

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146941.2 - phase: 0 /pseudo
(240 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
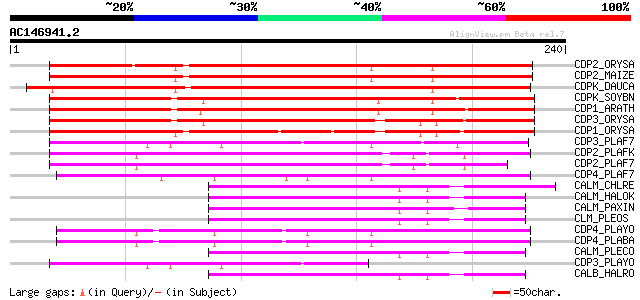
Score E
Sequences producing significant alignments: (bits) Value
CDP2_ORYSA (P53683) Calcium-dependent protein kinase, isoform 2 ... 217 2e-56
CDP2_MAIZE (P49101) Calcium-dependent protein kinase 2 (EC 2.7.1... 217 2e-56
CDPK_DAUCA (P28582) Calcium-dependent protein kinase (EC 2.7.1.-... 216 4e-56
CDPK_SOYBN (P28583) Calcium-dependent protein kinase SK5 (EC 2.7... 191 2e-48
CDP1_ARATH (Q06850) Calcium-dependent protein kinase, isoform AK... 188 1e-47
CDP3_ORYSA (P53684) Calcium-dependent protein kinase, isoform 11... 158 1e-38
CDP1_ORYSA (P53682) Calcium-dependent protein kinase, isoform 1 ... 139 8e-33
CDP3_PLAF7 (Q9NJU9) Calcium-dependent protein kinase 3 (EC 2.7.1... 99 7e-21
CDP2_PLAFK (O15865) Calcium-dependent protein kinase 2 (EC 2.7.1... 91 2e-18
CDP2_PLAF7 (Q8ICR0) Calcium-dependent protein kinase 2 (EC 2.7.1... 82 9e-16
CDP4_PLAF7 (Q8IBS5) Calcium-dependent protein kinase 4 (EC 2.7.1... 81 3e-15
CALM_CHLRE (P04352) Calmodulin (CaM) 81 3e-15
CALM_HALOK (Q95NI4) Calmodulin (CaM) 80 6e-15
CALM_PAXIN (Q8X187) Calmodulin (CaM) 79 1e-14
CLM_PLEOS (O94739) Calmodulin (CaM) 78 2e-14
CDP4_PLAYO (Q7RJG2) Calcium-dependent protein kinase 4 (EC 2.7.1... 77 3e-14
CDP4_PLABA (P62345) Calcium-dependent protein kinase 4 (EC 2.7.1... 77 3e-14
CALM_PLECO (P11120) Calmodulin (CaM) 77 3e-14
CDP3_PLAYO (Q7RAV5) Calcium-dependent protein kinase 3 (EC 2.7.1... 77 4e-14
CALB_HALRO (O96081) Calmodulin B (CaM B) 77 4e-14
>CDP2_ORYSA (P53683) Calcium-dependent protein kinase, isoform 2 (EC
2.7.1.-) (CDPK 2)
Length = 533
Score = 217 bits (553), Expect = 2e-56
Identities = 129/219 (58%), Positives = 154/219 (69%), Gaps = 13/219 (5%)
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNR------- 70
WPSIS SAKDLVRKML DPK+RI++ +VL HPW++ DGEA D P+D+AVL+R
Sbjct: 309 WPSISESAKDLVRKMLTQDPKKRITSAQVLQHPWLR-DGEASDKPIDSAVLSRMKQFRAM 367
Query: 71 NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
N L++ + K++ S L+EEEI GLKQMF MDTDNSGTIT EELK GL K G++LSE +
Sbjct: 368 NKLKK--MALKVIASNLNEEEIKGLKQMFTNMDTDNSGTITYEELKAGLAKLGSKLSEAE 425
Query: 131 VKQLMEAADADGNGIIDYDEFITAS--QQLCTRTD*TEKNMFILPSNTSIKIT-TELEQA 187
VKQLMEAAD DGNG IDY EFITA+ + R + K + S IT ELE A
Sbjct: 426 VKQLMEAADVDGNGSIDYVEFITATMHRHKLERDEHLFKAFQYFDKDNSGFITRDELESA 485
Query: 188 LHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKG 226
L E+ M D IK+IISEVD DNDGRINY+EF AMMR G
Sbjct: 486 LIEHEMGDTSTIKDIISEVDTDNDGRINYEEFCAMMRGG 524
>CDP2_MAIZE (P49101) Calcium-dependent protein kinase 2 (EC 2.7.1.-)
(CDPK 2)
Length = 513
Score = 217 bits (552), Expect = 2e-56
Identities = 127/219 (57%), Positives = 153/219 (68%), Gaps = 12/219 (5%)
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNR------- 70
WPSIS SAKDLVRKML DPK+R+++ +VL H W++E GEA D P+D+AVL+R
