

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
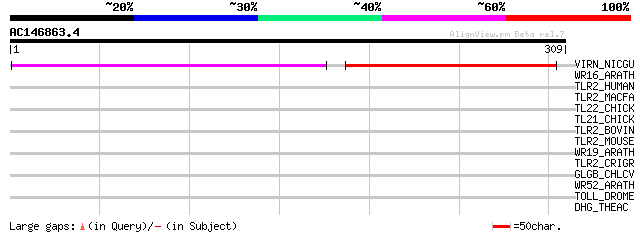
Query= AC146863.4 + phase: 0
(309 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
VIRN_NICGU (Q40392) TMV resistance protein N 137 3e-32
WR16_ARATH (Q9FL92) Probable WRKY transcription factor 16 (WRKY ... 42 0.001
TLR2_HUMAN (O60603) Toll-like receptor 2 precursor (Toll/interle... 36 0.11
TLR2_MACFA (Q95M53) Toll-like receptor 2 precursor 36 0.14
TL22_CHICK (Q9DGB6) Toll-like receptor 2 type 2 precursor 35 0.18
TL21_CHICK (Q9DD78) Toll-like receptor 2 type 1 precursor 35 0.18
TLR2_BOVIN (Q95LA9) Toll-like receptor 2 precursor 35 0.24
TLR2_MOUSE (Q9QUN7) Toll-like receptor 2 precursor 33 0.69
WR19_ARATH (Q9SZ67) Probable WRKY transcription factor 19 (WRKY ... 33 1.2
TLR2_CRIGR (Q9R1F8) Toll-like receptor 2 precursor 33 1.2
GLGB_CHLCV (Q823Y5) 1,4-alpha-glucan branching enzyme (EC 2.4.1.... 32 2.0
WR52_ARATH (Q9FH83) Probable WRKY transcription factor 52 (WRKY ... 30 7.6
TOLL_DROME (P08953) Toll protein precursor 30 7.6
DHG_THEAC (P13203) Glucose 1-dehydrogenase (EC 1.1.1.47) 30 7.6
>VIRN_NICGU (Q40392) TMV resistance protein N
Length = 1144
Score = 137 bits (345), Expect = 3e-32
Identities = 74/176 (42%), Positives = 103/176 (58%), Gaps = 1/176 (0%)
Query: 2 TSPRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAI 61
+S + + VF+SFRG DTR FT HL+ L K I TF+DD L+ G +I +L +AI
Sbjct: 4 SSSSSRWSYDVFLSFRGEDTRKTFTSHLYEVLNDKGIKTFQDDKRLEYGATIPGELCKAI 63
Query: 62 EGSQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGES 121
E SQ IVVFS+NYA+S WCL EL I+ C + + V+P+FYDVD S VR Q + ++
Sbjct: 64 EESQFAIVVFSENYATSRWCLNELVKIMECKTRFKQTVIPIFYDVDPSHVRNQKESFAKA 123
Query: 122 FNYHGKRFQDHSNMVQRRRETLQLVGNISG-WDLRDKPHHAELENIIEHINILGCK 176
F H +++D +QR R L N+ G D RDK + I++ I+ CK
Sbjct: 124 FEEHETKYKDDVEGIQRWRIALNEAANLKGSCDNRDKTDADCIRQIVDQISSKLCK 179
Score = 119 bits (299), Expect = 7e-27
Identities = 58/117 (49%), Positives = 75/117 (63%)
Query: 188 NYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYI 247
+YDVF+SFRG DTR FT H L +GI F+DD +L+ G I L +AIE SQ I
Sbjct: 11 SYDVFLSFRGEDTRKTFTSHLYEVLNDKGIKTFQDDKRLEYGATIPGELCKAIEESQFAI 70
Query: 248 VVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
VVFS+NYA+S WCL EL I+ C + + V+P+FYDVDPS V+ Q + A +H
Sbjct: 71 VVFSENYATSRWCLNELVKIMECKTRFKQTVIPIFYDVDPSHVRNQKESFAKAFEEH 127
>WR16_ARATH (Q9FL92) Probable WRKY transcription factor 16 (WRKY
DNA-binding protein 16)
Length = 1372
Score = 42.4 bits (98), Expect = 0.001
Identities = 31/120 (25%), Positives = 60/120 (49%), Gaps = 9/120 (7%)
Query: 20 DTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIVVFSKNYASST 79
+ R++F HL ALQRK + +D + +S++++ +E ++V +++ N T
Sbjct: 15 EVRYSFVSHLSKALQRKGV----NDVFIDSDDSLSNESQSMVERARVSVMILPGN---RT 67
Query: 80 WCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFN--YHGKRFQDHSNMVQ 137
L +L +L+C + V+PV Y V SE S + F+ +H ++ S +V+
