

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.23 + phase: 0 /pseudo
(320 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
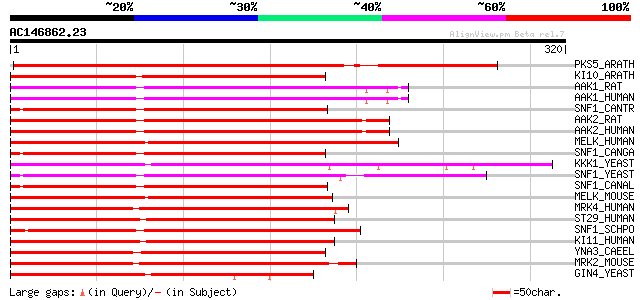
Score E
Sequences producing significant alignments: (bits) Value
PKS5_ARATH (O22932) SOS2-like protein kinase PKS5 (EC 2.7.1.37) 258 2e-68
KI10_ARATH (Q38997) SNF1-related protein kinase KIN10 (EC 2.7.1.... 186 7e-47
AAK1_RAT (P54645) 5'-AMP-activated protein kinase, catalytic alp... 184 3e-46
AAK1_HUMAN (Q13131) 5'-AMP-activated protein kinase, catalytic a... 182 7e-46
SNF1_CANTR (O94168) Carbon catabolite derepressing protein kinas... 182 1e-45
AAK2_RAT (Q09137) 5'-AMP-activated protein kinase, catalytic alp... 182 1e-45
AAK2_HUMAN (P54646) 5'-AMP-activated protein kinase, catalytic a... 182 1e-45
MELK_HUMAN (Q14680) Maternal embryonic leucine zipper kinase (EC... 181 2e-45
SNF1_CANGA (Q00372) Carbon catabolite derepressing protein kinas... 181 3e-45
KKK1_YEAST (P34244) Probable serine/threonine-protein kinase YKL... 181 3e-45
SNF1_YEAST (P06782) Carbon catabolite derepressing protein kinas... 180 5e-45
SNF1_CANAL (P52497) Carbon catabolite derepressing protein kinas... 180 5e-45
MELK_MOUSE (Q61846) Maternal embryonic leucine zipper kinase (EC... 176 5e-44
MRK4_HUMAN (Q96L34) MAP/microtubule affinity-regulating kinase 4... 175 1e-43
ST29_HUMAN (Q8IWQ3) Serine/threonine-protein kinase 29 (EC 2.7.1... 174 3e-43
SNF1_SCHPO (O74536) SNF1-like protein kinase ssp2 (EC 2.7.1.-) 174 3e-43
KI11_HUMAN (Q8TDC3) Probable serine/threonine-protein kinase KIA... 172 8e-43
YNA3_CAEEL (P45894) Putative serine/threonine-protein kinase PAR... 171 2e-42
MRK2_MOUSE (Q05512) MAP/microtubule affinity-regulating kinase 2... 168 1e-41
GIN4_YEAST (Q12263) Serine/threonine-protein kinase GIN4 (EC 2.7... 168 1e-41
>PKS5_ARATH (O22932) SOS2-like protein kinase PKS5 (EC 2.7.1.37)
Length = 435
Score = 258 bits (658), Expect = 2e-68
Identities = 132/282 (46%), Positives = 189/282 (66%), Gaps = 18/282 (6%)
Query: 3 LEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAKGN 62
+E+V G ELF+KI+ G+L E R+ FQQLI V YCH++GV+HRDLK EN+L+D GN
Sbjct: 99 MEFVKGGELFNKISKHGRLSEDLSRRYFQQLISAVGYCHARGVYHRDLKPENLLIDENGN 158
Query: 63 LKITDFGLSALPQQFRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYVVLT 122
LK++DFGLSAL Q R DGLLHT CG+ YVAPEIL+ +GY GA DVWSCG++L+V++
Sbjct: 159 LKVSDFGLSALTDQIRPDGLLHTLCGTPAYVAPEILSKKGYEGAKVDVWSCGIVLFVLVA 218
Query: 123 GFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDLWF- 181
G+LPF+D N+ +Y+K+ KG+++ P+W+S + + R+LD NP+TRIT+ EI +D WF
Sbjct: 219 GYLPFNDPNVMNMYKKIYKGEYRFPRWMSPDLKRFVSRLLDINPETRITIDEILKDPWFV 278
Query: 182 KEGYNQ-AYHEDEEDGVYVDDQAFNIHELSHEEEKSSSQSPIRINAFQLIGMSSCLDLSG 240
+ G+ Q +H+DE ++DQ + +SS ++ +NAF LI SS LDLSG
Sbjct: 279 RGGFKQIKFHDDE-----IEDQ----------KVESSLEAVKSLNAFDLISYSSGLDLSG 323
