

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.7 - phase: 0
(198 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
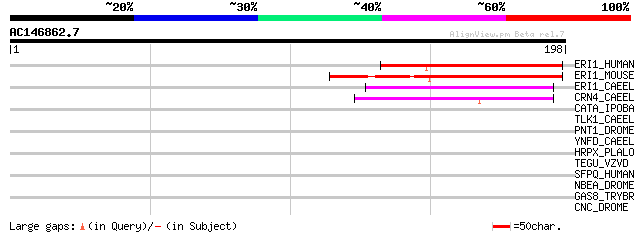
Score E
Sequences producing significant alignments: (bits) Value
ERI1_HUMAN (Q8IV48) 3'-5' exonuclease ERI1 (EC 3.1.-.-) (Eri-1 h... 69 5e-12
ERI1_MOUSE (Q7TMF2) 3'-5' exonuclease ERI1 (EC 3.1.-.-) (Eri-1 h... 66 6e-11
ERI1_CAEEL (O44406) 3'-5' exonuclease eri-1 (EC 3.1.-.-) (Enhanc... 56 5e-08
CRN4_CAEEL (Q10905) Cell death-related nuclease 4 precursor (EC ... 47 3e-05
CATA_IPOBA (P07145) Catalase (EC 1.11.1.6) 31 2.1
TLK1_CAEEL (P34314) Serine/threonine-protein kinase tousled-like... 30 2.8
PNT1_DROME (P51022) ETS-like protein pointed, isoform P1 (D-ETS-2) 30 3.6
YNFD_CAEEL (Q21828) Hypothetical UPF0041 protein R07E5.13 in chr... 29 6.2
HRPX_PLALO (P04929) Histidine-rich glycoprotein precursor 29 6.2
TEGU_VZVD (P09278) Large tegument protein 29 8.1
SFPQ_HUMAN (P23246) Splicing factor, proline-and glutamine-rich ... 29 8.1
NBEA_DROME (Q9W4E2) Neurobeachin protein (Rugose protein) (A-kin... 29 8.1
GAS8_TRYBR (O15697) Trypanin (T lymphocyte-triggering factor) (T... 29 8.1
CNC_DROME (P20482) Segmentation protein cap'n'collar 29 8.1
>ERI1_HUMAN (Q8IV48) 3'-5' exonuclease ERI1 (EC 3.1.-.-) (Eri-1
homolog) (Protein 3'hExo)
Length = 348
Score = 69.3 bits (168), Expect = 5e-12
Identities = 36/66 (54%), Positives = 45/66 (67%), Gaps = 1/66 (1%)
Query: 133 VIDFEATCDKDKNPH-PQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDFCKDLTG 191
+IDFEATC++ P EIIEFP V++++ T ++E FQ YVRP N LSDFC LTG
Sbjct: 131 IIDFEATCEEGNPPEFVHEIIEFPVVLLNTHTLEIEDTFQQYVRPEINTQLSDFCISLTG 190
Query: 192 IQQIQV 197
I Q QV
Sbjct: 191 ITQDQV 196
>ERI1_MOUSE (Q7TMF2) 3'-5' exonuclease ERI1 (EC 3.1.-.-) (Eri-1
homolog)
Length = 345
Score = 65.9 bits (159), Expect = 6e-11
Identities = 38/85 (44%), Positives = 53/85 (61%), Gaps = 5/85 (5%)
Query: 115 MVSQGYPREQYQEFQNFVVIDFEATCDKDKNPHP--QEIIEFPSVIVSSVTGQLEACFQT 172
M+ + + Y ++ +IDFEATC++ NP EIIEFP V++++ T ++E FQ
Sbjct: 112 MLKESSAGDSYYDY--ICIIDFEATCEEG-NPAEFLHEIIEFPVVLLNTHTLEIEDTFQQ 168
Query: 173 YVRPTCNQHLSDFCKDLTGIQQIQV 197
YVRP N LS+FC LTGI Q QV
Sbjct: 169 YVRPEVNAQLSEFCIGLTGITQDQV 193
>ERI1_CAEEL (O44406) 3'-5' exonuclease eri-1 (EC 3.1.-.-) (Enhanced