Sbjct: 68 VSLDKLVKVLDCQKNKDQVVVPVLYGVRSSETEWLSALDSKGFSSVHHSRKECSDSQLVK 127
Score = 42.4 bits (98), Expect = 0.001
Identities = 26/93 (27%), Positives = 46/93 (48%), Gaps = 7/93 (7%)
Query: 199 DTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIVVFSKNYASST 258
+ R++F H ALQR+G+N D + + ++ +E ++V +++ N T
Sbjct: 15 EVRYSFVSHLSKALQRKGVN----DVFIDSDDSLSNESQSMVERARVSVMILPGN---RT 67
Query: 259 WCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQ 291
L +L +L C K + V+PV Y V SE +
Sbjct: 68 VSLDKLVKVLDCQKNKDQVVVPVLYGVRSSETE 100
>TLR2_HUMAN (O60603) Toll-like receptor 2 precursor
(Toll/interleukin 1 receptor-like protein 4)
Length = 784
Score = 36.2 bits (82), Expect = 0.11
Identities = 29/101 (28%), Positives = 41/101 (39%), Gaps = 24/101 (23%)
Query: 176 KFPSRTKDLVEINYDVFVSFRGPDTRF----------NFTDHFCAALQRRGINAFRDDTK 225
K PSR I YD FVS+ D + NF F L +R
Sbjct: 633 KAPSRN-----ICYDAFVSYSERDAYWVENLMVQELENFNPPFKLCLHKRDFIP------ 681
Query: 226 LKKGEFIAPGLFRAIEASQVYIVVFSKNYASSTWCLRELEY 266
G++I + +IE S + V S+N+ S WC EL++
Sbjct: 682 ---GKWIIDNIIDSIEKSHKTVFVLSENFVKSEWCKYELDF 719
Score = 31.6 bits (70), Expect = 2.6
Identities = 14/38 (36%), Positives = 21/38 (54%)
Query: 50 GNSIASDLIQAIEGSQVFIVVFSKNYASSTWCLRELAY 87
G I ++I +IE S + V S+N+ S WC EL +
Sbjct: 682 GKWIIDNIIDSIEKSHKTVFVLSENFVKSEWCKYELDF 719
>TLR2_MACFA (Q95M53) Toll-like receptor 2 precursor
Length = 784
Score = 35.8 bits (81), Expect = 0.14
Identities = 25/91 (27%), Positives = 38/91 (41%), Gaps = 19/91 (20%)
Query: 186 EINYDVFVSFRGPDTRF----------NFTDHFCAALQRRGINAFRDDTKLKKGEFIAPG 235
+I YD FVS+ D + NF F L +R G++I
Sbjct: 638 DICYDAFVSYSERDAYWVENLMVQELENFNPPFKLCLHKRDFIP---------GKWIIDN 688
Query: 236 LFRAIEASQVYIVVFSKNYASSTWCLRELEY 266
+ +IE S + V S+N+ S WC EL++
Sbjct: 689 IIDSIEKSHKTVFVLSENFVKSEWCKYELDF 719
Score = 31.6 bits (70), Expect = 2.6
Identities = 14/38 (36%), Positives = 21/38 (54%)
Query: 50 GNSIASDLIQAIEGSQVFIVVFSKNYASSTWCLRELAY 87
G I ++I +IE S + V S+N+ S WC EL +
Sbjct: 682 GKWIIDNIIDSIEKSHKTVFVLSENFVKSEWCKYELDF 719
>TL22_CHICK (Q9DGB6) Toll-like receptor 2 type 2 precursor
Length = 781
Score = 35.4 bits (80), Expect = 0.18
Identities = 22/84 (26%), Positives = 39/84 (46%), Gaps = 5/84 (5%)
Query: 186 EINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFR---DDTKLKKGEFIAPGLFRAIEA 242
+I YD FVS+ D+ N+ ++ + FR G++I + +IE
Sbjct: 635 DICYDAFVSYSENDS--NWVENIMVQQLEQACPPFRLCLHKRDFVPGKWIVDNIIDSIEK 692
Query: 243 SQVYIVVFSKNYASSTWCLRELEY 266
S + V S+++ S WC EL++
Sbjct: 693 SHKTLFVLSEHFVQSEWCKYELDF 716
>TL21_CHICK (Q9DD78) Toll-like receptor 2 type 1 precursor
Length = 793
Score = 35.4 bits (80), Expect = 0.18
Identities = 22/84 (26%), Positives = 39/84 (46%), Gaps = 5/84 (5%)
Query: 186 EINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFR---DDTKLKKGEFIAPGLFRAIEA 242
+I YD FVS+ D+ N+ ++ + FR G++I + +IE