Sbjct: 289 WPSISESAKDLVRKMLTRDPKKRLTSAQVLQHQWLREGGEASDKPIDSAVLSRMKQFRAM 348
Query: 71 NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
N L++ + K++ S L+EEEI GLKQMF MDTDNSGTIT EELK GL K G++LSE +
Sbjct: 349 NKLKK--MALKVIASNLNEEEIKGLKQMFMNMDTDNSGTITYEELKAGLAKLGSKLSEAE 406
Query: 131 VKQLMEAADADGNGIIDYDEFITAS--QQLCTRTD*TEKNMFILPSNTSIKIT-TELEQA 187
VKQLMEAAD DGNG IDY EFITA+ + R + K + S IT ELE A
Sbjct: 407 VKQLMEAADVDGNGSIDYVEFITATMHRHKLERDEHLFKAFQYFDKDNSGFITRDELESA 466
Query: 188 LHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKG 226
L E+ M D I+EIISEVD DNDGRINY+EF AMMR G
Sbjct: 467 LIEHEMGDTSTIREIISEVDTDNDGRINYEEFCAMMRGG 505
>CDPK_DAUCA (P28582) Calcium-dependent protein kinase (EC 2.7.1.-)
(CDPK)
Length = 532
Score = 216 bits (550), Expect = 4e-56
Identities = 124/227 (54%), Positives = 156/227 (68%), Gaps = 11/227 (4%)
Query: 8 LEAMLTFQVI-WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNA 66
LE ++ F+ WPS+S SAKDLVRKML DP++RI++ +VL+HPW++E GEA D P+D+A
Sbjct: 294 LEGVIDFESEPWPSVSNSAKDLVRKMLTQDPRRRITSAQVLDHPWMREGGEASDKPIDSA 353
Query: 67 VLNR-------NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GL 119
VL+R N L+Q + K++ LSEEEI GLK MF MDTD SGTIT EELK GL
Sbjct: 354 VLSRMKQFRAMNKLKQL--ALKVIAESLSEEEIKGLKSMFANMDTDKSGTITYEELKSGL 411
Query: 120 VKQGTRLSEQQVKQLMEAADADGNGIIDYDEFITASQQLCTRTD*TEKNMFILPSNTSIK 179
+ G++LSE +V+QLM+AAD DGNG IDY EFITA+ + + S
Sbjct: 412 ARLGSKLSEVEVQQLMDAADVDGNGTIDYLEFITATMHRHKLESYEHQAFQYFDKDNSGF 471
Query: 180 IT-TELEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRK 225
IT ELE A+ EY M D IK+IISEVD+DNDGRINYDEF AMMR+
Sbjct: 472 ITKDELESAMKEYGMGDEATIKDIISEVDSDNDGRINYDEFCAMMRR 518
>CDPK_SOYBN (P28583) Calcium-dependent protein kinase SK5 (EC
2.7.1.-) (CDPK)
Length = 508
Score = 191 bits (484), Expect = 2e-48
Identities = 111/220 (50%), Positives = 147/220 (66%), Gaps = 13/220 (5%)
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T 77
WPSIS SAKDL+RKML+ +PK R++A+EVL HPWI +D APD PLD+AVL+R L+Q +
Sbjct: 258 WPSISDSAKDLIRKMLDQNPKTRLTAHEVLRHPWIVDDNIAPDKPLDSAVLSR--LKQFS 315
Query: 78 DSRKL-------L*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
KL + LSEEEI GLK++FK +DTDNSGTIT +ELK GL + G+ L E +
Sbjct: 316 AMNKLKKMALRVIAERLSEEEIGGLKELFKMIDTDNSGTITFDELKDGLKRVGSELMESE 375
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQL--CTRTD*TEKNMFILPSNTSIKIT-TELEQA 187
+K LM+AAD D +G IDY EFI A+ L R + + S IT E++QA
Sbjct: 376 IKDLMDAADIDKSGTIDYGEFIAATVHLNKLEREENLVSAFSYFDKDGSGYITLDEIQQA 435
Query: 188 LHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGN 227
++ + D +I ++I E+D DNDG+I+Y EF AMMRKGN
Sbjct: 436 CKDFGL-DDIHIDDMIKEIDQDNDGQIDYGEFAAMMRKGN 474
>CDP1_ARATH (Q06850) Calcium-dependent protein kinase, isoform AK1
(EC 2.7.1.-) (CDPK)
Length = 610
Score = 188 bits (477), Expect = 1e-47
Identities = 111/220 (50%), Positives = 145/220 (65%), Gaps = 13/220 (5%)
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T 77
WPSIS SAKDLVRKML DPK+R++A++VL HPW++ DG APD PLD+AVL+R ++Q +
