

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146941.4 + phase: 0
(328 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
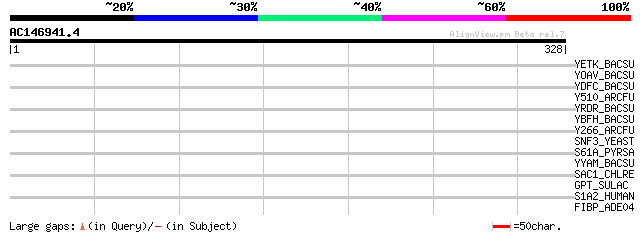
YETK_BACSU (O31540) Hypothetical transport protein yetK 42 0.002
YOAV_BACSU (O34416) Hypothetical transport protein yoaV 41 0.004
YDFC_BACSU (P96680) Hypothetical transport protein ydfC 40 0.011
Y510_ARCFU (O29740) Hypothetical transport protein AF0510 37 0.089
YRDR_BACSU (O07086) Hypothetical transport protein yrdR 35 0.34
YBFH_BACSU (O31448) Hypothetical transport protein ybhF 34 0.58
Y266_ARCFU (O29973) Hypothetical transport protein AF0266 33 0.98
SNF3_YEAST (P10870) High-affinity glucose transporter SNF3 33 1.3
S61A_PYRSA (P38379) Protein transport protein Sec61 alpha subunit 32 1.7
YYAM_BACSU (P37511) Hypothetical transport protein yyaM 32 2.9
SAC1_CHLRE (Q39593) Putative sulfur deprivation response regulator 32 2.9
GPT_SULAC (P39465) Putative UDP-N-acetylglucosamine--dolichyl-ph... 32 2.9
S1A2_HUMAN (P46721) Solute carrier organic anion transporter fam... 31 4.9
FIBP_ADE04 (P36844) Fiber protein 30 8.3
>YETK_BACSU (O31540) Hypothetical transport protein yetK
Length = 330
Score = 42.0 bits (97), Expect = 0.002
Identities = 28/135 (20%), Positives = 60/135 (43%), Gaps = 7/135 (5%)
Query: 24 LIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVLTKIFL 83
+ + G+ ++ ++VE +F + R L+A+V + L E+G P +V + +
Sbjct: 37 MAIVGSSVVVGKLMVERIPVFLSSGLRFLIASVVLLMLLFCIEKGFPALTKKDVFV-LLV 95
Query: 84 NGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNGRAKCV 143
F G+ + + YG++ T+ T + + P+ + + E + T +
Sbjct: 96 QSFTGVFLFSICLLYGVQYTTGTESGILTSTTPMLIGILSFFLLREKIEKKT------LI 149
Query: 144 GAILCVAGTLAARLY 158
G +L V G +A L+
Sbjct: 150 GILLAVCGVMAINLF 164
>YOAV_BACSU (O34416) Hypothetical transport protein yoaV
Length = 292
Score = 41.2 bits (95), Expect = 0.004
Identities = 49/230 (21%), Positives = 93/230 (40%), Gaps = 24/230 (10%)
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
+ K L ++ MS++M L Y GI TY + F++ + + + ++ + N
Sbjct: 59 IQKEHLKSYIIMSLLMGLGYMGI----LTYGMQFVDSGKTSVLVYTMPIFVTVISHFSLN 114
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFHSVAAHKTQMLRGTLFLIGACFSY-- 195
+ + V G GKE + L G L ++ A S+
Sbjct: 115 EKMNVYKTMGLVCGLFGLLFIFGKEMLNIDQSA-----------LFGELCVLVAALSWGI 163
Query: 196 -SAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITILY 254
+ + +Q K +++ + W +M+ VM + S + A V S++ ++ W+L L+
Sbjct: 164 ANVFSKLQFKHIDIIHMNAWHLMMGAVMLLVFSFIFEA-VPSAEWTYQAVWSL-----LF 217
Query: 255 SGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTV 304
+G LST F + W + + SM V F + L E +T+
Sbjct: 218 NGLLSTGFTFVVWFWVLNQIQASKASMALMFVPVLALFFGWLQLHEQITI 267
Score = 30.0 bits (66), Expect = 8.3
Identities = 19/109 (17%), Positives = 45/109 (40%), Gaps = 6/109 (5%)
Query: 43 IFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVLTKIFLNGFVGMSMVMVLYYYGIRD 102
I + ++ +++ AV + + FE ++ + + + NG + V++++ +