Query: 241 FFEK-EDVSERKIRFTANLSAKVLMEKIEDSVTEMEFRVQKK 281
F + S RF + S ++L E++E E R++KK
Sbjct: 324 LFAGCSNSSGESERFLSEKSPEMLAEEVEGFAREENLRMKKK 365
>KI10_ARATH (Q38997) SNF1-related protein kinase KIN10 (EC 2.7.1.-)
(AKIN10)
Length = 535
Score = 186 bits (472), Expect = 7e-47
Identities = 92/182 (50%), Positives = 122/182 (66%), Gaps = 3/182 (1%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
+V+EYV ELFD I KG+L E E R FQQ+I GV YCH V HRDLK EN+L+D+K
Sbjct: 117 LVMEYVNSGELFDYIVEKGRLQEDEARNFFQQIISGVEYCHRNMVVHRDLKPENLLLDSK 176
Query: 61 GNLKITDFGLSALPQQFRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYVV 120
N+KI DFGLS + R L T+CGS NY APE+++ + Y G DVWSCGVILY +
Sbjct: 177 CNVKIADFGLSNI---MRDGHFLKTSCGSPNYAAPEVISGKLYAGPEVDVWSCGVILYAL 233
Query: 121 LTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDLW 180
L G LPFDD N+ L++K+ G + P LS GA+++I R+L +P R+T+ EI++ W
Sbjct: 234 LCGTLPFDDENIPNLFKKIKGGIYTLPSHLSPGARDLIPRMLVVDPMKRVTIPEIRQHPW 293
Query: 181 FK 182
F+
Sbjct: 294 FQ 295
>AAK1_RAT (P54645) 5'-AMP-activated protein kinase, catalytic
alpha-1 chain (EC 2.7.1.-) (AMPK alpha-1 chain)
Length = 548
Score = 184 bits (467), Expect = 3e-46
Identities = 106/242 (43%), Positives = 145/242 (59%), Gaps = 17/242 (7%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
MV+EYV+G ELFD I G+L E E R+LFQQ++ GV YCH V HRDLK ENVL+DA
Sbjct: 91 MVMEYVSGGELFDYICKNGRLDEKESRRLFQQILSGVDYCHRHMVVHRDLKPENVLLDAH 150
Query: 61 GNLKITDFGLSALPQQFRADG-LLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYV 119
N KI DFGLS + +DG L T+CGS NY APE+++ R Y G D+WS GVILY
Sbjct: 151 MNAKIADFGLSNM----MSDGEFLRTSCGSPNYAAPEVISGRLYAGPEVDIWSSGVILYA 206
Query: 120 VLTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDL 179
+L G LPFDD ++ L++K+ G F P++L+ +++K +L +P R T+ +I+E
Sbjct: 207 LLCGTLPFDDDHVPTLFKKICDGIFYTPQYLNPSVISLLKHMLQVDPMKRATIKDIREHE 266
Query: 180 WFKEGY-NQAYHEDEE-DGVYVDDQAF----NIHELSHEEEKS-----SSQSPIRINAFQ 228
WFK+ + ED +DD+A E S EE S + Q P+ + A+
Sbjct: 267 WFKQDLPKYLFPEDPSYSSTMIDDEALKEVCEKFECSEEEVLSCLYNRNHQDPLAV-AYH 325
Query: 229 LI 230
LI
Sbjct: 326 LI 327
>AAK1_HUMAN (Q13131) 5'-AMP-activated protein kinase, catalytic
alpha-1 chain (EC 2.7.1.-) (AMPK alpha-1 chain)
Length = 550
Score = 182 bits (463), Expect = 7e-46
Identities = 105/242 (43%), Positives = 145/242 (59%), Gaps = 17/242 (7%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
MV+EYV+G ELFD I G+L E E R+LFQQ++ GV YCH V HRDLK ENVL+DA
Sbjct: 93 MVMEYVSGGELFDYICKNGRLDEKESRRLFQQILSGVDYCHRHMVVHRDLKPENVLLDAH 152
Query: 61 GNLKITDFGLSALPQQFRADG-LLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYV 119
N KI DFGLS + +DG L T+CGS NY APE+++ R Y G D+WS GVILY
Sbjct: 153 MNAKIADFGLSNM----MSDGEFLRTSCGSPNYAAPEVISGRLYAGPEVDIWSSGVILYA 208
Query: 120 VLTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDL 179
