

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146866.2 + phase: 0
(504 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
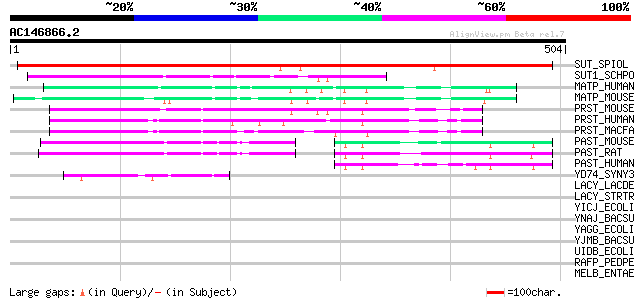
Score E
Sequences producing significant alignments: (bits) Value
SUT_SPIOL (Q03411) Sucrose transport protein (Sucrose permease) ... 515 e-145
SUT1_SCHPO (O14091) General alpha-glucoside permease 144 4e-34
MATP_HUMAN (Q9UMX9) Membrane-associated transporter protein (AIM... 120 6e-27
MATP_MOUSE (P58355) Membrane-associated transporter protein (AIM... 120 1e-26
PRST_MOUSE (Q8K0H7) Prostein 112 2e-24
PRST_HUMAN (Q96JT2) Prostein 110 8e-24
PRST_MACFA (Q95KI5) Prostein (QmoA-10594/QtrA-11310) 110 1e-23
PAST_MOUSE (Q8BIV7) Proton-associated sugar transporter A (PAST-... 110 1e-23
PAST_RAT (Q8K4S3) Proton-associated sugar transporter A (PAST-A) 109 1e-23
PAST_HUMAN (Q9Y2W3) Proton-associated sugar transporter A (PAST-... 54 1e-06
YD74_SYNY3 (P74168) Hypothetical symporter sll1374 48 7e-05
LACY_LACDE (P22733) Lactose permease (Lactose-proton symporter) ... 42 0.005
LACY_STRTR (P23936) Lactose permease (Lactose-proton symporter) ... 40 0.011
YICJ_ECOLI (P31435) Hypothetical symporter yicJ 40 0.014
YNAJ_BACSU (P94488) Hypothetical symporter ynaJ 40 0.018
YAGG_ECOLI (P75683) Hypothetical symporter yagG 38 0.053
YJMB_BACSU (O34961) Hypothetical symporter yjmB 38 0.070
UIDB_ECOLI (P30868) Glucuronide carrier protein (Glucuronide per... 37 0.16
RAFP_PEDPE (P43466) Raffinose carrier protein (Raffinose permease) 37 0.16
MELB_ENTAE (O07366) Melibiose carrier protein (Thiomethylgalacto... 37 0.16
>SUT_SPIOL (Q03411) Sucrose transport protein (Sucrose permease)
(Sucrose-proton symporter)
Length = 525
Score = 515 bits (1326), Expect = e-145
Identities = 270/503 (53%), Positives = 333/503 (65%), Gaps = 17/503 (3%)
Query: 8 NPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLG 67
N ++ S+ P P + L++L VASVA+G+QFGWALQLSLLTPYVQ LG
Sbjct: 8 NGENNKIAGSSLHLEKNPTTPPEAEATLKKLGLVASVAAGVQFGWALQLSLLTPYVQLLG 67
Query: 68 IPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAA 127
IPH WA+ IWLCGP+SG+ VQPLVG+ SDRC+SRFGRRRPFI GAA + VAV +IG+AA
Sbjct: 68 IPHTWAAYIWLCGPISGMIVQPLVGYYSDRCTSRFGRRRPFIAAGAALVAVAVGLIGFAA 127
Query: 128 DIGYLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANA 187
DIG GD +P AI VFV+GFWILDVANN QGPCRALLAD+ +TR ANA
Sbjct: 128 DIGAASGDPTGNVAKPRAIAVFVVGFWILDVANNTLQGPCRALLADMAAGSQTKTRYANA 187
Query: 188 YFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTY--L 245
+FS FMA+GNI GYA GSYS Y +F FT T AC + CANLKS FF+ + ++V T L
Sbjct: 188 FFSFFMALGNIGGYAAGSYSRLYTVFPFTKTAACDVYCANLKSCFFISITLLIVLTILAL 247
Query: 246 SIVSAHEVPLSSSGAGE--------SGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWI 297
S+V ++ + E SG A F +L G K P+ I+L VTAL WI
Sbjct: 248 SVVKERQITIDEIQEEEDLKNRNNSSGCARLPFFGQLIGALKDLPKPMLILLLVTALNWI 307
Query: 298 GWFPFNLFDTDWMGREIYGGDPEGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRK- 356
WFPF LFDTDWMG+E+YGG G +YD GV GALGL++NSVVL V SL +E L R
Sbjct: 308 AWFPFLLFDTDWMGKEVYGGTVGEGKLYDQGVHAGALGLMINSVVLGVMSLSIEGLARMV 367
Query: 357 RGAGFVWGISNIFMAICFIAMLVLTYAA------NSIGYVSKGQPPPTGIVIAALAIFTI 410
GA +WGI NI +A+C +++T +A + I + PPP G+ ALAIF +
Sbjct: 368 GGAKRLWGIVNIILAVCLAMTVLVTKSAEHFRDSHHIMGSAVPPPPPAGVKGGALAIFAV 427