Sbjct: 647 DICYDAFVSYSENDS--NWVENIMVQQLEQACPPFRLCLHKRDFVPGKWIVDNIIDSIEK 704
Query: 243 SQVYIVVFSKNYASSTWCLRELEY 266
S + V S+++ S WC EL++
Sbjct: 705 SHKTLFVLSEHFVQSEWCKYELDF 728
>TLR2_BOVIN (Q95LA9) Toll-like receptor 2 precursor
Length = 784
Score = 35.0 bits (79), Expect = 0.24
Identities = 25/88 (28%), Positives = 40/88 (45%), Gaps = 13/88 (14%)
Query: 186 EINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLK-------KGEFIAPGLFR 238
+I YD FVS+ D+ ++ L + + F KL G++I +
Sbjct: 638 DICYDAFVSYSERDS------YWVENLMVQELEHFNPPFKLCLHKRDFIPGKWIIDNIID 691
Query: 239 AIEASQVYIVVFSKNYASSTWCLRELEY 266
+IE S I V S+N+ S WC EL++
Sbjct: 692 SIEKSHKTIFVLSENFVKSEWCKYELDF 719
Score = 32.0 bits (71), Expect = 2.0
Identities = 15/38 (39%), Positives = 21/38 (54%)
Query: 50 GNSIASDLIQAIEGSQVFIVVFSKNYASSTWCLRELAY 87
G I ++I +IE S I V S+N+ S WC EL +
Sbjct: 682 GKWIIDNIIDSIEKSHKTIFVLSENFVKSEWCKYELDF 719
>TLR2_MOUSE (Q9QUN7) Toll-like receptor 2 precursor
Length = 784
Score = 33.5 bits (75), Expect = 0.69
Identities = 23/91 (25%), Positives = 38/91 (41%), Gaps = 19/91 (20%)
Query: 186 EINYDVFVSFRGPDTRF----------NFTDHFCAALQRRGINAFRDDTKLKKGEFIAPG 235
++ YD FVS+ D+ + N F L +R G++I
Sbjct: 638 DVCYDAFVSYSEQDSHWVENLMVQQLENSDPPFKLCLHKRDF---------VPGKWIIDN 688
Query: 236 LFRAIEASQVYIVVFSKNYASSTWCLRELEY 266
+ +IE S + V S+N+ S WC EL++
Sbjct: 689 IIDSIEKSHKTVFVLSENFVRSEWCKYELDF 719
Score = 31.2 bits (69), Expect = 3.4
Identities = 14/38 (36%), Positives = 21/38 (54%)
Query: 50 GNSIASDLIQAIEGSQVFIVVFSKNYASSTWCLRELAY 87
G I ++I +IE S + V S+N+ S WC EL +
Sbjct: 682 GKWIIDNIIDSIEKSHKTVFVLSENFVRSEWCKYELDF 719
>WR19_ARATH (Q9SZ67) Probable WRKY transcription factor 19 (WRKY
DNA-binding protein 19)
Length = 1895
Score = 32.7 bits (73), Expect = 1.2
Identities = 28/111 (25%), Positives = 46/111 (41%), Gaps = 17/111 (15%)
Query: 188 NYDVFVSFRGPD-TRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVY 246
+YDV + + D + +F H A+L RRGI+ + ++ A+ +V
Sbjct: 667 DYDVVIRYGRADISNEDFISHLRASLCRRGISVYEKFNEVD-----------ALPKCRVL 715
Query: 247 IVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGY 297
I+V + Y S L IL + V P+FY + P + S Y
Sbjct: 716 IIVLTSTYVPSN-----LLNILEHQHTEDRVVYPIFYRLSPYDFVCNSKNY 761
>TLR2_CRIGR (Q9R1F8) Toll-like receptor 2 precursor
Length = 784
Score = 32.7 bits (73), Expect = 1.2
Identities = 24/91 (26%), Positives = 38/91 (41%), Gaps = 19/91 (20%)
Query: 186 EINYDVFVSFRGPDTRF----------NFTDHFCAALQRRGINAFRDDTKLKKGEFIAPG 235
+I YD FVS+ D+ + N F L +R G++I
Sbjct: 638 DICYDAFVSYSEQDSYWVENLMVQQLENSEPPFKLCLHKRDF---------VPGKWIIDN 688
Query: 236 LFRAIEASQVYIVVFSKNYASSTWCLRELEY 266
+ +IE S + V S+N+ S WC EL++
Sbjct: 689 IIDSIEKSHKTLFVLSENFVRSEWCKYELDF 719
Score = 30.8 bits (68), Expect = 4.5
Identities = 14/38 (36%), Positives = 21/38 (54%)
Query: 50 GNSIASDLIQAIEGSQVFIVVFSKNYASSTWCLRELAY 87
G I ++I +IE S + V S+N+ S WC EL +
Sbjct: 682 GKWIIDNIIDSIEKSHKTLFVLSENFVRSEWCKYELDF 719