Sbjct: 177 IIHMNAWHLMMGAVMLLVFSFIFEAVPSAEWTYQAVWSLLFNGLLSTGFTFVVWFWVLNQ 236
Query: 103 TSATYALNFLNLIPICTFLTAILFRMENLNIHTWNGRAKCVGAILCVAG 151
A+ A L +P+ L E + I+ +GA+L G
Sbjct: 237 IQASKASMALMFVPVLALFFGWLQLHEQITINI------ILGALLICCG 279
>YDFC_BACSU (P96680) Hypothetical transport protein ydfC
Length = 306
Score = 39.7 bits (91), Expect = 0.011
Identities = 36/164 (21%), Positives = 69/164 (41%), Gaps = 25/164 (15%)
Query: 46 LTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSA 105
L +R+L+ ++ + A+ + P + + IFL GF+G + +L G + SA
Sbjct: 38 LALFRLLIGSMALLLFAVLTQMRLP---DLKDIPAIFLLGFLGFAFYHILLNIGEKTVSA 94
Query: 106 TYALNFLNLIPICTFLTAILFRMENLNIHTWNGRAKCVGAILCVAGTLAARLYKGKEFYI 165
A + PI + + + LF E+ W +G+++ + G L G +F
Sbjct: 95 GVASLLVTTAPIFSAMLSRLFYKEHFGFTKW------LGSMISLLGVLLIAFGAG-DF-- 145
Query: 166 AQYHSFHSVAAHKTQMLRGTLFLIGACFSYSAWFFMQVKLVEVF 209
T + G L ++ A FS S +F Q + ++ +
Sbjct: 146 -------------TYSMSGILVILLAAFSESIYFVFQARYIKKY 176
>Y510_ARCFU (O29740) Hypothetical transport protein AF0510
Length = 289
Score = 36.6 bits (83), Expect = 0.089
Identities = 22/106 (20%), Positives = 45/106 (41%), Gaps = 6/106 (5%)
Query: 46 LTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSA 105
LT+Y + + + P A+ P F+ E + + + V++YY + + +
Sbjct: 180 LTAYAFALGTIFLIPFALMSGFANPVTFNPETVAALLYLSILCSVFAYVVWYYALTNADS 239
Query: 106 TYALNFLNLIPICTFLTAILFRMENLNIHTWNGRAKCVGAILCVAG 151
T ++ L+P+ T + A E + T +G I+ +AG
Sbjct: 240 TSVAVYVYLVPLFTAIFAFYALNEKPDFFT------AIGGIITIAG 279
>YRDR_BACSU (O07086) Hypothetical transport protein yrdR
Length = 321
Score = 34.7 bits (78), Expect = 0.34
Identities = 38/167 (22%), Positives = 76/167 (44%), Gaps = 15/167 (8%)
Query: 5 DVKEWFVLSSILWSMIFVQLIVTGTQI--LSRIILVEGTFIFALTSYRVLVAAVCVAPLA 62
++ +F L L+S++F+ + V G + + I E + + T + A+ A
Sbjct: 149 NISFFFSLGDHLFSILFIFIGVLGWVVYTMGGQIFREWSTLRYSTLTCLFGTAITGIMTA 208
Query: 63 IYFERGQPKNFSCEVLTKI-----FLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPI 117
I +G S +V+ I F+ G+ + ++ + YGI+ S+ + F+N +PI
Sbjct: 209 ILTAQGYVSVPSIKVIAAIKYDFLFMITLPGI-IALLSWNYGIKILSSINGILFINFVPI 267
Query: 118 CTFLTAILFRMENLNIHTWNGRAKCVGAILCVAGTLAARLYKGKEFY 164
T L ++ + NI ++ VG + + G + +Y+ KE Y
Sbjct: 268 TTLLIMVI---KGYNITAFD----IVGTLFVIIGLILNNIYQRKEDY 307
>YBFH_BACSU (O31448) Hypothetical transport protein ybhF
Length = 306
Score = 33.9 bits (76), Expect = 0.58
Identities = 48/208 (23%), Positives = 86/208 (41%), Gaps = 45/208 (21%)
Query: 22 VQLIVTGTQILSRIILVEGTFIFALTSYRVL---VAAVCVAPLAIYFERGQPKNFSCEVL 78
V +++ GT +S +L+ + YR L +A + V P I F +N+ E+L
Sbjct: 15 VTILIWGTTFISTKVLLADFSPMEILFYRFLMGFIALILVRPNMIPF-----RNWRQELL 69
Query: 79 TKIFLNGFVGMSMVMVLYYYGIRDTSATY--ALNFLNLIPICTFLTAIL--FRMEN---- 130
F G + V Y+ + + + TY A N ++ I +TA+L F +E
Sbjct: 70 -------FAGAGLFGVTLYFLLENIALTYTYASNVGMIVSIIPMITAVLAHFLLEGEKLR 122
Query: 131 -------------LNIHTWNG----RAKCVGAILCVAGTLAARLYKGKEFYIAQY--HSF 171
L + T+NG R +G I+ AA ++ G ++ + + +
Sbjct: 123 LTFLIGFISALIGLLLITFNGNVVLRLNPLGDIMAAG---AALVFGGYSIFMKKLSAYEY 179