Sbjct: 374 WPSISESAKDLVRKMLVRDPKKRLTAHQVLCHPWVQVDGVAPDKPLDSAVLSR--MKQFS 431
Query: 78 DSRK-------LL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
K ++ LSEEEI GLK+MF +D D SG IT EELK GL + G L E +
Sbjct: 432 AMNKFKKMALRVIAESLSEEEIAGLKEMFNMIDADKSGQITFEELKAGLKRVGANLKESE 491
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQL--CTRTD*TEKNMFILPSNTSIKIT-TELEQA 187
+ LM+AAD D +G IDY EFI A+ L R D + S IT EL+QA
Sbjct: 492 ILDLMQAADVDNSGTIDYKEFIAATLHLNKIEREDHLFAAFTYFDKDGSGYITPDELQQA 551
Query: 188 LHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGN 227
E+ + D R I+E++ +VD DNDGRI+Y+EFVAMM+KG+
Sbjct: 552 CEEFGVEDVR-IEELMRDVDQDNDGRIDYNEFVAMMQKGS 590
>CDP3_ORYSA (P53684) Calcium-dependent protein kinase, isoform 11
(EC 2.7.1.-) (CDPK 11)
Length = 542
Score = 158 bits (400), Expect = 1e-38
Identities = 101/224 (45%), Positives = 138/224 (61%), Gaps = 21/224 (9%)
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNRNSLEQ*T 77
WP IS SAKDL+RKML+ P +R+ A+EVL HPWI E+G A D LD +V++R L+Q +
Sbjct: 303 WPKISDSAKDLIRKMLSHCPSERLKAHEVLRHPWICENGVATDQALDPSVISR--LKQFS 360
Query: 78 DSRKL-------L*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
KL + LSEEEI GL++MFK +DT N G IT EL+ GL + G + +
Sbjct: 361 AMNKLKKLALRVIAERLSEEEIAGLREMFKAVDTKNRGVITFGELREGLRRFGAEFKDTE 420
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQLCTRTD*TEKNMFILPSNT------SIKITTE- 183
+ +MEAA D N I Y+EFI A+ L + E+ +L + T S IT +
Sbjct: 421 IGDIMEAAHNDNNVTIHYEEFIAATLPL----NKIEREEHLLAAFTYFDKDGSGYITVDK 476
Query: 184 LEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGN 227
L++A E+NM D ++EIISEVD +NDG+I+Y EFVAMM+ N
Sbjct: 477 LQRACGEHNMEDS-LLEEIISEVDQNNDGQIDYAEFVAMMQGSN 519
>CDP1_ORYSA (P53682) Calcium-dependent protein kinase, isoform 1 (EC
2.7.1.-) (CDPK 1)
Length = 534
Score = 139 bits (349), Expect = 8e-33
Identities = 93/224 (41%), Positives = 136/224 (60%), Gaps = 23/224 (10%)
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLDNAVLNR------- 70
WP IS SAKDL++KML P +R+ A+EVL HPWI ++G A + LD +VL R
Sbjct: 297 WPRISDSAKDLIKKMLCPYPLERLKAHEVLKHPWICDNGVATNRALDPSVLPRLKQFSAM 356
Query: 71 NSLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
N L++ S +++ LSEEEI+GL++MFK MDT N +T ELK GL + + + +
Sbjct: 357 NRLKK--LSLQIIAERLSEEEIVGLREMFKAMDTKNRSVVTFGELK-GLKRYSSVFKDTE 413
Query: 131 VKQLMEAADADGNGIIDYDEFITASQQLCTRTD*TEKNMFILPSNT------SIKITTE- 183
+ LMEAAD D I+++EFI A+ L + E+ ++ + T S IT +
Sbjct: 414 INDLMEAAD-DTTSTINWEEFIAAAVSL----NKIEREKHLMAAFTYFDKDGSGFITVDK 468
Query: 184 LEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGN 227
L++A E NM D +++E+I EVD +NDG+I+Y EFV MM+ N
Sbjct: 469 LQKACMERNMED-TFLEEMILEVDQNNDGQIDYAEFVTMMQSNN 511
>CDP3_PLAF7 (Q9NJU9) Calcium-dependent protein kinase 3 (EC
2.7.1.37) (PfCDPK3)
Length = 562
Score = 99.4 bits (246), Expect = 7e-21
Identities = 76/223 (34%), Positives = 115/223 (51%), Gaps = 18/223 (8%)
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEA--PDTPLDNAVL-NRNSLE 74