>GLGB_CHLCV (Q823Y5) 1,4-alpha-glucan branching enzyme (EC 2.4.1.18)
(Glycogen branching enzyme) (BE)
(1,4-alpha-D-glucan:1,4-alpha-D-glucan
6-glucosyl-transferase)
Length = 719
Score = 32.0 bits (71), Expect = 2.0
Identities = 18/61 (29%), Positives = 29/61 (47%), Gaps = 2/61 (3%)
Query: 148 NISGWDLRDKPHHAELENIIEHINILGCKFPSRTKDLVEINYDVFVSFRGPDTRFNFTDH 207
N W L D P+HA L + +N L C+ P K + ++V F+ DT+ N +
Sbjct: 577 NSLDWHLLDNPYHASLHKCVAKMNALYCELPYFWKGDHKQESFLWVDFK--DTKNNVISY 634
Query: 208 F 208
+
Sbjct: 635 Y 635
>WR52_ARATH (Q9FH83) Probable WRKY transcription factor 52 (WRKY
DNA-binding protein 52) (Disease resistance protein
RRS1) (Resistance to Ralstonia solanacearum 1 protein)
Length = 1378
Score = 30.0 bits (66), Expect = 7.6
Identities = 26/91 (28%), Positives = 41/91 (44%), Gaps = 12/91 (13%)
Query: 20 DTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIVVFSKNYASST 79
+ R++F HL AL+RK I D ++ + + + IE + V ++V N S
Sbjct: 18 EVRYSFVSHLSEALRRKGINNVVVDVDID--DLLFKESQAKIEKAGVSVMVLPGNCDPSE 75
Query: 80 WCLRELAYILNC----------SVLYGKRVL 100
L + A +L C SVLYG +L
Sbjct: 76 VWLDKFAKVLECQRNNKDQAVVSVLYGDSLL 106
>TOLL_DROME (P08953) Toll protein precursor
Length = 1097
Score = 30.0 bits (66), Expect = 7.6
Identities = 25/107 (23%), Positives = 44/107 (40%), Gaps = 5/107 (4%)
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFR---DDTKLKKGEFIAPGLFRAIEASQV 245
+D F+S+ D F D+ L+ G F+ + G I + R++ S+
Sbjct: 859 FDAFISYSHKDQSF-IEDYLVPQLEH-GPQKFQLCVHERDWLVGGHIPENIMRSVADSRR 916
Query: 246 YIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQK 292
I+V S+N+ S W E + G+ + V D +V+K
Sbjct: 917 TIIVLSQNFIKSEWARLEFRAAHRSALNEGRSRIIVIIYSDIGDVEK 963
>DHG_THEAC (P13203) Glucose 1-dehydrogenase (EC 1.1.1.47)
Length = 360
Score = 30.0 bits (66), Expect = 7.6
Identities = 32/111 (28%), Positives = 48/111 (42%), Gaps = 10/111 (9%)
Query: 97 KRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRFQDHSNMVQRRRETL--QLVGNISGWDL 154
K V+ F D+ V K+S +G+ GKR ++ E G G+D+
Sbjct: 158 KNVMKAFEVFDV--VSKRSIFFGDDSTLIGKRMV----IIGSGSEAFLYSFAGVDRGFDV 211
Query: 155 RDKPHHAELENIIEHINILGCKFPSRTKDLVEINYDVFVSFRG-PDTRFNF 204
H E EN ++ ++ G KF + KD+ E D+ V G P T F F
Sbjct: 212 TMVNRHDETENKLKIMDEFGVKFANYLKDMPE-KIDLLVDTSGDPTTTFKF 261
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.139 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,484,670
Number of Sequences: 164201
Number of extensions: 1570960
Number of successful extensions: 2705
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 2687
Number of HSP's gapped (non-prelim): 23
length of query: 309
length of database: 59,974,054
effective HSP length: 110
effective length of query: 199
effective length of database: 41,911,944
effective search space: 8340476856
effective search space used: 8340476856
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 65 (29.6 bits)
Medicago: description of AC146863.4