Query: 172 HSVAAHKTQMLRGTLFLIGACFSYSAWF 199
H + + L G LF++ A F + F
Sbjct: 180 HIIELTQRVFLYGLLFMVPALFLFDFHF 207
>Y266_ARCFU (O29973) Hypothetical transport protein AF0266
Length = 276
Score = 33.1 bits (74), Expect = 0.98
Identities = 35/150 (23%), Positives = 59/150 (39%), Gaps = 23/150 (15%)
Query: 6 VKEWFVLSSIL--WSMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAI 63
+K+W VL+ + W + F I + LS I + F A T + V++
Sbjct: 1 MKKWIVLALTVTFWGLAFTA-IKYSVRFLSPIAIASLRFAIANTLFAVIIIL-------- 51
Query: 64 YFERGQPKNFSCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTA 123
K + L K+F G G+S+ V G S+ A ++L PI + +
Sbjct: 52 ------GKRIKWKDLPKVFALGIFGVSVYHVFLNLGEVYISSGVASVVISLAPIFVLILS 105
Query: 124 ILFRMENLNIHTWNGRAKCVGAILCVAGTL 153
+F E + +K VG I+ G +
Sbjct: 106 AIFLRERITY------SKVVGIIIAFLGVV 129
>SNF3_YEAST (P10870) High-affinity glucose transporter SNF3
Length = 818
Score = 32.7 bits (73), Expect = 1.3
Identities = 39/157 (24%), Positives = 68/157 (42%), Gaps = 20/157 (12%)
Query: 67 RGQPKNFSCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILF 126
+ +PK + + T I L F S + ++YYG+ + T N L+ T+ ++F
Sbjct: 281 KSRPKQ-TLRMFTGIALQAFQQFSGINFIFYYGVNFFNKTGVSNSY-LVSFITYAVNVVF 338
Query: 127 RMENLNIHTWNGRAKCV---GAILCVAGTLAARLYKGKEFYIAQYHSFHSVAAHKTQMLR 183
+ L + GR K + G I+ +A + A + S +VAA K +
Sbjct: 339 NVPGLFFVEFFGRRKVLVVGGVIMTIANFIVAIV----------GCSLKTVAAAKVMIAF 388
Query: 184 GTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQC 220
LF+ A FS + + V E++PL G+ +C
Sbjct: 389 ICLFI--AAFSATWGGVVWVISAELYPL---GVRSKC 420
>S61A_PYRSA (P38379) Protein transport protein Sec61 alpha subunit
Length = 494
Score = 32.3 bits (72), Expect = 1.7
Identities = 43/180 (23%), Positives = 74/180 (40%), Gaps = 26/180 (14%)
Query: 23 QLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV-LTKI 81
QL++T T + L E + L + L+A V V L +YF+ F E+ +T
Sbjct: 222 QLMITKTDKVRA--LQEAFYRQNLPNVTNLLATVLVFVLVVYFQ-----GFQVELPITPA 274
Query: 82 FLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRM--ENLNIHT---W 136
G G + L+Y TS + L+ F++ IL++ EN+ IH W
Sbjct: 275 KSKGMAGQFYPIKLFY-----TSNMPIILQTALVSNLYFISQILYKRYPENIIIHILGRW 329
Query: 137 NGRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFHSVAAHKTQMLRGTLFLIGACFSYS 196
V + G +A +YI+ +SF + + L +F++ +C +S
Sbjct: 330 EEPEMSVSGQMRPVGGIA--------YYISPLNSFAEIVSDPVHALLYIIFILASCALFS 381
>YYAM_BACSU (P37511) Hypothetical transport protein yyaM
Length = 305
Score = 31.6 bits (70), Expect = 2.9
Identities = 27/126 (21%), Positives = 57/126 (44%), Gaps = 9/126 (7%)
Query: 31 ILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQP-KNFSCEVLTKIFLNGFVGM 89
+L R + +GT + + T+Y +++ AV + ++++ + N V I F
Sbjct: 171 VLGRRFVKDGTPL-STTTYTMVIGAVSLIVVSLFTSKPVSLSNIPIGVWGAIAFMAFFTS 229
Query: 90 SMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNGRAKCVGAILCV 149
+ + + GI++ A+ F NL+P+ T + + + T + +GA+L +
Sbjct: 230 VLGYLWWNQGIKEIGASKTSLFFNLVPVVTMIISFA-------VGTPIKVFQVIGAVLVI 282