W +IS AKDL+++ L D +RI A E L HPW K+ A D +D VL N +
Sbjct: 339 WNNISEEAKDLIKRCLTMDADKRICASEALQHPWFKKKKYAFNMDMKMDIHVLENFKNYG 398
Query: 75 Q*TDSRKLL*SCLSEE----EIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
+KL + ++++ ++ LK F +D D G IT E+LK GL K G +L
Sbjct: 399 LLLKFQKLAMTIIAQQSNDYDVEKLKSTFLVLDEDGKGYITKEQLKKGLEKDGLKL-PYN 457
Query: 131 VKQLMEAADADGNGIIDYDEFITAS---QQLCTRTD*TEKNMFILPSNTSIKITTELEQA 187
L++ D+DG+G IDY EFI A+ +QL + +F + ++ I T EL
Sbjct: 458 FDLLLDQIDSDGSGKIDYTEFIAAALDRKQLSKKLIYCAFRVFDVDNDGEI-TTAELAHI 516
Query: 188 LHEYN------MHDGRYIKEIISEVDADNDGRINYDEFVAMMR 224
L+ N D +K +I +VD +NDG+I++ EF MM+
Sbjct: 517 LYNGNKKGNITQRDVNRVKRMIRDVDKNNDGKIDFHEFSEMMK 559
>CDP2_PLAFK (O15865) Calcium-dependent protein kinase 2 (EC
2.7.1.37) (PfCDPK2)
Length = 512
Score = 91.3 bits (225), Expect = 2e-18
Identities = 66/224 (29%), Positives = 111/224 (49%), Gaps = 20/224 (8%)
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIK------EDGEAPDTPLDNAVLNRN 71
W SIS AK+L+ K+L +P +R + E LNHPWI E E T L N +
Sbjct: 291 WGSISSDAKNLITKLLTYNPNERCTIEEALNHPWITQMTKSHEHVELSSTLLKNLKNFKK 350
Query: 72 SLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQV 131
E + ++ L + EI L+ +F +D DNSGT++ +E+ GL K G + +
Sbjct: 351 ENELKKIALTIIAKHLCDVEINNLRNIFIALDVDNSGTLSSQEILDGLKKIGYQKIPPDI 410
Query: 132 KQLMEAADADGNGIIDYDEFITASQQLCTRTD*TEKNMFILP-------SNTSIKITTEL 184
Q++ D++ +G I Y +F+ A+ T +K + ++P N I + EL
Sbjct: 411 HQVLRDIDSNASGQIHYTDFLAATIDKQTY---LKKEVCLIPFKFFDIDGNGKISV-EEL 466
Query: 185 EQALHEYNMHD---GRYIKEIISEVDADNDGRINYDEFVAMMRK 225
++ ++ + + I ++ EVD + DG I++ EF+ MM K
Sbjct: 467 KRIFGRDDIENPLIDKAIDSLLQEVDLNGDGEIDFHEFMLMMSK 510
>CDP2_PLAF7 (Q8ICR0) Calcium-dependent protein kinase 2 (EC
2.7.1.37) (PfCDPK2)
Length = 508
Score = 82.4 bits (202), Expect = 9e-16
Identities = 61/214 (28%), Positives = 104/214 (48%), Gaps = 20/214 (9%)
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIK------EDGEAPDTPLDNAVLNRN 71
W SIS AK+L+ K+L +P +R + E LNHPWI E E T L N +
Sbjct: 291 WGSISSDAKNLITKLLTYNPNERCTIEEALNHPWITQMTKSHEHVELSSTLLKNLKNFKK 350
Query: 72 SLEQ*TDSRKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQV 131
E + ++ L + EI L+ +F +D DNSGT++ +E+ GL K G + +
Sbjct: 351 ENELKKIALTIIAKHLCDVEINNLRNIFIALDVDNSGTLSSQEILDGLKKIGYQKIPPDI 410
Query: 132 KQLMEAADADGNGIIDYDEFITASQQLCTRTD*TEKNMFILP-------SNTSIKITTEL 184
Q++ D++ +G I Y +F+ A+ T +K + ++P N I + EL
Sbjct: 411 HQVLRDIDSNASGQIHYTDFLAATIDKQTY---LKKEVCLIPFKFFDIDGNGKISV-EEL 466
Query: 185 EQALHEYNMHD---GRYIKEIISEVDADNDGRIN 215
++ ++ + + I ++ EVD + DG +N
Sbjct: 467 KRIFGRDDIENPLIDKAIDSLLQEVDLNGDGEVN 500
>CDP4_PLAF7 (Q8IBS5) Calcium-dependent protein kinase 4 (EC
2.7.1.37)
Length = 527
Score = 80.9 bits (198), Expect = 3e-15
Identities = 70/228 (30%), Positives = 108/228 (46%), Gaps = 23/228 (10%)
Query: 21 ISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEAPDTPLD--NAVLNRNSLEQ*TD 78
IS AKDL++KML RISA + L H WIK + +D + L+ ++ Q
Sbjct: 296 ISDKAKDLIKKMLMYTSAVRISARDALEHEWIKMMTSKDNLNIDIPSLELSIANIRQFQS 355