Query: 411 LGFPMAITYSVPYALISTHIEPLGLGQGLSMGVLNLAIVVPQIVVSLGSGPWDQLFGGGN 470
LG P+AIT+S+P+AL S G GQGLS+GVLNLAIVVPQ+ VS+ SGPWD +FGGGN
Sbjct: 428 LGIPLAITFSIPFALASIFSASSGSGQGLSLGVLNLAIVVPQMFVSVTSGPWDAMFGGGN 487
Query: 471 SPAFAVAAVAALLSGLLALLAIP 493
PAF V AVAA S +L+ +P
Sbjct: 488 LPAFVVGAVAATASAVLSFTLLP 510
>SUT1_SCHPO (O14091) General alpha-glucoside permease
Length = 553
Score = 144 bits (364), Expect = 4e-34
Identities = 99/332 (29%), Positives = 165/332 (48%), Gaps = 21/332 (6%)
Query: 17 STSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASII 76
S+ S P + P L+ + G+Q W+++L TPY+ LG+ +W SII
Sbjct: 15 SSENEASSPFKESIPSRSSLYLIALTVSLLGVQLTWSVELGYGTPYLFSLGLRKEWTSII 74
Query: 77 WLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIG-YLIGD 135
W+ GP++G+ +QP+ G LSDR +SR GRRRPF+L + ++ ++G+A DI ++ +
Sbjct: 75 WIAGPLTGILIQPIAGILSDRVNSRIGRRRPFMLCASLLGTFSLFLMGWAPDICLFIFSN 134
Query: 136 DITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAV 195
++ + IV+ I ++LDVA NV R+L+ D +D + AN++ + V
Sbjct: 135 EVLM--KRVTIVLATISIYLLDVAVNVVMASTRSLIVDSVRSDQQHE--ANSWAGRMIGV 190
Query: 196 GNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPL 255
GN+LGY G Y Y+IF+F + C + L V + T ++S V
Sbjct: 191 GNVLGYLLG-YLPLYRIFSFLNFTQLQVFCVLASISLVLTVT--ITTIFVSERRFPPVEH 247
Query: 256 SSSGAGESGSAEEAFMWELFGTFK--YFSMPVWI--VLSVTALTWIGWFPFNLFDTDWMG 311
S AGE ++E F T + ++P + + V + GWFPF + T ++G
Sbjct: 248 EKSVAGE--------IFEFFTTMRQSITALPFTLKRICFVQFFAYFGWFPFLFYITTYVG 299
Query: 312 REIYGGDPEG-GLIYDTGVRMGALGLLLNSVV 342
P+G +D R G+ LLL +++
Sbjct: 300 ILYLRHAPKGHEEDWDMATRQGSFALLLFAII 331
Score = 35.8 bits (81), Expect = 0.27
Identities = 21/56 (37%), Positives = 31/56 (54%), Gaps = 2/56 (3%)
Query: 404 ALAIFTILGFPMAITYSVPYALISTHIEPLGL--GQGLSMGVLNLAIVVPQIVVSL 457
A A+ I G A T +PY+L S+ I LGL G +GV N+ I PQ++ ++
Sbjct: 451 AQAMVAICGLSWACTLWIPYSLFSSEIGKLGLRESSGKMIGVHNVFISAPQVLSTI 506
>MATP_HUMAN (Q9UMX9) Membrane-associated transporter protein (AIM-1
protein) (Melanoma antigen AIM1)
Length = 530
Score = 120 bits (302), Expect = 6e-27
Identities = 130/494 (26%), Positives = 199/494 (39%), Gaps = 90/494 (18%)
Query: 31 PRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPL 90
P+ P +L+ + G +F +A++ + +TP + +G+P SI+W P+ G +QP+
Sbjct: 28 PKRPTSRLIMHSMAMFGREFCYAVEAAYVTPVLLSVGLPSSLYSIVWFLSPILGFLLQPV 87
Query: 91 VGHLSDRCSSRFGRRRPFIL-VGAASIVVAVVIIGYAADIGYLIGDDITQNYRPFAIVVF 149
VG SD C SR+GRRRP+IL +G +V + + A + LI + + +AI V
Sbjct: 88 VGSASDHCRSRWGRRRPYILTLGVMMLVGMALYLNGATVVAALIAN--PRRKLVWAISVT 145
Query: 150 VIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGYATGSYSGW 209
+IG + D A + GP +A L D+ + + + Y +LF G LGY G+ W
Sbjct: 146 MIGVVLFDFAADFIDGPIKAYLFDVCSHQDKEKGL--HYHALFTGFGGALGYLLGAID-W 202
Query: 210 YKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEV-------------PLS 256
+ L SA L + F V +S EV PLS
Sbjct: 203 AHLELGRLL-GTEFQVMFFFSALVLTLCFTVHLCSISEAPLTEVAKGIPPQQTPQDPPLS 261
Query: 257 SSGAGESGSAEE--------------------AFMWELFGTFKYF------SMPVWIVLS 290
S G E GS E+ A T K P + L
Sbjct: 262 SDGMYEYGSIEKVKNGYVNPELAMQGAKNKNHAEQTRRAMTLKSLLRALVNMPPHYRYLC 321
Query: 291 VTALTWIGWFPF---NLFDTDWMGREIYGGDPEGG------LIYDTGVRMGALGLLLNSV 341
++ L IGW F LF TD+MG+ +Y GDP LIY+ GV +G G +NSV
Sbjct: 322 ISHL--IGWTAFLSNMLFFTDFMGQIVYRGDPYSAHNSTEFLIYERGVEVGCWGFCINSV 379
