

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146866.12 + phase: 0
(273 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
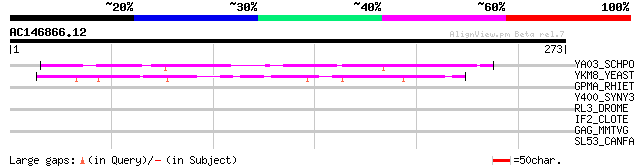
Score E
Sequences producing significant alignments: (bits) Value
YA03_SCHPO (Q09676) Hypothetical protein C5H10.03 in chromosome I 58 2e-08
YKM8_YEAST (P36069) Hypothetical 33.8 kDa protein in MYO3-PGM1 i... 52 1e-06
GPMA_RHIET (Q8KL44) 2,3-bisphosphoglycerate-dependent phosphogly... 33 0.97
Y400_SYNY3 (Q55129) Hypothetical protein sll0400 31 3.7
RL3_DROME (O16797) 60S ribosomal protein L3 30 6.3
IF2_CLOTE (Q895J8) Translation initiation factor IF-2 30 6.3
GAG_MMTVG (P03343) Gag polyprotein [Contains: Protein p10; Phosp... 30 6.3
SL53_CANFA (P31637) Sodium/myo-inositol cotransporter (Na(+)/myo... 30 8.2
>YA03_SCHPO (Q09676) Hypothetical protein C5H10.03 in chromosome I
Length = 219
Score = 58.2 bits (139), Expect = 2e-08
Identities = 60/228 (26%), Positives = 98/228 (42%), Gaps = 40/228 (17%)
Query: 16 KTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHVKAIGLSKKIEL 75
KT++L+RH Q HNV +++H + D LT G +Q E L K ++ SK+I +
Sbjct: 7 KTVYLIRHGQAQHNVGPDEDH------NIRDPVLTSEGIEQCEALAKELE----SKQIPI 56
Query: 76 --VVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLNCPPFVAVELC 133
+V SP+ RT+QT G +K P+ I PF
Sbjct: 57 DGIVCSPMRRTLQTMEIALKKYLAEGGPDKVPVYIS----------------PFF----- 95
Query: 134 REQMGLHPCDKRRTVSEYRHMFPGIDFSLIETDDDTWWKPEREKKEEVT---GRGLKFLE 190
+++G PCD + + ++P +F D + + +VT R + LE
Sbjct: 96 -QEVGHLPCDIGLELDKLNKLYPKYNFQ--SCQDGIYPEKRDIYASDVTISAIRSKEALE 152
Query: 191 WLCTRKEKEIAVVTHSSFLFNTLSAFGNDCHPNIKTEMCAHFANCELR 238
+L +++IAV+THS+F+ L + + F NCE R
Sbjct: 153 YLAALPQQQIAVITHSAFIRFLLKKMVKAADIDFLPPQLS-FKNCEFR 199
>YKM8_YEAST (P36069) Hypothetical 33.8 kDa protein in MYO3-PGM1
intergenic region
Length = 295
Score = 52.4 bits (124), Expect = 1e-06
Identities = 67/246 (27%), Positives = 102/246 (41%), Gaps = 62/246 (25%)
Query: 14 HSKTIHLVRHAQGVHNVE----GEKNHDAYLSY-------DFFDANLTPLGWQQVENLQK 62
H K + L RH QG HN G + DAY S ++ D+ LTPLG QV
Sbjct: 52 HYKLLILARHGQGYHNAAILRYGMEKWDAYWSLLSGDEHGEWLDSKLTPLGKDQVRRTGS 111
Query: 63 HVKAIGLSKKIELV----VVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPA 118
+V + ++K++ ++ SP+ R ++T + + P++ E + PA
Sbjct: 112 NV-LLPMAKQLGMLPHVFFSSPMRRCLETFIESW-----------TPVLAET---QELPA 156
Query: 119 VSSLNCPPFVAVELCREQMGLHPCDKR----RTVSEYRHMFPGIDFSL------------ 162
+ ++ +E RE +G H CDKR V EY+ DFS
Sbjct: 157 GTKISTR---IIEGLRETLGSHTCDKRVAHSMAVDEYQ------DFSTESGHTVHWQYVP 207
Query: 163 -IETDDDTWWKPEREKKEEVTGRGLKFLEWL---CTRKEKEIAVVTHSSFLFNTLSAFGN 218
DD+ W RE E+ R L L L + +EK I++ HS + + L N
Sbjct: 208 DYPEDDELWLPDHRETCAEMDKRTLNGLFELFNQLSSEEKFISLTCHSGVIQSVLR---N 264
Query: 219 DCHPNI 224
HP I
Sbjct: 265 LQHPPI 270
>GPMA_RHIET (Q8KL44) 2,3-bisphosphoglycerate-dependent
phosphoglycerate mutase (EC 5.4.2.1)
(Phosphoglyceromutase) (PGAM) (BPG-dependent PGAM)
(dPGM)
Length = 209