Query: 79 SRKLL*SCL--------SEEEIMGLKQMFKGMDTDNSGTITIEELK*G----LVKQGTRL 126
++KL + L + +E L ++FK MD + G + EL G L +G
Sbjct: 356 TQKLAQAALLYMGSKLTTIDETKELTKIFKKMDKNGDGQLDRNELIIGYKELLKLKGEDT 415
Query: 127 S-------EQQVKQLMEAADADGNGIIDYDEFITAS--QQLCTRTD*TEKNMFILPSNTS 177
S E +V Q++ + D D NG I+Y EF+T S ++L T+ EK + + S
Sbjct: 416 SDLDNAAIEYEVDQILNSIDLDQNGYIEYSEFLTVSIDRKLLLSTERLEKAFKLFDKDGS 475
Query: 178 IKITTELEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRK 225
KI+ L + K ++ EVD +NDG I++ EF M+ K
Sbjct: 476 GKISANELAQLFGLSDVSSECWKTVLKEVDQNNDGEIDFKEFRDMLVK 523
Score = 31.2 bits (69), Expect = 2.3
Identities = 29/82 (35%), Positives = 42/82 (50%), Gaps = 12/82 (14%)
Query: 80 RKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*--GLVKQGTRLSEQQVKQLMEA 137
RKLL LS E L++ FK D D SG I+ EL GL + +S + K +++
Sbjct: 454 RKLL---LSTER---LEKAFKLFDKDGSGKISANELAQLFGL----SDVSSECWKTVLKE 503
Query: 138 ADADGNGIIDYDEFITASQQLC 159
D + +G ID+ EF +LC
Sbjct: 504 VDQNNDGEIDFKEFRDMLVKLC 525
>CALM_CHLRE (P04352) Calmodulin (CaM)
Length = 162
Score = 80.9 bits (198), Expect = 3e-15
Identities = 53/161 (32%), Positives = 78/161 (47%), Gaps = 17/161 (10%)
Query: 87 LSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGII 146
L+EE+I K+ F D D GTIT +EL + G +E +++ ++ DADGNG I
Sbjct: 7 LTEEQIAEFKEAFALFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMISEVDADGNGTI 66
Query: 147 DYDEFITASQQLCTRTD*TEK-----NMFILPSNTSIK------ITTELEQALHEYNMHD 195
D+ EF+ + TD ++ +F N I + T L + L E
Sbjct: 67 DFPEFLMLMARKMKETDHEDELREAFKVFDKDGNGFISAAELRHVMTNLGEKLSE----- 121
Query: 196 GRYIKEIISEVDADNDGRINYDEFVAMMRKGNPEAHTKKRH 236
+ E+I E D D DG++NY+EFV MM G + KK H
Sbjct: 122 -EEVDEMIREADVDGDGQVNYEEFVRMMTSGATDDKDKKGH 161
>CALM_HALOK (Q95NI4) Calmodulin (CaM)
Length = 148
Score = 79.7 bits (195), Expect = 6e-15
Identities = 52/148 (35%), Positives = 75/148 (50%), Gaps = 17/148 (11%)
Query: 87 LSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGII 146
LSEE+I K+ F D D GTIT +EL + G +E +++ ++ DADGNG I
Sbjct: 4 LSEEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTI 63
Query: 147 DYDEFITASQQLCTRTD*TEK-----NMFILPSNTSIK------ITTELEQALHEYNMHD 195
D+ EF+T + TD E+ +F N I + T L + L +
Sbjct: 64 DFPEFLTMMARKMKETDSEEEIREAFRVFDKDGNGFISAAELRHVMTNLGEKLTD----- 118
Query: 196 GRYIKEIISEVDADNDGRINYDEFVAMM 223
+ E+I E D D DG++NY+EFVAMM
Sbjct: 119 -EEVDEMIREADIDGDGQVNYEEFVAMM 145
Score = 34.7 bits (78), Expect = 0.21
Identities = 30/112 (26%), Positives = 48/112 (42%), Gaps = 35/112 (31%)
Query: 126 LSEQQVKQLMEAA---DADGNGIIDYDEFITASQQLCTRTD*TEKNMFILPSNTSIKITT 182
LSE+Q+ + EA D DG+G I E T + L T
Sbjct: 4 LSEEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNP-------------------T 44
Query: 183 ELEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGNPEAHTKK 234
E E ++++I+EVDAD +G I++ EF+ MM + E +++
Sbjct: 45 EAE-------------LQDMINEVDADGNGTIDFPEFLTMMARKMKETDSEE 83
>CALM_PAXIN (Q8X187) Calmodulin (CaM)
Length = 148