Query: 342 VLAVTSLLMERLCRKRGAGFVWGISNIFMAICFIAMLVLTYAANSIGYVSKGQPPPTGIV 401
++ S + L G + F L+ IG V
Sbjct: 380 FSSLYSYFQKVLVSYIG----------LKGLYFTGYLLFGLGTGFIGLFPN--------V 421
Query: 402 IAALAIFTILGFPMAITYSVPYALISTHIE---------PLG------LGQGLSMGVLNL 446
+ L + ++ G + Y+VP+ LI+ + P G G+G+ L
Sbjct: 422 YSTLVLCSLFGVMSSTLYTVPFNLITEYHREEEKERQQAPGGDPDNSVRGKGMDCATLTC 481
Query: 447 AIVVPQIVVSLGSG 460
+ + QI+V G G
Sbjct: 482 MVQLAQILVGGGLG 495
>MATP_MOUSE (P58355) Membrane-associated transporter protein (AIM-1
protein) (Melanoma antigen AIM1) (Underwhite protein)
Length = 530
Score = 120 bits (300), Expect = 1e-26
Identities = 131/534 (24%), Positives = 213/534 (39%), Gaps = 122/534 (22%)
Query: 4 PTTTNPHRSRTRSSTSTSTSRPVQPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYV 63
PT T+ ++S S QP+ +L+ + G +F +A++ + +TP +
Sbjct: 7 PTDTHTYQSLAEDCPFGSVE------QPKRSTGRLVMHSMAMFGREFCYAVEAAYVTPVL 60
Query: 64 QQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVII 123
+G+P S++WL P+ G +QP+VG SD C +R+GRRRP+IL A +++ + +
Sbjct: 61 LSVGLPKSLYSMVWLLSPILGFLLQPVVGSASDHCRARWGRRRPYILTLAIMMLLGMAL- 119
Query: 124 GYAADIGYLIGDDITQ----NYRP---FAIVVFVIGFWILDVANNVTQGPCRALLADLTC 176
YL GD + N R +AI + ++G + D + + GP +A L D+
Sbjct: 120 -------YLNGDAVVSALVANPRQKLIWAISITMVGVVLFDFSADFIDGPIKAYLFDVCS 172
Query: 177 NDARRTRVANAYFSLFMAVGNILGYATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDV 236
+ + + Y +LF G LGY G+ W + L + FF
Sbjct: 173 HQDKEKGL--HYHALFTGFGGALGYILGAID-WVHLDLGRLLG------TEFQVMFFFSA 223
Query: 237 AFIVVTTYLSIVSAHEVP------------------LSSSGAGESGSAEEA--------- 269
+++ + S E P LS+SG E GS E+
Sbjct: 224 LVLILCFITHLCSIPEAPLRDAATDPPSQQDPQGSSLSASGMHEYGSIEKVKNGGADTEQ 283
Query: 270 ------------------FMWELFGTFKYFSMPV-WIVLSVTALTWIGWFPF---NLFDT 307
M L +MP + L V+ L IGW F LF T
Sbjct: 284 PVQEWKNKKPSGQSQRTMSMKSLLRAL--VNMPSHYRCLCVSHL--IGWTAFLSNMLFFT 339
Query: 308 DWMGREIYGGDPEGG------LIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGF 361
D+MG+ +Y GDP G LIY+ GV +G GL +NSV +V S + + G
Sbjct: 340 DFMGQIVYHGDPYGAHNSTEFLIYERGVEVGCWGLCINSVFSSVYSYFQKAMVSYIG--- 396
Query: 362 VWGISNIFMAICFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSV 421
+ F+ L+ IG V + L + ++ G + Y+V
Sbjct: 397 -------LKGLYFMGYLLFGLGTGFIGLFPN--------VYSTLVLCSMFGVMSSTLYTV 441
Query: 422 PYALISTH---------------IEPLGLGQGLSMGVLNLAIVVPQIVVSLGSG 460
P+ LI+ + + G G+G+ L + + QI+V G G
Sbjct: 442 PFNLIAEYHREEEKEKGQEAPGGPDNQGRGKGVDCAALTCMVQLAQILVGGGLG 495
>PRST_MOUSE (Q8K0H7) Prostein
Length = 553
Score = 112 bits (281), Expect = 2e-24
Identities = 105/410 (25%), Positives = 178/410 (42%), Gaps = 40/410 (9%)
Query: 37 QLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSD 96
QLL V + G++ A ++ + P + ++G+ K+ +++ GPV GL PL+G SD
Sbjct: 17 QLLLVNLLTFGLEVCLAAGITYVPPLLLEVGVEEKFMTMVLGIGPVLGLVSVPLLGSASD 76
Query: 97 RCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYRPFAIVVFVIGFWIL 156
+ R+GRRRPFI + +++++ +I A + L+ D RP + + ++G +L
Sbjct: 77 QWRGRYGRRRPFIWALSLGVLLSLFLIPRAGWLAGLLYPDT----RPLELALLILGVGLL 132
Query: 157 DVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGYATGSYSGWYKIFTFT 216
D V P ALL+DL D R A + ++ +++G LGY + +
Sbjct: 133 DFCGQVCFTPLEALLSDL-FRDPDHCRQAFSVYAFMISLGGCLGYLLPAIDWDTSVLAPY 191
Query: 217 LTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVP---LSSSGAGESGSAEEAFMWE 273