Score = 32.7 bits (73), Expect = 0.97
Identities = 26/107 (24%), Positives = 44/107 (40%), Gaps = 12/107 (11%)
Query: 17 TIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKHVKAIGLSKKIELV 76
T+ +VRH Q N GE + D LT GW + + +G+S ++
Sbjct: 3 TLVIVRHGQSEGNARGEFTGTS-------DVPLTQEGWSESRRAGSLLANLGIS--FDIA 53
Query: 77 VVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLN 123
S LLRT+ T + T+G P+ + D+ ++ +N
Sbjct: 54 FSSALLRTVDTCRAILN---ETNGDLLEPIRRTELNERDYGQLTGIN 97
>Y400_SYNY3 (Q55129) Hypothetical protein sll0400
Length = 164
Score = 30.8 bits (68), Expect = 3.7
Identities = 15/43 (34%), Positives = 27/43 (61%), Gaps = 2/43 (4%)
Query: 46 DANLTPLGWQQVENLQKHVKAIGLSKKIELVVVSPLLRTMQTA 88
D LT G + + + + ++AIG+ + +L++ SPL+R QTA
Sbjct: 22 DRQLTKKGKDKTQRVAQRLQAIGV--EFDLILTSPLVRAQQTA 62
>RL3_DROME (O16797) 60S ribosomal protein L3
Length = 415
Score = 30.0 bits (66), Expect = 6.3
Identities = 28/93 (30%), Positives = 46/93 (49%), Gaps = 11/93 (11%)
Query: 6 GQSLYPLHHSKTIH--LVRHAQGVHNVEGEK-NHDAYLSYDFFDANLTPLG----WQQVE 58
GQ Y HH I+ + R G+H +G+ ++A YD D ++TP+G + +V
Sbjct: 269 GQKGY--HHRTEINKKIYRIGAGIHTKDGKVIKNNASTEYDLTDKSITPMGGFPHYGEVN 326
Query: 59 NLQKHVK--AIGLSKKIELVVVSPLLRTMQTAV 89
N +K IG K+I + S L T ++A+
Sbjct: 327 NDFVMIKGCCIGSKKRIITLRKSLLKHTKRSAL 359
>IF2_CLOTE (Q895J8) Translation initiation factor IF-2
Length = 685
Score = 30.0 bits (66), Expect = 6.3
Identities = 16/55 (29%), Positives = 28/55 (50%)
Query: 69 LSKKIELVVVSPLLRTMQTAVGVFGGEANTDGVNKPPLMIENVGHSDHPAVSSLN 123
L +K E +VV + ++ +V + E +G K P ++ +GH DH S L+
Sbjct: 151 LGEKFEAIVVQREVDVLEESVEQYIEEEEEEGTVKRPPVVTVMGHVDHGKTSLLD 205
>GAG_MMTVG (P03343) Gag polyprotein [Contains: Protein p10;
Phosphorylated protein pp21; Protein p3; Protein p8;
Major core protein p27] (Fragment)
Length = 353
Score = 30.0 bits (66), Expect = 6.3
Identities = 14/51 (27%), Positives = 25/51 (48%)
Query: 13 HHSKTIHLVRHAQGVHNVEGEKNHDAYLSYDFFDANLTPLGWQQVENLQKH 63
H K +V+ + + EK+ A+L+ D+ D +L+P W +E H
Sbjct: 169 HVRKVKKIVQRKENSEHKRKEKDQKAFLATDWNDDDLSPEDWDNLEEQAAH 219
>SL53_CANFA (P31637) Sodium/myo-inositol cotransporter
(Na(+)/myo-inositol cotransporter)
Length = 718
Score = 29.6 bits (65), Expect = 8.2
Identities = 12/28 (42%), Positives = 18/28 (63%)
Query: 174 EREKKEEVTGRGLKFLEWLCTRKEKEIA 201
ER+K+ E GR KF++W C K K ++
Sbjct: 643 ERKKETEDGGRYWKFIDWFCGFKSKSLS 670
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.135 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,768,634
Number of Sequences: 164201
Number of extensions: 1433038
Number of successful extensions: 3032
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 3025
Number of HSP's gapped (non-prelim): 8
length of query: 273
length of database: 59,974,054
effective HSP length: 109
effective length of query: 164
effective length of database: 42,076,145
effective search space: 6900487780
effective search space used: 6900487780
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC146866.12