Score = 79.0 bits (193), Expect = 1e-14
Identities = 53/148 (35%), Positives = 74/148 (49%), Gaps = 17/148 (11%)
Query: 87 LSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGII 146
LSEE+I K+ F D D GTIT +EL + G +E +++ ++ DADGNG I
Sbjct: 4 LSEEQISEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEGELQDMINEVDADGNGTI 63
Query: 147 DYDEFITASQQLCTRTD*TEK-----NMFILPSNTSIK------ITTELEQALHEYNMHD 195
D+ EF+T + TD E+ +F N I + T L + L +
Sbjct: 64 DFPEFLTMMARKMRDTDSEEEIKEAFKVFDKDGNGYISAAELRHVMTNLGEKLTDTE--- 120
Query: 196 GRYIKEIISEVDADNDGRINYDEFVAMM 223
+ E+I E D D DG+INYDEFV MM
Sbjct: 121 ---VDEMIREADVDGDGQINYDEFVKMM 145
Score = 33.9 bits (76), Expect = 0.36
Identities = 28/104 (26%), Positives = 45/104 (42%), Gaps = 35/104 (33%)
Query: 125 RLSEQQVKQLMEAA---DADGNGIIDYDEFITASQQLCTRTD*TEKNMFILPSNTSIKIT 181
+LSE+Q+ + EA D DG+G I E T + L
Sbjct: 3 QLSEEQISEFKEAFSLFDKDGDGTITTKELGTVMRSL----------------------- 39
Query: 182 TELEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRK 225
N +G ++++I+EVDAD +G I++ EF+ MM +
Sbjct: 40 --------GQNPTEGE-LQDMINEVDADGNGTIDFPEFLTMMAR 74
>CLM_PLEOS (O94739) Calmodulin (CaM)
Length = 148
Score = 77.8 bits (190), Expect = 2e-14
Identities = 52/148 (35%), Positives = 74/148 (49%), Gaps = 17/148 (11%)
Query: 87 LSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGII 146
LSEE+I K+ F D D GTIT +EL + G +E +++ ++ DADGNG I
Sbjct: 4 LSEEQISEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTI 63
Query: 147 DYDEFITASQQLCTRTD*TEK-----NMFILPSNTSIK------ITTELEQALHEYNMHD 195
D+ EF+T + TD E+ +F N I + T L + L +
Sbjct: 64 DFPEFLTMMARKMRDTDSEEEIKEAFKVFDKDGNGYISAAELRHVMTNLGEKLTD----- 118
Query: 196 GRYIKEIISEVDADNDGRINYDEFVAMM 223
+ E+I E D D DG+INY+EFV MM
Sbjct: 119 -NEVDEMIREADVDGDGQINYEEFVKMM 145
Score = 34.7 bits (78), Expect = 0.21
Identities = 29/104 (27%), Positives = 45/104 (42%), Gaps = 35/104 (33%)
Query: 125 RLSEQQVKQLMEAA---DADGNGIIDYDEFITASQQLCTRTD*TEKNMFILPSNTSIKIT 181
+LSE+Q+ + EA D DG+G I E T + L
Sbjct: 3 QLSEEQISEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNP------------------- 43
Query: 182 TELEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRK 225
TE E ++++I+EVDAD +G I++ EF+ MM +
Sbjct: 44 TEAE-------------LQDMINEVDADGNGTIDFPEFLTMMAR 74
>CDP4_PLAYO (Q7RJG2) Calcium-dependent protein kinase 4 (EC
2.7.1.37)
Length = 527
Score = 77.4 bits (189), Expect = 3e-14
Identities = 69/231 (29%), Positives = 109/231 (46%), Gaps = 29/231 (12%)
Query: 21 ISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIK----EDGEAPDTPLDNAVLNRNSLEQ* 76
IS AKDL++KML RISA + L H WI+ +D D P + L+ +++Q
Sbjct: 296 ISDKAKDLIKKMLMYTSAVRISARDALEHEWIRLMTSKDNVNIDIP--SLELSITNIKQF 353
Query: 77 TDSRKLL*SCL--------SEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLS- 127
++KL + L + +E L ++FK MD + G + EL G K+ +L
Sbjct: 354 QSTQKLAQAALLYMGSKLTTIDETKELTKIFKKMDKNGDGQLDRNELIIG-YKELLKLKG 412
Query: 128 -----------EQQVKQLMEAADADGNGIIDYDEFITAS--QQLCTRTD*TEKNMFILPS 174
E +V Q++ + D D NG I+Y EF+T + ++L T+ EK +