L L F + +A + T +++ E L S+ + +
Sbjct: 192 LGTQEECLFGLLTLIFLICMAATLFVTEEAVLGPPEPAEGLLVSAVSRRCCPCHVGLAFR 251
Query: 274 LFGTF------KYFSMPVWI--VLSVTALTWIGWFPFNLFDTDWMGREIYGGDP------ 319
GT MP + + +W+ F LF TD++G +Y G P
Sbjct: 252 NLGTLFPRLQQLCCRMPRTLRRLFVAELCSWMALMTFTLFYTDFVGEGLYQGVPRAEPGT 311
Query: 320 EGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAICFIAMLV 379
E YD G+RMG+LGL L + V SL+M+RL +K G V+ + + F
Sbjct: 312 EARRHYDEGIRMGSLGLFLQCAISLVFSLVMDRLVQKFGTRSVY----LASVMTFPVAAA 367
Query: 380 LTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTH 429
T ++S+ +V+ A A T GF + +PY L S +
Sbjct: 368 ATCLSHSV------------VVVTASAALT--GFTFSALQILPYTLASLY 403
>PRST_HUMAN (Q96JT2) Prostein
Length = 553
Score = 110 bits (275), Expect = 8e-24
Identities = 110/413 (26%), Positives = 182/413 (43%), Gaps = 46/413 (11%)
Query: 37 QLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSD 96
QLL V + G++ A ++ + P + ++G+ K+ +++ GPV GL PL+G SD
Sbjct: 17 QLLLVNLLTFGLEVCLAAGITYVPPLLLEVGVEEKFMTMVLGIGPVLGLVCVPLLGSASD 76
Query: 97 RCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYRPFAIVVFVIGFWIL 156
R+GRRRPFI + I++++ +I A G+L G + + RP + + ++G +L
Sbjct: 77 HWRGRYGRRRPFIWALSLGILLSLFLIPRA---GWLAG-LLCPDPRPLELALLILGVGLL 132
Query: 157 DVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGY----------ATGSY 206
D V P ALL+DL D R A + ++ +++G LGY A Y
Sbjct: 133 DFCGQVCFTPLEALLSDL-FRDPDHCRQAYSVYAFMISLGGCLGYLLPAIDWDTSALAPY 191
Query: 207 SGWYKIFTFTLTPACSISC--ANLKSAFFLDVAFIVVTTYLSI--VSAHEVPLSSSGAGE 262
G + F L ++C A L A + LS +S H P + A
Sbjct: 192 LGTQEECLFGLLTLIFLTCVAATLLVAEEAALGPTEPAEGLSAPSLSPHCCPCRARLAFR 251
Query: 263 SGSAEEAFMWELFGTFKYFSMPVWIVLSVTALTWIGWFPFNLFDTDWMGREIYGGDP--- 319
+ A + +L +++ +W+ F LF TD++G +Y G P
Sbjct: 252 NLGALLPRLHQLCCRMPRTLRRLFV---AELCSWMALMTFTLFYTDFVGEGLYQGVPRAE 308
Query: 320 ---EGGLIYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAICFIA 376
E YD GVRMG+LGL L + V SL+M+RL ++ G V+ + +A
Sbjct: 309 PGTEARRHYDEGVRMGSLGLFLQCAISLVFSLVMDRLVQRFGTRAVY--------LASVA 360
Query: 377 MLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTH 429
+ A + + +V A+ A + GF + +PY L S +
Sbjct: 361 AFPVAAGATCLSH-------SVAVVTASAA---LTGFTFSALQILPYTLASLY 403
>PRST_MACFA (Q95KI5) Prostein (QmoA-10594/QtrA-11310)
Length = 553
Score = 110 bits (274), Expect = 1e-23
Identities = 108/426 (25%), Positives = 178/426 (41%), Gaps = 72/426 (16%)
Query: 37 QLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSD 96
QLL + + G++ A ++ + P + ++G+ K+ +++ GPV GL PL+G SD
Sbjct: 17 QLLLINLLTFGLEVCLAAGITYVPPLLLEVGVEEKFMTMVLGIGPVLGLVSVPLLGSASD 76
Query: 97 RCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYRPFAIVVFVIGFWIL 156
R+GRRRPFI + I++++ +I A G+L G + + RP + + ++G +L
Sbjct: 77 HWRGRYGRRRPFIWALSLGILLSLFLIPRA---GWLAG-LLCPDPRPLELALLILGVGLL 132
Query: 157 DVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGYATGSYSGWYKIFTFT 216
D V P ALL+DL D R A + ++ +++G LGY + W T
Sbjct: 133 DFCGQVCFTPLEALLSDL-FRDPDHCRQAYSVYAFMISLGGCLGYLLPAID-W---DTSA 187
Query: 217 LTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSSSGAGESGSAEEAFMWELFG 276
L P + F L + +++ A E L + E SA
Sbjct: 188 LAPYLG---TQEECLFGLLTLIFLTCVAATLLVAEEAALGPAEPAEGLSAP--------- 235
Query: 277 TFKYFSMPVWIVLSVTAL---------------------------TWIGWFPFNLFDTDW 309
+ P W L+ L +W+ F LF TD+
