

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146972.10 - phase: 0
(106 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
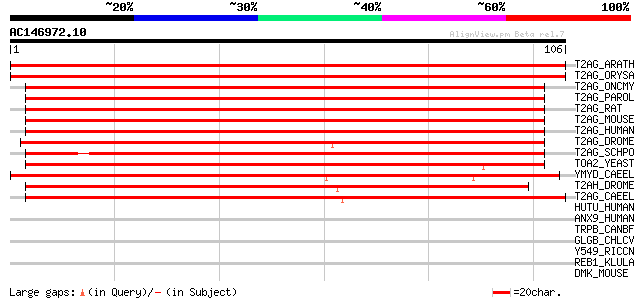
Score E
Sequences producing significant alignments: (bits) Value
T2AG_ARATH (Q39236) Transcription initiation factor IIA gamma ch... 198 2e-51
T2AG_ORYSA (Q94HL5) Transcription initiation factor IIA gamma ch... 191 3e-49
T2AG_ONCMY (Q90YG6) Transcription initiation factor IIA gamma ch... 114 4e-26
T2AG_PAROL (Q9IA78) Transcription initiation factor IIA gamma ch... 113 7e-26
T2AG_RAT (O08950) Transcription initiation factor IIA gamma chai... 112 1e-25
T2AG_MOUSE (Q80ZM7) Putative transcription initiation factor IIA... 112 1e-25
T2AG_HUMAN (P52657) Transcription initiation factor IIA gamma ch... 112 2e-25
T2AG_DROME (P52656) Transcription initiation factor IIA gamma ch... 107 5e-24
T2AG_SCHPO (O74948) Probable transcription initiation factor IIA... 107 7e-24
TOA2_YEAST (P32774) Transcription initiation factor IIA small ch... 91 4e-19
YMYD_CAEEL (Q9BIB4) Transcription initiation factor IIA small ch... 86 1e-17
T2AH_DROME (Q9W5B9) Transcription initiation factor IIA gamma-2 ... 85 3e-17
T2AG_CAEEL (Q9NEX2) Probable transcription initiation factor IIA... 80 1e-15
HUTU_HUMAN (Q96N76) Probable urocanate hydratase (EC 4.2.1.49) (... 29 2.3
ANX9_HUMAN (O76027) Annexin A9 (Annexin 31) (Annexin XXXI) 29 2.3
TRPB_CANBF (Q7VR00) Tryptophan synthase beta chain (EC 4.2.1.20) 28 3.9
GLGB_CHLCV (Q823Y5) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 28 3.9
Y549_RICCN (Q92I71) Hypothetical protein RC0549 27 6.6
REB1_KLULA (Q05950) DNA-binding protein REB1 (QBP) 27 6.6
DMK_MOUSE (P54265) Myotonin-protein kinase (EC 2.7.1.-) (Myotoni... 27 8.6
>T2AG_ARATH (Q39236) Transcription initiation factor IIA gamma chain
(TFIIA-gamma)
Length = 106
Score = 198 bits (503), Expect = 2e-51
Identities = 96/106 (90%), Positives = 104/106 (97%)
Query: 1 MATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK 60
MATFELYRRSTIGMCLTETLDEMVQ+GTLSPE+AIQVLVQFDKSMTEALE+QVK+KVSIK
Sbjct: 1 MATFELYRRSTIGMCLTETLDEMVQSGTLSPELAIQVLVQFDKSMTEALESQVKTKVSIK 60
Query: 61 GHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSKLLSQ 106
GHLHTYRFCDNVWTFILQDA+FK++D QENV RVKIVACDSKLL+Q
Sbjct: 61 GHLHTYRFCDNVWTFILQDAMFKSDDRQENVSRVKIVACDSKLLTQ 106
>T2AG_ORYSA (Q94HL5) Transcription initiation factor IIA gamma chain
(TFIIA-gamma)
Length = 106
Score = 191 bits (484), Expect = 3e-49
Identities = 94/106 (88%), Positives = 99/106 (92%)
Query: 1 MATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK 60
MATFELYRRSTIGMCLTETLDEMV +GTLSPE+AIQVLVQFDKSMTEALE QVKSKVSIK
Sbjct: 1 MATFELYRRSTIGMCLTETLDEMVSSGTLSPELAIQVLVQFDKSMTEALENQVKSKVSIK 60
Query: 61 GHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSKLLSQ 106
GHLHTYRFCDNVWTFIL +A FKNE+ E VG+VKIVACDSKLLSQ
Sbjct: 61 GHLHTYRFCDNVWTFILTEASFKNEETTEQVGKVKIVACDSKLLSQ 106
>T2AG_ONCMY (Q90YG6) Transcription initiation factor IIA gamma chain
(TFIIA P12 subunit) (TFIIA-12) (TFIIAS) (TFIIA-gamma)
Length = 108
Score = 114 bits (285), Expect = 4e-26