Sbjct: 413 DDTTDLDNAAIEVEVDQILSSIDLDQNGYIEYSEFLTVAIDRKLLLSTERLEKAFKLFDK 472
Query: 175 NTSIKITTELEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRK 225
+ S KI+ L K ++ EVD +NDG I++ EF M+ K
Sbjct: 473 DGSGKISANELAQLFGLGDVSSDCWKTVLKEVDQNNDGEIDFKEFRDMLIK 523
Score = 30.4 bits (67), Expect = 3.9
Identities = 29/80 (36%), Positives = 40/80 (49%), Gaps = 8/80 (10%)
Query: 80 RKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAAD 139
RKLL LS E L++ FK D D SG I+ EL L G +S K +++ D
Sbjct: 454 RKLL---LSTER---LEKAFKLFDKDGSGKISANELA-QLFGLGD-VSSDCWKTVLKEVD 505
Query: 140 ADGNGIIDYDEFITASQQLC 159
+ +G ID+ EF +LC
Sbjct: 506 QNNDGEIDFKEFRDMLIKLC 525
>CDP4_PLABA (P62345) Calcium-dependent protein kinase 4 (EC
2.7.1.37) (PbCDPK4)
Length = 527
Score = 77.4 bits (189), Expect = 3e-14
Identities = 69/231 (29%), Positives = 109/231 (46%), Gaps = 29/231 (12%)
Query: 21 ISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIK----EDGEAPDTPLDNAVLNRNSLEQ* 76
IS AKDL++KML RISA + L H WI+ +D D P + L+ +++Q
Sbjct: 296 ISDKAKDLIKKMLMYTSAVRISARDALEHEWIRLMTSKDNVNIDIP--SLELSITNIKQF 353
Query: 77 TDSRKLL*SCL--------SEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLS- 127
++KL + L + +E L ++FK MD + G + EL G K+ +L
Sbjct: 354 QSTQKLAQAALLYMGSKLTTIDETKELTKIFKKMDKNGDGQLDRNELIIG-YKELLKLKG 412
Query: 128 -----------EQQVKQLMEAADADGNGIIDYDEFITAS--QQLCTRTD*TEKNMFILPS 174
E +V Q++ + D D NG I+Y EF+T + ++L T+ EK +
Sbjct: 413 DDTTDLDNAAIEVEVDQILSSIDLDQNGYIEYSEFLTVAIDRKLLLSTERLEKAFKLFDK 472
Query: 175 NTSIKITTELEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRK 225
+ S KI+ L K ++ EVD +NDG I++ EF M+ K
Sbjct: 473 DGSGKISANELAQLFGLGDVSSDCWKTVLKEVDQNNDGEIDFKEFRDMLIK 523
Score = 30.4 bits (67), Expect = 3.9
Identities = 29/80 (36%), Positives = 40/80 (49%), Gaps = 8/80 (10%)
Query: 80 RKLL*SCLSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAAD 139
RKLL LS E L++ FK D D SG I+ EL L G +S K +++ D
Sbjct: 454 RKLL---LSTER---LEKAFKLFDKDGSGKISANELA-QLFGLGD-VSSDCWKTVLKEVD 505
Query: 140 ADGNGIIDYDEFITASQQLC 159
+ +G ID+ EF +LC
Sbjct: 506 QNNDGEIDFKEFRDMLIKLC 525
>CALM_PLECO (P11120) Calmodulin (CaM)
Length = 148
Score = 77.4 bits (189), Expect = 3e-14
Identities = 52/148 (35%), Positives = 74/148 (49%), Gaps = 17/148 (11%)
Query: 87 LSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGII 146
LSEE+I K+ F D D GTIT +EL + G +E +++ ++ DADGNG I
Sbjct: 4 LSEEQISEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTI 63
Query: 147 DYDEFITASQQLCTRTD*TEK-----NMFILPSNTSIK------ITTELEQALHEYNMHD 195
D+ EF+T + TD E+ +F N I + T L + L +
Sbjct: 64 DFPEFLTMMARKMRDTDSEEEIKEAFKVFDKDGNGYISAAELRHVMTNLGEKLTD----- 118
Query: 196 GRYIKEIISEVDADNDGRINYDEFVAMM 223
+ E+I E D D DG+INY+EFV MM
Sbjct: 119 -NEVDEMIREADIDGDGQINYEEFVKMM 145
Score = 34.7 bits (78), Expect = 0.21
Identities = 29/104 (27%), Positives = 45/104 (42%), Gaps = 35/104 (33%)
Query: 125 RLSEQQVKQLMEAA---DADGNGIIDYDEFITASQQLCTRTD*TEKNMFILPSNTSIKIT 181
+LSE+Q+ + EA D DG+G I E T + L
