

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
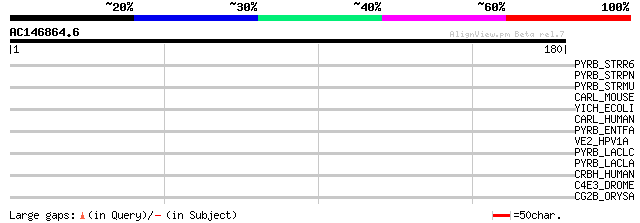
Query= AC146864.6 - phase: 0 /pseudo
(180 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
PYRB_STRR6 (Q8DPH9) Aspartate carbamoyltransferase (EC 2.1.3.2) ... 31 1.8
PYRB_STRPN (Q97QE2) Aspartate carbamoyltransferase (EC 2.1.3.2) ... 31 1.8
PYRB_STRMU (Q8DUP5) Aspartate carbamoyltransferase (EC 2.1.3.2) ... 30 3.0
CARL_MOUSE (Q9D5J6) Carbohydrate kinase-like protein (EC 2.7.1.-) 30 3.0
YICH_ECOLI (P31433) Hypothetical protein yicH 30 3.9
CARL_HUMAN (Q9UHJ6) Carbohydrate kinase-like protein (EC 2.7.1.-) 30 3.9
PYRB_ENTFA (Q9L4T8) Aspartate carbamoyltransferase (EC 2.1.3.2) ... 29 5.2
VE2_HPV1A (P03118) Regulatory protein E2 29 6.7
PYRB_LACLC (Q9L4N6) Aspartate carbamoyltransferase (EC 2.1.3.2) ... 29 6.7
PYRB_LACLA (Q9CF79) Aspartate carbamoyltransferase (EC 2.1.3.2) ... 29 6.7
CRBH_HUMAN (P82279) Crumbs protein homolog 1 precursor 29 6.7
C4E3_DROME (Q9VL92) Cytochrome P450 4e3 (EC 1.14.-.-) (CYPIVE3) 29 6.7
CG2B_ORYSA (Q40671) G2/mitotic-specific cyclin 2 (B-like cyclin)... 28 8.8
>PYRB_STRR6 (Q8DPH9) Aspartate carbamoyltransferase (EC 2.1.3.2)
(Aspartate transcarbamylase) (ATCase)
Length = 307
Score = 30.8 bits (68), Expect = 1.8
Identities = 15/52 (28%), Positives = 25/52 (47%)
Query: 64 PVGGNLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSI 115
P+ NL F +S H+ + LGLE + +T + YDT+L++
Sbjct: 44 PIVSNLFFEDSTRTHKSFEVAEIKLGLERLDFDVKTSSVNKGETLYDTILTL 95
>PYRB_STRPN (Q97QE2) Aspartate carbamoyltransferase (EC 2.1.3.2)
(Aspartate transcarbamylase) (ATCase)
Length = 307
Score = 30.8 bits (68), Expect = 1.8
Identities = 15/52 (28%), Positives = 25/52 (47%)
Query: 64 PVGGNLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSI 115
P+ NL F +S H+ + LGLE + +T + YDT+L++
Sbjct: 44 PIVSNLFFEDSTRTHKSFEVAEIKLGLERLDFDVKTSSVNKGETLYDTILTL 95
>PYRB_STRMU (Q8DUP5) Aspartate carbamoyltransferase (EC 2.1.3.2)
(Aspartate transcarbamylase) (ATCase)
Length = 308
Score = 30.0 bits (66), Expect = 3.0
Identities = 15/48 (31%), Positives = 23/48 (47%)
Query: 68 NLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSI 115
NL F S H+ + LGL+ E A+T + YDT+L++
Sbjct: 51 NLFFEASTRTHKSFEVAEKKLGLDVIEFDAKTSSVNKGETLYDTILTM 98
>CARL_MOUSE (Q9D5J6) Carbohydrate kinase-like protein (EC 2.7.1.-)
Length = 476
Score = 30.0 bits (66), Expect = 3.0
Identities = 20/57 (35%), Positives = 33/57 (57%), Gaps = 4/57 (7%)
Query: 66 GGNLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTE 122
GGN+L + +H Q++ + LGLE EES ++ +++A DT L+I + L E
Sbjct: 321 GGNVL---ATFVHMLVQWMAD-LGLEVEESTVYSRMIQAAAQQKDTHLTITPTVLGE 373
>YICH_ECOLI (P31433) Hypothetical protein yicH
Length = 569
Score = 29.6 bits (65), Expect = 3.9
Identities = 16/44 (36%), Positives = 23/44 (51%), Gaps = 7/44 (15%)
Query: 7 VIRASGLYLLLETNYGQVDHGLLIAFSERWHSETSSFHLPVGEM 50
++ +GLY LL+T +G +H W SE S +HL G M
Sbjct: 16 LVAIAGLYFLLQTRWG-AEH------ISAWVSENSDYHLAFGAM 52
>CARL_HUMAN (Q9UHJ6) Carbohydrate kinase-like protein (EC 2.7.1.-)
Length = 478
Score = 29.6 bits (65), Expect = 3.9
Identities = 20/57 (35%), Positives = 33/57 (57%), Gaps = 4/57 (7%)
Query: 66 GGNLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTSYLTE 122
GGN+L + +H Q++ + LGLE EES ++ +++A DT L+I + L E
Sbjct: 323 GGNVL---ATFVHMLVQWMAD-LGLEVEESTVYSRMIQAAVQQRDTHLTITPTVLGE 375
>PYRB_ENTFA (Q9L4T8) Aspartate carbamoyltransferase (EC 2.1.3.2)
(Aspartate transcarbamylase) (ATCase)