Identities = 50/99 (50%), Positives = 75/99 (75%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
++LYR +T+G L E+LDE++Q ++P++A+QVL+QFDK++ AL +V+++V+ KG L
Sbjct: 3 YQLYRNTTLGNSLQESLDELIQTQQITPQLALQVLLQFDKAINTALANRVRNRVNFKGSL 62
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
+TYRFCDNVWTF+L D F+ + V +VKIVACD K
Sbjct: 63 NTYRFCDNVWTFVLNDVEFREVTDLVKVDKVKIVACDGK 101
>T2AG_PAROL (Q9IA78) Transcription initiation factor IIA gamma chain
(TFIIA P12 subunit) (TFIIA-12) (TFIIAS) (TFIIA-gamma)
Length = 111
Score = 113 bits (283), Expect = 7e-26
Identities = 49/99 (49%), Positives = 76/99 (76%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
++LYR +T+G L E+LDE++Q ++P++A+QVL+QFDK++ AL ++V+++V+ +G L
Sbjct: 3 YQLYRNTTLGNSLQESLDELIQTQQITPQLALQVLLQFDKAINTALASRVRNRVNFRGSL 62
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
+TYRFCDNVWTF+L D F+ + V +VKIVACD K
Sbjct: 63 NTYRFCDNVWTFVLNDVEFREVTDLVKVDKVKIVACDGK 101
>T2AG_RAT (O08950) Transcription initiation factor IIA gamma chain
(TFIIA P12 subunit) (TFIIA-12) (TFIIAS) (TFIIA-gamma)
Length = 109
Score = 112 bits (281), Expect = 1e-25
Identities = 49/99 (49%), Positives = 75/99 (75%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
++LYR +T+G L E+LDE++Q+ ++P++A+QVL+QFDK++ AL +V+++V+ +G L
Sbjct: 3 YQLYRNTTLGNSLQESLDELIQSQQITPQLALQVLLQFDKAINSALAQRVRNRVNFRGSL 62
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
+TYRFCDNVWTF+L D F+ V +VKIVACD K
Sbjct: 63 NTYRFCDNVWTFVLNDVEFREVTELIKVDKVKIVACDGK 101
>T2AG_MOUSE (Q80ZM7) Putative transcription initiation factor IIA
gamma chain (TFIIA P12 subunit) (TFIIA-12) (TFIIAS)
(TFIIA-gamma)
Length = 109
Score = 112 bits (281), Expect = 1e-25
Identities = 49/99 (49%), Positives = 75/99 (75%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
++LYR +T+G L E+LDE++Q+ ++P++A+QVL+QFDK++ AL +V+++V+ +G L
Sbjct: 3 YQLYRNTTLGNSLQESLDELIQSQQITPQLALQVLLQFDKAINSALAQRVRNRVNFRGSL 62
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
+TYRFCDNVWTF+L D F+ V +VKIVACD K
Sbjct: 63 NTYRFCDNVWTFVLNDVEFREVTELIKVDKVKIVACDGK 101
>T2AG_HUMAN (P52657) Transcription initiation factor IIA gamma chain
(TFIIA P12 subunit) (TFIIA-12) (TFIIAS) (TFIIA-gamma)
Length = 109
Score = 112 bits (280), Expect = 2e-25
Identities = 49/99 (49%), Positives = 75/99 (75%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
++LYR +T+G L E+LDE++Q+ ++P++A+QVL+QFDK++ AL +V+++V+ +G L
Sbjct: 3 YQLYRNTTLGNSLQESLDELIQSQQITPQLALQVLLQFDKAINAALAQRVRNRVNFRGSL 62
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
+TYRFCDNVWTF+L D F+ V +VKIVACD K
Sbjct: 63 NTYRFCDNVWTFVLNDVEFREVTELIKVDKVKIVACDGK 101
>T2AG_DROME (P52656) Transcription initiation factor IIA gamma chain
(TFIIA P14 subunit) (TFIIA-14) (dTFIIA-S) (TFIIA-gamma)
Length = 106
Score = 107 bits (267), Expect = 5e-24
Identities = 51/101 (50%), Positives = 73/101 (71%), Gaps = 1/101 (0%)
Query: 3 TFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIK-G 61
+++LYR +T+G L E+LDE++Q G ++P +A +VL+QFDKS+ AL +VK++V+ K G
Sbjct: 2 SYQLYRNTTLGNTLQESLDELIQYGQITPGLAFKVLLQFDKSINNALNQRVKARVTFKAG 61
Query: 62 HLHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
