P+D T+ ASV KT R V V+E Q GVGA++ A + E + L+AP+ R+A +D
Sbjct: 240 QPIDIDTLVASVEKTGRAVVVQEAQRQAGVGAQVAAELAERAILSLEAPIARVAASDTIY 299
Query: 328 PYAANLERLAVPQIEDIVRAAK 349
P+ E + +P +DI+ AK
Sbjct: 300 PF-TQAENVWLPNKKDIIEQAK 320
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.134 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 40,303,432
Number of Sequences: 164201
Number of extensions: 1705480
Number of successful extensions: 4661
Number of sequences better than 10.0: 150
Number of HSP's better than 10.0 without gapping: 125
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 4373
Number of HSP's gapped (non-prelim): 160
length of query: 362
length of database: 59,974,054
effective HSP length: 111
effective length of query: 251
effective length of database: 41,747,743
effective search space: 10478683493
effective search space used: 10478683493
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC146972.11