RNAi protein)
Length = 582
Score = 56.2 bits (134), Expect = 5e-08
Identities = 28/67 (41%), Positives = 39/67 (57%)
Query: 128 FQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDFCK 187
F + IDFE TC + +P EIIE P+V++ ++ + F+TYVRP N LS+FC
Sbjct: 148 FDYLIAIDFECTCVEIIYDYPHEIIELPAVLIDVREMKIISEFRTYVRPVRNPKLSEFCM 207
Query: 188 DLTGIQQ 194
T I Q
Sbjct: 208 QFTKIAQ 214
>CRN4_CAEEL (Q10905) Cell death-related nuclease 4 precursor (EC
3.1.-.-)
Length = 298
Score = 47.0 bits (110), Expect = 3e-05
Identities = 24/73 (32%), Positives = 37/73 (49%), Gaps = 2/73 (2%)
Query: 124 QYQEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQL--EACFQTYVRPTCNQH 181
Q+ F +++DFE T D +P E+I+F V ++ + F YV+P N+
Sbjct: 4 QHCPFDTLLILDFETTSDAANQDYPCEVIQFAIVAYDVPNDKIREDISFNKYVKPVLNRT 63
Query: 182 LSDFCKDLTGIQQ 194
L+ C D TGI Q
Sbjct: 64 LTKNCVDFTGIPQ 76
>CATA_IPOBA (P07145) Catalase (EC 1.11.1.6)
Length = 492
Score = 30.8 bits (68), Expect = 2.1
Identities = 22/89 (24%), Positives = 38/89 (41%), Gaps = 2/89 (2%)
Query: 81 HFHPHKMQQCQMNAFENHFYPHPVENQFQYAPINMVSQGYPREQYQEFQNFVVIDFEATC 140
+F KM Q ++ A+ + + H + + P+N + Y + NFV D E
Sbjct: 334 YFSDDKMLQARVFAYADT-HRHRLGPNYMLLPVNAPKCAHHNNSYDGYMNFVHRDEEVDY 392
Query: 141 DKDKNPHPQEIIEFPSVIVSSVTGQLEAC 169
K + + FP+ + VTGQ + C
Sbjct: 393 FPSKFDNTRNAERFPTPL-RIVTGQRDKC 420
>TLK1_CAEEL (P34314) Serine/threonine-protein kinase tousled-like 1
(EC 2.7.1.37) (Tousled-like kinase 1)
Length = 965
Score = 30.4 bits (67), Expect = 2.8
Identities = 16/78 (20%), Positives = 30/78 (37%)
Query: 50 NHRLGSEFLEPSNEFHNKPTYHHDYSTWTACHFHPHKMQQCQMNAFENHFYPHPVENQFQ 109
++ G + P + HH + H H +MQQ + + + ++
Sbjct: 140 DYNSGVHHMHPHQMQMQQQQQHHQQQYNMSYHNHQQQMQQMHYHQQQQQYQQQQAQHHQM 199
Query: 110 YAPINMVSQGYPREQYQE 127
YAP Q P++Q Q+
Sbjct: 200 YAPQIQQQQQQPQQQSQQ 217
Score = 29.3 bits (64), Expect = 6.2
Identities = 16/54 (29%), Positives = 22/54 (40%), Gaps = 3/54 (5%)
Query: 81 HFHPHKMQQCQMNAFENHFYP---HPVENQFQYAPINMVSQGYPREQYQEFQNF 131
H HPH+MQ Q Y H + Q Q + Q Y ++Q Q Q +
Sbjct: 147 HMHPHQMQMQQQQQHHQQQYNMSYHNHQQQMQQMHYHQQQQQYQQQQAQHHQMY 200
>PNT1_DROME (P51022) ETS-like protein pointed, isoform P1 (D-ETS-2)
Length = 623
Score = 30.0 bits (66), Expect = 3.6
Identities = 20/79 (25%), Positives = 36/79 (45%), Gaps = 9/79 (11%)