L+TYRFCDNVWT +L D F+ V +VKIVACD K
Sbjct: 62 KLNTYRFCDNVWTLMLNDVEFREVHEIVKVDKVKIVACDGK 102
>T2AG_SCHPO (O74948) Probable transcription initiation factor IIA
small chain
Length = 107
Score = 107 bits (266), Expect = 7e-24
Identities = 48/99 (48%), Positives = 74/99 (74%), Gaps = 2/99 (2%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
+ELYRRS I LT+ LD+++ G +SP++A++VL FDKSMTEAL +V+S+++ KGHL
Sbjct: 5 YELYRRSRIS--LTDALDDLISQGKISPQLAMKVLFNFDKSMTEALAEKVRSRLTFKGHL 62
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSK 102
TYRFCD VWTFI+++ F+ ++ +++IVAC ++
Sbjct: 63 DTYRFCDEVWTFIIKNPSFRFDNETVTSNKIRIVACATR 101
>TOA2_YEAST (P32774) Transcription initiation factor IIA small chain
(TFIIA 13.5 kDa subunit)
Length = 122
Score = 91.3 bits (225), Expect = 4e-19
Identities = 45/113 (39%), Positives = 71/113 (62%), Gaps = 14/113 (12%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHL 63
+ELYRRSTIG L + LD ++ +G + +A++VL FDK + E L+ +SK+++KG+L
Sbjct: 7 YELYRRSTIGNSLVDALDTLISDGRIEASLAMRVLETFDKVVAETLKDNTQSKLTVKGNL 66
Query: 64 HTYRFCDNVWTFILQDALFKNEDNQE--------------NVGRVKIVACDSK 102
TY FCD+VWTFI+++ ED+ +V +++IVAC+SK
Sbjct: 67 DTYGFCDDVWTFIVKNCQVTVEDSHRDASQNGSGDSQSVISVDKLRIVACNSK 119
>YMYD_CAEEL (Q9BIB4) Transcription initiation factor IIA small chain
homolog
Length = 139
Score = 86.3 bits (212), Expect = 1e-17
Identities = 44/108 (40%), Positives = 67/108 (61%), Gaps = 3/108 (2%)
Query: 1 MATFELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSI- 59
M+++ LYR +T+G L +TL++M G L+ +A +VL QFDKSM + + K K++
Sbjct: 1 MSSYALYRGTTLGQALDKTLEDMESEGLLTKSLASKVLQQFDKSMNKQISRLPKEKMNFC 60
Query: 60 KGHLHTYRFCDNVWTFILQDALFKNEDN--QENVGRVKIVACDSKLLS 105
L TYR+CDNVWTFIL + K+ E + ++K+VACD + S
Sbjct: 61 ATQLLTYRYCDNVWTFILNNVTLKDPQRSFDEPIDKLKVVACDGRQTS 108
>T2AH_DROME (Q9W5B9) Transcription initiation factor IIA gamma-2
chain (TFIIA-gamma-2)
Length = 107
Score = 84.7 bits (208), Expect = 3e-17
Identities = 43/97 (44%), Positives = 65/97 (66%), Gaps = 1/97 (1%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKG-H 62
++ YR +T+G L +TLDEM++ G ++ +IA VL+++DKS++ AL+ S +S
Sbjct: 3 YQHYRATTLGRTLQDTLDEMMERGDITKKIANLVLLRYDKSISTALKDHGTSNMSFTAER 62
Query: 63 LHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVAC 99
L T+R CDNVWT IL+DA F+ + + V VKIVAC
Sbjct: 63 LETFRCCDNVWTLILKDAEFREDQHSLKVDVVKIVAC 99
>T2AG_CAEEL (Q9NEX2) Probable transcription initiation factor IIA
gamma chain (TFIIA P12 subunit) (TFIIA-12) (TFIIAS)
(TFIIA-gamma)
Length = 113
Score = 79.7 bits (195), Expect = 1e-15
Identities = 38/104 (36%), Positives = 64/104 (61%), Gaps = 1/104 (0%)
Query: 4 FELYRRSTIGMCLTETLDEMVQNGTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGH- 62
++LYR +T+G L +TLD+ V + + ++ +++ FDKS+ + L + K+KV+ +
Sbjct: 6 YQLYRNTTLGQALQKTLDDFVGDQMIPDSLSKKIMDSFDKSINKILPHKAKNKVNFRADK 65
Query: 63 LHTYRFCDNVWTFILQDALFKNEDNQENVGRVKIVACDSKLLSQ 106
L YR+CDNVWTFI++ ++ V R+KIVACD + Q
Sbjct: 66 LRAYRYCDNVWTFIVEQIDLRDAVEGGTVDRLKIVACDGQTKGQ 109
>HUTU_HUMAN (Q96N76) Probable urocanate hydratase (EC 4.2.1.49)
(Urocanase) (Imidazolonepropionate hydrolase)
Length = 676
Score = 28.9 bits (63), Expect = 2.3