Sbjct: 236 SLPSHCCPCWARLAFRNLGALLPRLHQLCCRMPRTLRRLFVAELCSWMALMTFTLFYTDF 295
Query: 310 MGREIYGGDPEGGL------IYDTGVRMGALGLLLNSVVLAVTSLLMERLCRKRGAGFVW 363
+G +Y G P L YD GVRMG+LGL L + V SL+M+RL ++ G V+
Sbjct: 296 VGEGLYQGVPRAELGTEARRHYDEGVRMGSLGLFLQCAISLVFSLVMDRLVQRFGTRAVY 355
Query: 364 GISNIFMAICFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPY 423
+ +A + A + + +V A+ A + GF + +PY
Sbjct: 356 --------LASVAAFPVAAGATCLSH-------SVAVVTASAA---LTGFTFSALQILPY 397
Query: 424 ALISTH 429
L S +
Sbjct: 398 TLASLY 403
>PAST_MOUSE (Q8BIV7) Proton-associated sugar transporter A (PAST-A)
(Deleted in neuroblastoma 5 protein homolog) (DNb-5
homolog)
Length = 751
Score = 110 bits (274), Expect = 1e-23
Identities = 70/231 (30%), Positives = 116/231 (49%), Gaps = 11/231 (4%)
Query: 29 VQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLFVQ 88
+ P+ +LL + GI+F +A++ + +TP + Q+G+P + S++W P+ G +Q
Sbjct: 78 LHPQRSFWELLFNGCILFGIEFSYAMETAYVTPVLLQMGLPDQLYSLVWFISPILGFLLQ 137
Query: 89 PLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYRPFAIVV 148
PL+G SDRC+SRFGRRRPFILV A ++ + ++ DIG + D T + + I++
Sbjct: 138 PLLGAWSDRCTSRFGRRRPFILVLAIGALLGLSLLLNGRDIGMALADTATNH--KWGILL 195
Query: 149 FVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGYATGSYSG 208
V G ++D + + P A + D+ C + R N + +L +G GY G
Sbjct: 196 TVCGVVLMDFSADSADNPSHAXMMDV-CGPVDQDRGLNIH-ALMAGLGGGFGYVVGGIH- 252
Query: 209 WYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSSSG 259
W K T L+ + + VTT L+++S E PL G
Sbjct: 253 WDK------TSFGRALGGQLRVIYVFTAITLSVTTVLTLISIPERPLRPLG 297
Score = 51.6 bits (122), Expect = 5e-06
Identities = 51/220 (23%), Positives = 88/220 (39%), Gaps = 40/220 (18%)
Query: 296 WIGWFPFN---LFDTDWMGREIYGGDP------EGGLIYDTGVRMGALGLLLNSVVLAVT 346
++GW F LF TD+MG ++ GDP E Y++GV MG G+ + + A
Sbjct: 534 FLGWLSFEGMLLFYTDFMGEVVFQGDPKAPHTSEAYQKYNSGVTMGCWGMCIYAFSAAFY 593
Query: 347 SLLMERLCRKRGAGFVWGISNIFMAICFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALA 406
S ++E+L + + + Y A +G G + + L+
Sbjct: 594 SAILEKLEE---------------CLSVRTLYFIAYLAFGLG---TGLATLSRNLYVVLS 635
Query: 407 IFTILGFPMAITYSVPYALISTHIEPLGL----------GQGLSMGVLNLAIVVPQIVVS 456
+ T G + ++PY+L+ + + G G+ + +L+ + QI+VS
Sbjct: 636 LCTTYGILFSTLCTLPYSLLCDYYQSKKFAGSSADGTRRGMGVDISLLSCQYFLAQILVS 695
Query: 457 LGSGPWDQLFGGGNSPAF---AVAAVAALLSGLLALLAIP 493
L GP G N + V+ + L S L IP
Sbjct: 696 LVLGPLTSAVGSANGVMYFSSLVSFLGCLYSSLCVTYEIP 735
>PAST_RAT (Q8K4S3) Proton-associated sugar transporter A (PAST-A)
Length = 751
Score = 109 bits (273), Expect = 1e-23
Identities = 70/233 (30%), Positives = 116/233 (49%), Gaps = 11/233 (4%)
Query: 27 QPVQPRTPLRQLLRVASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLF 86
+ + P+ +LL + GI+F +A++ + +TP + Q+G+P + S++W P+ G
Sbjct: 76 EDLHPQRSFWELLFNGCILFGIEFSYAMETAYVTPVLLQMGLPDQLYSLVWFISPILGFL 135
Query: 87 VQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYRPFAI 146
+QPL+G SDRC+SRFGRRRPFILV A ++ + ++ DIG + D T + + I
Sbjct: 136 LQPLLGAWSDRCTSRFGRRRPFILVLAIGALLGLSLLLNGRDIGMALADTATNH--KWGI 193
Query: 147 VVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGYATGSY 206
++ V G ++D + + P A + D+ C + R N + +L +G GY G
Sbjct: 194 LLTVCGVVLMDFSADSADNPSHAYMMDV-CGPVDQDRGLNIH-ALMAGLGGGFGYVVGGI 251
Query: 207 SGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSSSG 259