Query: 49 PNHRLGSEFLEPSNEFHNKPTYHHDYSTWTACHFHPHKMQQCQMNAFENHFYPHPVENQF 108
PN+ + + E +H +HH + H HPH Q Q +H H ++ Q
Sbjct: 55 PNYNMVLQSYENYPSYHLAQHHHHHHHH----HSHPHH-QLAQQTGLHSH---HTMQQQL 106
Query: 109 QYAPINMVSQGYPREQYQE 127
Q +++ Q +P++Q Q+
Sbjct: 107 QQQQQSLLLQ-HPQQQQQQ 124
>YNFD_CAEEL (Q21828) Hypothetical UPF0041 protein R07E5.13 in
chromosome III
Length = 160
Score = 29.3 bits (64), Expect = 6.2
Identities = 13/28 (46%), Positives = 15/28 (53%)
Query: 79 ACHFHPHKMQQCQMNAFENHFYPHPVEN 106
ACHF Q Q+ F NH Y H VE+
Sbjct: 84 ACHFANFSAQGAQLGRFVNHNYLHYVED 111
>HRPX_PLALO (P04929) Histidine-rich glycoprotein precursor
Length = 351
Score = 29.3 bits (64), Expect = 6.2
Identities = 19/79 (24%), Positives = 34/79 (42%), Gaps = 8/79 (10%)
Query: 38 PGNHPSGNVPEPNHRLG--SEFLEPSNEFHNKPTYHHDYSTWTACHFHPHKMQQCQMNAF 95
P +H + P +H LG ++ H++ +HH + A H H H+ +A
Sbjct: 100 PHHHHHHHPPHHHHHLGHHHHHHHAAHHHHHEEHHHHHH----AAHHHHHEEHHHHHHAA 155
Query: 96 ENH--FYPHPVENQFQYAP 112
+H F+ H + +AP
Sbjct: 156 HHHPWFHHHHLGYHHHHAP 174
Score = 29.3 bits (64), Expect = 6.2
Identities = 19/65 (29%), Positives = 25/65 (38%), Gaps = 11/65 (16%)
Query: 41 HPSGNVPEPNHRLGSEFLEPSNE--FHNKPTYHHDYSTWTACHFHPHKMQQCQMNAFENH 98
HP E +H E EP +E H+ P HH+ H HPH +H
Sbjct: 53 HPEHLHEEHHHHHPEEHHEPHHEEHHHHHPEEHHEPHHEEHHHHHPHP---------HHH 103
Query: 99 FYPHP 103
+ HP
Sbjct: 104 HHHHP 108
>TEGU_VZVD (P09278) Large tegument protein
Length = 2763
Score = 28.9 bits (63), Expect = 8.1
Identities = 19/64 (29%), Positives = 24/64 (36%), Gaps = 1/64 (1%)
Query: 45 NVPEPNHRLGSEFLEPSNEFHNKPTYHH-DYSTWTACHFHPHKMQQCQMNAFENHFYPHP 103
N P+ N L F +P E + PT Y TW K Q + + HP
Sbjct: 2548 NNPQQNQTLNQAFSKPILEITSIPTDDSISYRTWIEKSNQTQKRHQNDPRMYNSKTVFHP 2607
Query: 104 VENQ 107
V NQ
Sbjct: 2608 VNNQ 2611
>SFPQ_HUMAN (P23246) Splicing factor, proline-and glutamine-rich
(Polypyrimidine tract-binding protein-associated
splicing factor) (PTB-associated splicing factor) (PSF)
(DNA-binding p52/p100 complex, 100 kDa subunit)
Length = 707
Score = 28.9 bits (63), Expect = 8.1
Identities = 20/62 (32%), Positives = 24/62 (38%), Gaps = 6/62 (9%)
Query: 13 QGKGPPFTFQCNGSSMEVFPELNNEPGNHPSGNVPEPNHRLGSEFLEPSNEFHNKPTYHH 72
QG GP G M P+ PG G P+P HR G EP + P YH
Sbjct: 200 QGPGPGGP---KGGKMPGGPKPGGGPGLSTPGGHPKPPHRGGG---EPRGGRQHHPPYHQ 253