Identities = 15/53 (28%), Positives = 30/53 (56%), Gaps = 4/53 (7%)
Query: 24 VQNGTLSP---EIAIQ-VLVQFDKSMTEALETQVKSKVSIKGHLHTYRFCDNV 72
V+ +LSP ++A++ L F + E L + ++ + GH++ YRFC ++
Sbjct: 31 VRTPSLSPVEKQLALRNALRYFPPDVQELLAPEFAQELQLYGHIYMYRFCPDI 83
>ANX9_HUMAN (O76027) Annexin A9 (Annexin 31) (Annexin XXXI)
Length = 338
Score = 28.9 bits (63), Expect = 2.3
Identities = 22/82 (26%), Positives = 40/82 (47%), Gaps = 6/82 (7%)
Query: 30 SPEIAIQVLVQFDKSMTEALETQVKSK------VSIKGHLHTYRFCDNVWTFILQDALFK 83
+PE I+V Q+ +S + LE V+++ V++ G + + L AL +
Sbjct: 219 NPEHLIRVFDQYQRSTGQELEEAVQNRFHGDAQVALLGLASVIKNTPLYFADKLHQALQE 278
Query: 84 NEDNQENVGRVKIVACDSKLLS 105
E N + + R+ I C++ LLS
Sbjct: 279 TEPNYQVLIRILISRCETDLLS 300
>TRPB_CANBF (Q7VR00) Tryptophan synthase beta chain (EC 4.2.1.20)
Length = 398
Score = 28.1 bits (61), Expect = 3.9
Identities = 18/67 (26%), Positives = 28/67 (40%), Gaps = 8/67 (11%)
Query: 27 GTLSPEIAIQVLVQFDKSMTEALETQVKSKVSIKGHLHTY-------RFCDNVWTFILQD 79
G P+I I L+Q +K EA++ + + + K LH Y C N+ +
Sbjct: 14 GMYVPQILIPALIQLEKEFIEAMK-DISFQTTFKNLLHHYAGRPTPLTLCQNLTSGTCSK 72
Query: 80 ALFKNED 86
K ED
Sbjct: 73 LYLKRED 79
>GLGB_CHLCV (Q823Y5) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 719
Score = 28.1 bits (61), Expect = 3.9
Identities = 16/39 (41%), Positives = 19/39 (48%), Gaps = 3/39 (7%)
Query: 47 EALETQVKSKVSIKGHLHTYRF---CDNVWTFILQDALF 82
EAL V K + H HTY F C V F++ ALF
Sbjct: 349 EALYESVDHKDPLHPHWHTYTFDYRCSEVVNFLIGSALF 387
>Y549_RICCN (Q92I71) Hypothetical protein RC0549
Length = 315
Score = 27.3 bits (59), Expect = 6.6
Identities = 17/46 (36%), Positives = 19/46 (40%)
Query: 42 DKSMTEALETQVKSKVSIKGHLHTYRFCDNVWTFILQDALFKNEDN 87
D L Q+ SK KG L D V L+DALF DN
Sbjct: 46 DLKNINGLNLQISSKCFAKGKLPFLASSDEVRFNCLRDALFDESDN 91
>REB1_KLULA (Q05950) DNA-binding protein REB1 (QBP)
Length = 595
Score = 27.3 bits (59), Expect = 6.6
Identities = 13/34 (38%), Positives = 20/34 (58%), Gaps = 1/34 (2%)
Query: 41 FDKSMTEALETQVKSKVSIKGHLHTYRFCDNVWT 74
FD+S EALE +K I+G L + C+ +W+
Sbjct: 268 FDESEEEALEQFIKEYQKIRG-LSRRQICERIWS 300
>DMK_MOUSE (P54265) Myotonin-protein kinase (EC 2.7.1.-) (Myotonic
dystrophy protein kinase) (MDPK) (DM-kinase) (DMK)
(DMPK) (MT-PK)
Length = 631
Score = 26.9 bits (58), Expect = 8.6
Identities = 14/53 (26%), Positives = 30/53 (56%)
Query: 43 KSMTEALETQVKSKVSIKGHLHTYRFCDNVWTFILQDALFKNEDNQENVGRVK 95
+ + EALE +V ++ S+ L R + ++ LQ+A +N D + +V +++
Sbjct: 471 QQLQEALEEEVLTRQSLSRELEAIRTANQNFSSQLQEAEVRNRDLEAHVRQLQ 523
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.133 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,786,969
Number of Sequences: 164201
Number of extensions: 335270
Number of successful extensions: 840
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 819
Number of HSP's gapped (non-prelim): 22
length of query: 106
length of database: 59,974,054
effective HSP length: 82
effective length of query: 24
effective length of database: 46,509,572
effective search space: 1116229728
effective search space used: 1116229728
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146972.10