W K T L+ + + VTT ++VS E PL G
Sbjct: 252 H-WDK------TSFGRALGGQLRVIYIFTAITLSVTTVFTLVSIPERPLRPLG 297
Score = 50.1 bits (118), Expect = 1e-05
Identities = 51/220 (23%), Positives = 89/220 (40%), Gaps = 40/220 (18%)
Query: 296 WIGWFPFN---LFDTDWMGREIYGGDP------EGGLIYDTGVRMGALGLLLNSVVLAVT 346
++GW F LF TD+MG ++ GDP E Y++GV MG G+ + + A
Sbjct: 534 FLGWLSFEGMLLFYTDFMGEVVFQGDPKAPHASEAYQKYNSGVTMGCWGMCIYAFSAAFY 593
Query: 347 SLLMERLCRKRGAGFVWGISNIFMAICFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALA 406
S ++E+L +++ L + A + + G + + L+
Sbjct: 594 SAILEKLEE------------------CLSVRTLYFIAYLLFGLGTGLATLSRNLYVVLS 635
Query: 407 IFTILGFPMAITYSVPYALISTHIEPLGL----------GQGLSMGVLNLAIVVPQIVVS 456
+ T G + ++PY+L+ + + G G+ + +L+ + QI+VS
Sbjct: 636 LCTHYGILFSTLCTLPYSLLCDYYQSKKFAGSSADGTRRGMGVDISLLSCQYFLAQILVS 695
Query: 457 LGSGPWDQLFGGGNSP---AFAVAAVAALLSGLLALLAIP 493
L GP G N A V+ + L S L IP
Sbjct: 696 LVLGPLTSAVGSANGVMYFASLVSFLGCLYSSLCVTYEIP 735
>PAST_HUMAN (Q9Y2W3) Proton-associated sugar transporter A (PAST-A)
(Deleted in neuroblastoma 5 protein) (DNb-5) (Fragment)
Length = 509
Score = 53.5 bits (127), Expect = 1e-06
Identities = 54/228 (23%), Positives = 94/228 (40%), Gaps = 56/228 (24%)
Query: 296 WIGWFPFN---LFDTDWMGREIYGGDP------EGGLIYDTGVRMGALGLLLNSVVLAVT 346
++GW F LF TD+MG ++ GDP E Y++GV MG G+ + + A
Sbjct: 292 FLGWLSFEGMLLFYTDFMGEVVFQGDPKAPHTSEAYQKYNSGVTMGCWGMCIYAFSAAFY 351
Query: 347 SLLMERLCRKRGAGFVWGISNIFMAICFIAMLVLTYAANSIGYVSKGQPPPTGIVIAALA 406
S ++E+L F+++ + + Y A +G TG+ +
Sbjct: 352 SAILEKL-------------EEFLSV--RTLYFIAYLAFGLG---------TGLATLSRN 387
Query: 407 IFTILGFPMAITYSV--------PYALISTHIEPLGL----------GQGLSMGVLNLAI 448
++ +L + ITY + PY+L+ + + G G+ + +L+
Sbjct: 388 LYVVLS--LCITYGILFSTLCTLPYSLLCDYYQSKKFAGSSADGTRRGMGVDISLLSCQY 445
Query: 449 VVPQIVVSLGSGPWDQLFGGGNSPAF---AVAAVAALLSGLLALLAIP 493
+ QI+VSL GP G N + V+ + L S L + IP
Sbjct: 446 FLAQILVSLVLGPLTSAVGSANGVMYFSSLVSFLGCLYSSLFVIYEIP 493
>YD74_SYNY3 (P74168) Hypothetical symporter sll1374
Length = 544
Score = 47.8 bits (112), Expect = 7e-05
Identities = 45/157 (28%), Positives = 70/157 (43%), Gaps = 15/157 (9%)
Query: 50 FGWALQLSLLTPYV-----QQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGR 104
FG A+ ++L Y+ GIP A + + G + P++G LSDR SR+GR
Sbjct: 23 FGPAITANILVFYLLFFLTDVAGIPAALAGSVLMIGKIFDAINDPIIGLLSDRTRSRWGR 82
Query: 105 RRPFILVGAASIVVAVVIIGYAAD--IGYLIGDDITQNYRPFAIVVFVIGFWILDVANNV 162
R P++L G + Y A I + D +T + F + +V ++
Sbjct: 83 RLPWMLGGMIPFA-----LFYTAQWLIPHFSDDRLTNQWGLF--IYYVAIAMAFNLCYTT 135
Query: 163 TQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNIL 199
P AL +LT N RTR+ N++ F G+IL
Sbjct: 136 VNLPYTALTPELTQNYNERTRL-NSFRFAFSIGGSIL 171
>LACY_LACDE (P22733) Lactose permease (Lactose-proton symporter)
(Lactose transport protein)
Length = 627
Score = 41.6 bits (96), Expect = 0.005
Identities = 41/190 (21%), Positives = 84/190 (43%), Gaps = 13/190 (6%)
Query: 82 VSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNY 141
+ + + PL+G+ DR SR+G+ +P+++ G I+ ++ ++ D G I Q+
Sbjct: 58 IGEVLLDPLIGNAIDRTESRWGKFKPWVVGG--GIISSLALLALFTDFG-----GINQSK 110
Query: 142 RPFAIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGY 201
+V+F I + I+D+ + A++ L+ + R + S F VG+ +G
Sbjct: 111 PVVYLVIFGIVYLIMDIFYSFKDTGFWAMIPALSLDSREREKT-----STFARVGSTIGA 165