Query: 73 DY 74
+
Sbjct: 254 QH 255
>NBEA_DROME (Q9W4E2) Neurobeachin protein (Rugose protein) (A-kinase
anchor protein 550) (AKAP 550) (dAKAP550)
Length = 3584
Score = 28.9 bits (63), Expect = 8.1
Identities = 18/83 (21%), Positives = 37/83 (43%), Gaps = 14/83 (16%)
Query: 47 PEPNHRLGSEFLEPSNEFHNKPTYHHDYSTWTACHFHPHKMQQCQMNAFENHFYPHPVEN 106
P+ +H + +FL S++ HN H + QQ Q + + H+Y +
Sbjct: 1987 PQWSHHVYPQFLPESHQNHNSNMQHQ------------QQQQQQQQHQQQQHYYQQQQQQ 2034
Query: 107 Q--FQYAPINMVSQGYPREQYQE 127
Q ++ +M + Y ++Q+Q+
Sbjct: 2035 QVVHNHSHHHMTAAYYQQQQHQQ 2057
>GAS8_TRYBR (O15697) Trypanin (T lymphocyte-triggering factor)
(TLTF)
Length = 453
Score = 28.9 bits (63), Expect = 8.1
Identities = 30/134 (22%), Positives = 47/134 (34%), Gaps = 22/134 (16%)
Query: 1 MQITCEASLKCLQGKGPPFTFQCNGSSMEVFPELNNEPGNHPSGNVPEPNHRLGSEFLEP 60
+Q++CEA LK L+ + EL H N + + +
Sbjct: 182 LQVSCEAKLKVLRDE----------------LELRRRAEIHEIEE--RKNEHINALIKQH 223
Query: 61 SNEFHNKPTYHHDYSTWTACHFHPHKMQQCQMNAFENH----FYPHPVENQFQYAPINMV 116
+FH TY++ +T H K + QM + H Y ENQ AP+
Sbjct: 224 EEKFHEMKTYYNQITTNNLEIIHSLKEEIAQMKQNDEHNETLMYDIDRENQNLVAPLEEA 283
Query: 117 SQGYPREQYQEFQN 130
+ Q + QN
Sbjct: 284 QREVAELQQKRKQN 297
>CNC_DROME (P20482) Segmentation protein cap'n'collar
Length = 1383
Score = 28.9 bits (63), Expect = 8.1
Identities = 22/79 (27%), Positives = 33/79 (40%), Gaps = 8/79 (10%)
Query: 22 QCNGSSMEV--FPELNNEPGNHPSGNVPEPNHRLGSEFLEPSNEFHNKPTYHHDYSTWTA 79
+ +G S V F EL N G+ P ++ E + E L+ + P+YHH +
Sbjct: 543 KADGDSFSVSDFEELQNSVGS-PLFDLDEDAKKELDEMLQSA-----VPSYHHPHPHHGH 596
Query: 80 CHFHPHKMQQCQMNAFENH 98
H HPH M+ H
Sbjct: 597 PHAHPHSHHHASMHHAHAH 615
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.135 0.441
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,298,170
Number of Sequences: 164201
Number of extensions: 1234024
Number of successful extensions: 2203
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 2168
Number of HSP's gapped (non-prelim): 37
length of query: 198
length of database: 59,974,054
effective HSP length: 105
effective length of query: 93
effective length of database: 42,732,949
effective search space: 3974164257
effective search space used: 3974164257
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146862.7