Query: 150 AGTLAA 155
G L A
Sbjct: 283 LGVLTA 288
>SAC1_CHLRE (Q39593) Putative sulfur deprivation response regulator
Length = 585
Score = 31.6 bits (70), Expect = 2.9
Identities = 31/104 (29%), Positives = 43/104 (40%), Gaps = 10/104 (9%)
Query: 131 LNIHTWNGRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFHSVAAHKTQMLRGTLF--L 188
LN+ +W GRA V AI L A L G + Y +A ++G L+ +
Sbjct: 470 LNVFSWMGRAGPVAAIYASTSLLTALLSNGAAVTL-MYPIARDLAKQAGVSIKGPLYALM 528
Query: 189 IGACFSYSAWFFMQVKLVEVFPLRY-------WGIMLQCVMAAI 225
IGA +S Q L+ P Y +G+ LQ V A I
Sbjct: 529 IGASSDFSTPIGYQTNLMVSGPGGYRFLDFTRFGLPLQFVAALI 572
>GPT_SULAC (P39465) Putative
UDP-N-acetylglucosamine--dolichyl-phosphate
N-acetylglucosaminephosphotransferase (EC 2.7.8.15)
(GPT) (G1PT) (N-acetylglucosamine-1-phosphate
transferase) (GlcNAc-1-P transferase)
Length = 328
Score = 31.6 bits (70), Expect = 2.9
Identities = 36/140 (25%), Positives = 58/140 (40%), Gaps = 19/140 (13%)
Query: 38 VEGTFIFALTSYRV------LVAAVCVAPLAIYFERGQPKNFSCEVLTKIFLNGFVGMSM 91
V G+F F LTSY + +V ++ ++ L I F F+ T+ FL F S+
Sbjct: 61 VAGSFTFLLTSYNLSPGIENVVVSILLSSLIIGFLGLLDDIFNISQATRAFLPIFA--SI 118
Query: 92 VMVLYYYGIRDTSATY-------ALNFLNLIPICTFLTAILFRMENLNIHTWNGRAKCVG 144
++LY G S + L ++ ++P +TA F M + NG +G
Sbjct: 119 PLILYSVGHTIISIPFLGKVNFGILFYIIILPATLTITANAFNM----LEGLNGLGAGMG 174
Query: 145 AILCVAGTLAARLYKGKEFY 164
I+ +A G FY
Sbjct: 175 LIMALALAYIGLKSGGTSFY 194
>S1A2_HUMAN (P46721) Solute carrier organic anion transporter family
member 1A2 (Solute carrier family 21 member 3)
(Sodium-independent organic anion transporter) (Organic
anion transporting polypeptide 1) (OATP1) (OATP-A)
Length = 670
Score = 30.8 bits (68), Expect = 4.9
Identities = 19/58 (32%), Positives = 26/58 (44%), Gaps = 10/58 (17%)
Query: 71 KNFSCEVLTKIFL-------NGFVGMSMVMVLYY---YGIRDTSATYALNFLNLIPIC 118
K+ SC + +F+ N FV M M Y YGI + A + + NL PIC
Sbjct: 310 KSLSCNPIYMLFILVSVIQFNAFVNMISFMPKYLEQQYGISSSDAIFLMGIYNLPPIC 367
>FIBP_ADE04 (P36844) Fiber protein
Length = 426
Score = 30.0 bits (66), Expect = 8.3
Identities = 13/57 (22%), Positives = 30/57 (51%)
Query: 55 AVCVAPLAIYFERGQPKNFSCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNF 111
AV P + + + Q ++ ++++NG V M++ + G DT++ Y+++F
Sbjct: 343 AVGFMPNSTAYPKTQSSTTKNNIVGQVYMNGDVSKPMLLTITLNGTDDTTSAYSMSF 399
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.332 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,704,648
Number of Sequences: 164201
Number of extensions: 1301286
Number of successful extensions: 3564
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 3557
Number of HSP's gapped (non-prelim): 18
length of query: 328
length of database: 59,974,054
effective HSP length: 110
effective length of query: 218
effective length of database: 41,911,944
effective search space: 9136803792
effective search space used: 9136803792
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 66 (30.0 bits)
Medicago: description of AC146941.4