Length = 308
Score = 29.3 bits (64), Expect = 5.2
Identities = 15/48 (31%), Positives = 22/48 (45%)
Query: 68 NLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSI 115
NL F S H+ + LGLE E A ++ YDT+L++
Sbjct: 51 NLFFENSTRTHKSFEVAEKKLGLEVIEFEASRSSVQKGETLYDTVLTM 98
>VE2_HPV1A (P03118) Regulatory protein E2
Length = 401
Score = 28.9 bits (63), Expect = 6.7
Identities = 22/75 (29%), Positives = 33/75 (43%), Gaps = 6/75 (8%)
Query: 69 LLFHESLSIHQGT-QYLVNYLGLEFEESAAQTKRLRSA-----HITYDTLLSIYTSYLTE 122
+LF E S + T QY VNY G F + T R+A H Y TL T+ +
Sbjct: 171 VLFAEEASKYSTTGQYAVNYRGKRFTNVMSSTSSPRAAGAPAVHSDYPTLSESDTAQQST 230
Query: 123 AKSYANQPEEEDSME 137
+ Y P + ++ +
Sbjct: 231 SIDYTELPGQGETSQ 245
>PYRB_LACLC (Q9L4N6) Aspartate carbamoyltransferase (EC 2.1.3.2)
(Aspartate transcarbamylase) (ATCase)
Length = 310
Score = 28.9 bits (63), Expect = 6.7
Identities = 15/48 (31%), Positives = 20/48 (41%)
Query: 68 NLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSI 115
NL F S H LGL+ E AQ + YDT+L++
Sbjct: 51 NLFFENSTRTHHSFHIAERKLGLDVLEFDAQASSISKGETLYDTVLTL 98
>PYRB_LACLA (Q9CF79) Aspartate carbamoyltransferase (EC 2.1.3.2)
(Aspartate transcarbamylase) (ATCase)
Length = 310
Score = 28.9 bits (63), Expect = 6.7
Identities = 15/48 (31%), Positives = 20/48 (41%)
Query: 68 NLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSI 115
NL F S H LGL+ E AQ + YDT+L++
Sbjct: 51 NLFFENSTRTHHSFHIAERKLGLDVLEFDAQASSISKGETLYDTVLTL 98
>CRBH_HUMAN (P82279) Crumbs protein homolog 1 precursor
Length = 1376
Score = 28.9 bits (63), Expect = 6.7
Identities = 36/174 (20%), Positives = 62/174 (34%), Gaps = 26/174 (14%)
Query: 4 FWNVIRASGLYLLLETNYGQVDHGLLIAFSERWHSETSSFHLPVGE-----MTITLDDVS 58
F+ + + Y+L T+ V+ G WH T S P+ + M + +
Sbjct: 1009 FFQLQSGNSFYMLSLTSLQSVNDGT-------WHEVTLSMTDPLSQTSRWQMEVDNETPF 1061
Query: 59 CLLHIPVGGNLLFHESLSIHQGTQYLVNYLGLEFEESAAQTKRLRSAHITYDTLLSIYTS 118
I G ++ I+ G + + N GL+ S + + IY S
Sbjct: 1062 VTSTIATGSLNFLKDNTDIYVGDRAIDNIKGLQGCLSTIE-------------IGGIYLS 1108
Query: 119 YLTEAKSYANQPEEEDSMEWYRTRCIRAFLLYLVGCTLFSDKAGNSCCVVYLKY 172
Y + N+P+EE ++ T + L L C G +C +Y Y
Sbjct: 1109 YFENVHGFINKPQEEQFLK-ISTNSVVTGCLQLNVCNSNPCLHGGNCEDIYSSY 1161
>C4E3_DROME (Q9VL92) Cytochrome P450 4e3 (EC 1.14.-.-) (CYPIVE3)
Length = 526
Score = 28.9 bits (63), Expect = 6.7
Identities = 17/45 (37%), Positives = 23/45 (50%), Gaps = 7/45 (15%)
Query: 7 VIRASGLYLLLETNYGQVDHGLLIAFSERWHSE----TSSFHLPV 47
+I+ S +Y LL G HGLL +F +WH T SFH +
Sbjct: 98 LIQKSTIYDLLHPWLG---HGLLTSFGSKWHKHRKMITPSFHFNI 139
>CG2B_ORYSA (Q40671) G2/mitotic-specific cyclin 2 (B-like cyclin)
(CycOs2)
Length = 419
Score = 28.5 bits (62), Expect = 8.8
Identities = 12/43 (27%), Positives = 22/43 (50%)
Query: 133 EDSMEWYRTRCIRAFLLYLVGCTLFSDKAGNSCCVVYLKYFDD 175
E M ++ + A +Y CT+ K+ N CC ++ KY ++
Sbjct: 326 EYEMLKFQPSMLAAAAIYTAQCTINGFKSWNKCCELHTKYSEE 368
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.137 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,117,767
Number of Sequences: 164201
Number of extensions: 825100
Number of successful extensions: 1646
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1637
Number of HSP's gapped (non-prelim): 14
length of query: 180
length of database: 59,974,054
effective HSP length: 103
effective length of query: 77
effective length of database: 43,061,351
effective search space: 3315724027
effective search space used: 3315724027
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146864.6