Sbjct: 3 QLSEEQISEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNP------------------- 43
Query: 182 TELEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRK 225
TE E ++++I+EVDAD +G I++ EF+ MM +
Sbjct: 44 TEAE-------------LQDMINEVDADGNGTIDFPEFLTMMAR 74
>CDP3_PLAYO (Q7RAV5) Calcium-dependent protein kinase 3 (EC
2.7.1.37)
Length = 538
Score = 77.0 bits (188), Expect = 4e-14
Identities = 51/145 (35%), Positives = 79/145 (54%), Gaps = 8/145 (5%)
Query: 18 WPSISPSAKDLVRKMLNSDPKQRISAYEVLNHPWIKEDGEA--PDTPLDNAVL-NRNSLE 74
W +IS AKDL+++ L D +RI+A E L HPW K+ + D +D VL N +
Sbjct: 333 WNNISDEAKDLIKRCLTIDSGKRINASEALKHPWFKKKKGSFNLDVKMDIHVLENFKNYA 392
Query: 75 Q*TDSRKLL*SCLSEE----EIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQ 130
+KL + ++++ ++ LK +F +D D G IT +LK GL G +L Q
Sbjct: 393 LLLKLQKLAMTIIAQQSNDYDLQQLKAVFLYLDEDGKGNITKNQLKKGLENSGLKL-PQN 451
Query: 131 VKQLMEAADADGNGIIDYDEFITAS 155
L++ D+DG+G IDY EF+ A+
Sbjct: 452 FDVLLDQIDSDGSGRIDYTEFLAAA 476
>CALB_HALRO (O96081) Calmodulin B (CaM B)
Length = 148
Score = 77.0 bits (188), Expect = 4e-14
Identities = 50/148 (33%), Positives = 74/148 (49%), Gaps = 17/148 (11%)
Query: 87 LSEEEIMGLKQMFKGMDTDNSGTITIEELK*GLVKQGTRLSEQQVKQLMEAADADGNGII 146
L+EE+I K+ F D D GTIT +EL + G +E +++ ++ DADGNG I
Sbjct: 4 LTEEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTI 63
Query: 147 DYDEFITASQQLCTRTD*TEK-----NMFILPSNTSIK------ITTELEQALHEYNMHD 195
D+ EF+T + TD E+ +F N I + T L + L +
Sbjct: 64 DFPEFLTMMARKMKETDSEEEIREAFRVFDKDGNGFISAAELRHVMTNLGEKLTD----- 118
Query: 196 GRYIKEIISEVDADNDGRINYDEFVAMM 223
+ E+I E D D DG++NY+EFV MM
Sbjct: 119 -EEVDEMIREADIDGDGQVNYEEFVTMM 145
Score = 33.9 bits (76), Expect = 0.36
Identities = 29/113 (25%), Positives = 49/113 (42%), Gaps = 35/113 (30%)
Query: 125 RLSEQQVKQLMEAA---DADGNGIIDYDEFITASQQLCTRTD*TEKNMFILPSNTSIKIT 181
+L+E+Q+ + EA D DG+G I E T + L
Sbjct: 3 QLTEEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNP------------------- 43
Query: 182 TELEQALHEYNMHDGRYIKEIISEVDADNDGRINYDEFVAMMRKGNPEAHTKK 234
TE E ++++I+EVDAD +G I++ EF+ MM + E +++
Sbjct: 44 TEAE-------------LQDMINEVDADGNGTIDFPEFLTMMARKMKETDSEE 83
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.137 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,918,308
Number of Sequences: 164201
Number of extensions: 1048257
Number of successful extensions: 5521
Number of sequences better than 10.0: 996
Number of HSP's better than 10.0 without gapping: 787
Number of HSP's successfully gapped in prelim test: 213
Number of HSP's that attempted gapping in prelim test: 3213
Number of HSP's gapped (non-prelim): 1664
length of query: 240
length of database: 59,974,054
effective HSP length: 107
effective length of query: 133
effective length of database: 42,404,547
effective search space: 5639804751
effective search space used: 5639804751
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 64 (29.3 bits)
Medicago: description of AC146941.2