Query: 202 ATGSYSGWYKIFTFTLTPACSISCANLKSAFFLDVAFIVVTTYLSI-VSAHEVPLSSSGA 260
I F+ + A F L VA + + T +++ + HEV + +
Sbjct: 166 NLVGVVITPIILFFSASKANPNGDKQGWFFFALIVAIVGILTSITVGLGTHEVKSALRES 225
Query: 261 GESGSAEEAF 270
E + ++ F
Sbjct: 226 NEKTTLKQVF 235
>LACY_STRTR (P23936) Lactose permease (Lactose-proton symporter)
(Lactose transport protein)
Length = 634
Score = 40.4 bits (93), Expect = 0.011
Identities = 41/191 (21%), Positives = 91/191 (47%), Gaps = 21/191 (10%)
Query: 85 LFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYRPF 144
+F+ PL+G++ D ++++G+ +P+++ G I+ ++ ++ D+G L PF
Sbjct: 68 VFIDPLIGNMIDNTNTKYGKFKPWVVGG--GIISSITLLLLFTDLGGL------NKTNPF 119
Query: 145 A-IVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILGYAT 203
+V+F I + ++DV ++ +++ L+ + R ++A F +G+ +G
Sbjct: 120 LYLVLFGIIYLVMDVFYSIKDIGFWSMIPALSLDSHEREKMAT-----FARIGSTIGANI 174
Query: 204 GSYSGWYKIFTFTLTPACSISCANLKSAFF---LDVAFIVVTTYLSI-VSAHEVPLSSSG 259
+ + F++T + S + KS +F VA I V T +++ + EV
Sbjct: 175 VGVAIMPIVLFFSMT---NNSGSGDKSGWFWFAFIVALIGVITSIAVGIGTREVESKIRD 231
Query: 260 AGESGSAEEAF 270
E S ++ F
Sbjct: 232 NNEKTSLKQVF 242
>YICJ_ECOLI (P31435) Hypothetical symporter yicJ
Length = 460
Score = 40.0 bits (92), Expect = 0.014
Identities = 35/143 (24%), Positives = 64/143 (44%), Gaps = 11/143 (7%)
Query: 54 LQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGA 113
+ L ++ Y GIP + ++L P +G L+DR SR+G+ RP++L GA
Sbjct: 28 VMLYMMFFYTDIFGIPAGFVGTMFLVARALDAISDPCMGLLADRTRSRWGKFRPWVLFGA 87
Query: 114 ASIVVAVVIIGYAADIGYLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLAD 173
+ V ++ Y+ D++ N + ++ I + +L + V P AL
Sbjct: 88 LPFGI-VCVLAYST-------PDLSMNGK---MIYAAITYTLLTLLYTVVNIPYCALGGV 136
Query: 174 LTCNDARRTRVANAYFSLFMAVG 196
+T + +R + + F L A G
Sbjct: 137 ITNDPTQRISLQSWRFVLATAGG 159
>YNAJ_BACSU (P94488) Hypothetical symporter ynaJ
Length = 463
Score = 39.7 bits (91), Expect = 0.018
Identities = 26/84 (30%), Positives = 40/84 (46%), Gaps = 3/84 (3%)
Query: 58 LLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAASIV 117
LL Y G+ A ++L + P +G + DR +SRF R RP++L GA V
Sbjct: 35 LLFFYTDVFGLSAAAAGTMFLVVRIIDALADPFIGTIVDRTNSRFARFRPYLLFGAFPFV 94
Query: 118 VAVVIIGYA---ADIGYLIGDDIT 138
+ ++ +D+G LI IT
Sbjct: 95 ILAILCFTTPDFSDMGKLIYAYIT 118
>YAGG_ECOLI (P75683) Hypothetical symporter yagG
Length = 460
Score = 38.1 bits (87), Expect = 0.053
Identities = 25/80 (31%), Positives = 35/80 (43%), Gaps = 1/80 (1%)
Query: 50 FGWALQLSLLTP-YVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPF 108
F W + LL Y G+ ++L V PL+G L DR +R G+ RPF
Sbjct: 21 FVWQATMFLLAYFYTDVFGLSAGIMGTLFLVSRVLDAVTDPLMGLLVDRTRTRHGQFRPF 80
Query: 109 ILVGAASIVVAVVIIGYAAD 128
+L GA + V+ Y D
Sbjct: 81 LLWGAIPFGIVCVLTFYTPD 100
>YJMB_BACSU (O34961) Hypothetical symporter yjmB
Length = 459
Score = 37.7 bits (86), Expect = 0.070
Identities = 37/149 (24%), Positives = 68/149 (44%), Gaps = 24/149 (16%)
Query: 55 QLSLLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSD--RCSSRFGRRRPFILVG 112
Q+ LL + GIP A I+L + P+VG D + + G+ RP++L+G
Sbjct: 41 QIYLLKYFTDVAGIPAAMAGGIFLVSKLFAAITDPIVGSSIDYRKNIGKRGKFRPYLLIG 100
Query: 113 AASIVVAVVIIGYAADI---GYLIGDDITQNYRPFAIVVFVIGFWILDVANNVTQGPCRA 169
+ + V V+I + ++ G LI Y + +++ IG+ +++ P +
Sbjct: 101 SIVLAVLTVLIFLSPNVSTTGKLI-------YAYASYMIWGIGYSFVNI-------PYGS 146
Query: 170 LLADLTCNDARRTRVANAYFSLFMAVGNI 198
L A +T N RT + S F +G++
Sbjct: 147 LGAAMTQNSEDRTSI-----STFRQIGSL 170
>UIDB_ECOLI (P30868) Glucuronide carrier protein (Glucuronide
permease)
Length = 457
Score = 36.6 bits (83), Expect = 0.16
Identities = 23/83 (27%), Positives = 39/83 (46%), Gaps = 1/83 (1%)
Query: 41 VASVASGIQFGWALQLSLLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSS 100
+ VA+ F L LL+ Y G+ A + L V F G + D ++
Sbjct: 15 LGDVANNFAFAMGA-LFLLSYYTDVAGVGAAAAGTMLLLVRVFDAFADVFAGRVVDSVNT 73
Query: 101 RFGRRRPFILVGAASIVVAVVII 123
R+G+ RPF+L G A +++ V++
Sbjct: 74 RWGKFRPFLLFGTAPLMIFSVLV 96
Score = 32.0 bits (71), Expect = 3.8
Identities = 30/116 (25%), Positives = 57/116 (48%), Gaps = 12/116 (10%)
Query: 331 MGALGLLLNSVVLAVTSLLMERLCRKRGAGFVWGISNIFMAICFIAMLVLTYAANSI--G 388
+GAL +L+++ ++ +SL R +V + +F + + LV T A+ + G
Sbjct: 235 IGALCVLISTFAVSASSLFYVR--------YVLNDTGLFTVLVLVQNLVGTVASAPLVPG 286
Query: 389 YVSKGQPPPTGIVIAALAIFTILGFPMAITYSVPYALISTHIEPLGLGQGLSMGVL 444
V++ T ++ A L L F +S+P AL++ I +GQG++M V+
Sbjct: 287 MVARIGKKNTFLIGALLGTCGYLLFFWVSVWSLPVALVALAI--ASIGQGVTMTVM 340
>RAFP_PEDPE (P43466) Raffinose carrier protein (Raffinose permease)
Length = 641
Score = 36.6 bits (83), Expect = 0.16
Identities = 27/116 (23%), Positives = 54/116 (46%), Gaps = 12/116 (10%)
Query: 85 LFVQPLVGHLSDRCSSRFGRRRPFILVGAASIVVAVVIIGYAADIGYLIGDDITQNYRPF 144
LF+ P +G+ DR + G RP+++VG V +++++ ++G L +
Sbjct: 70 LFIDPFIGNAIDRTKNSPGHFRPWVVVGGT--VSSIILLLLFTNLGGLYAKNAM-----I 122
Query: 145 AIVVFVIGFWILDVANNVTQGPCRALLADLTCNDARRTRVANAYFSLFMAVGNILG 200
+VVF I + +D+ + ++L LT + R + A F +G+ +G
Sbjct: 123 YLVVFAILYITMDIFYSFKDVGFWSMLPSLTTDSREREKTAT-----FARLGSTIG 173
>MELB_ENTAE (O07366) Melibiose carrier protein
(Thiomethylgalactoside permease II) (Melibiose permease)
(Na+ (Li+)/melibiose symporter) (Melibiose transporter)
Length = 471
Score = 36.6 bits (83), Expect = 0.16
Identities = 45/220 (20%), Positives = 91/220 (40%), Gaps = 44/220 (20%)
Query: 58 LLTPYVQQLGIPHKWASIIWLCGPVSGLFVQPLVGHLSDRCSSRFGRRRPFILVGAA--S 115
L+ Y +G+ ++L + P++G + + SR+G+ +P+IL+G S
Sbjct: 29 LMYYYTDIVGLSVGVVGTLFLVARILDAIADPIMGWIVNCTRSRWGKFKPWILIGTITNS 88
Query: 116 IVVAVVIIGYAADIGYLIGDD-ITQNYRPFAIVVFVIGFWILDVANNVTQGPCRALLADL 174
+V+ ++ + G L+ +T F + + FW +L+ +
Sbjct: 89 VVLYMLFSAHHFSGGALLAWVWLTYLLWGFTYTIMDVPFW--------------SLVPTI 134
Query: 175 TCNDARRTRVAN-----AYFSLFMAVGNILGYAT---GSYSGW-YKIFTFTLTPACSISC 225
T + R ++ A + F+ G L + + G+ G+ +++FT L
Sbjct: 135 TLDKREREQLVPYPRFFASLAGFVTAGVTLPFVSAVGGADRGFGFQMFTLVL-------- 186
Query: 226 ANLKSAFFLDVAFIVVTTYLSIVSAHEVPLSSSGAGESGS 265
+AF V++T +++ + HEV S SG E S
Sbjct: 187 ----------IAFFVISTLVTLRNVHEVYSSDSGVSEDSS 216
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.140 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 58,904,816
Number of Sequences: 164201
Number of extensions: 2534945
Number of successful extensions: 8656
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 8598
Number of HSP's gapped (non-prelim): 54
length of query: 504
length of database: 59,974,054
effective HSP length: 115
effective length of query: 389
effective length of database: 41,090,939
effective search space: 15984375271
effective search space used: 15984375271
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC146866.2