+L G LPFDD ++ L++K+ G F P++L+ +++K +L +P R ++ +I+E
Sbjct: 209 LLCGTLPFDDDHVPTLFKKICDGIFYTPQYLNPSVISLLKHMLQVDPMKRASIKDIREHE 268
Query: 180 WFKEGY-NQAYHEDEE-DGVYVDDQAF----NIHELSHEEEKS-----SSQSPIRINAFQ 228
WFK+ + ED +DD+A E S EE S + Q P+ + A+
Sbjct: 269 WFKQDLPKYLFPEDPSYSSTMIDDEALKEVCEKFECSEEEVLSCLYNRNHQDPLAV-AYH 327
Query: 229 LI 230
LI
Sbjct: 328 LI 329
>SNF1_CANTR (O94168) Carbon catabolite derepressing protein kinase
(EC 2.7.1.-)
Length = 619
Score = 182 bits (462), Expect = 1e-45
Identities = 93/184 (50%), Positives = 125/184 (67%), Gaps = 6/184 (3%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
MV+E+ G+ELFD I +GK+PE E R+ FQQ+I V YCH + HRDLK EN+L+D +
Sbjct: 127 MVIEFA-GKELFDYIVQRGKMPEDEARRFFQQIIAAVEYCHRHKIVHRDLKPENLLLDDQ 185
Query: 61 GNLKITDFGLSALPQQFRADG-LLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYV 119
N+KI DFGLS + DG L T+CGS NY APE+++ + Y G DVWS GVILYV
Sbjct: 186 LNVKIADFGLSNI----MTDGNFLKTSCGSPNYAAPEVISGKLYAGPEVDVWSSGVILYV 241
Query: 120 VLTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDL 179
+L G LPFDD + L++K+ G + P +LS GA++++ R+L NP RIT+ EI ED
Sbjct: 242 MLCGRLPFDDEFIPALFKKISNGVYTLPNYLSPGAKHLLTRMLVVNPLNRITIHEIMEDE 301
Query: 180 WFKE 183
WFK+
Sbjct: 302 WFKQ 305
>AAK2_RAT (Q09137) 5'-AMP-activated protein kinase, catalytic
alpha-2 chain (EC 2.7.1.-) (AMPK alpha-2 chain)
Length = 552
Score = 182 bits (462), Expect = 1e-45
Identities = 98/222 (44%), Positives = 140/222 (62%), Gaps = 9/222 (4%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
MV+EYV+G ELFD I G++ E E R+LFQQ++ V YCH V HRDLK ENVL+DA+
Sbjct: 91 MVMEYVSGGELFDYICKHGRVEEVEARRLFQQILSAVDYCHRHMVVHRDLKPENVLLDAQ 150
Query: 61 GNLKITDFGLSALPQQFRADG-LLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYV 119
N KI DFGLS + +DG L T+CGS NY APE+++ R Y G D+WSCGVILY
Sbjct: 151 MNAKIADFGLSNM----MSDGEFLRTSCGSPNYAAPEVISGRLYAGPEVDIWSCGVILYA 206
Query: 120 VLTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDL 179
+L G LPFDD ++ L++K+ G F P++L+ ++ +L +P R T+ +I+E
Sbjct: 207 LLCGTLPFDDEHVPTLFKKIRGGVFYIPEYLNRSIATLLMHMLQVDPLKRATIKDIREHE 266
Query: 180 WFKEGY-NQAYHEDEE-DGVYVDDQAFNIHELSHEEEKSSSQ 219
WFK+ + + ED D +DD+A + E+ + E + S+
Sbjct: 267 WFKQDLPSYLFPEDPSYDANVIDDEA--VKEVCEKFECTESE 306
>AAK2_HUMAN (P54646) 5'-AMP-activated protein kinase, catalytic
alpha-2 chain (EC 2.7.1.-) (AMPK alpha-2 chain)
Length = 552
Score = 182 bits (461), Expect = 1e-45
Identities = 98/222 (44%), Positives = 139/222 (62%), Gaps = 9/222 (4%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
MV+EYV+G ELFD I G++ E E R+LFQQ++ V YCH V HRDLK ENVL+DA
Sbjct: 91 MVMEYVSGGELFDYICKHGRVEEMEARRLFQQILSAVDYCHRHMVVHRDLKPENVLLDAH 150
Query: 61 GNLKITDFGLSALPQQFRADG-LLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYV 119
N KI DFGLS + +DG L T+CGS NY APE+++ R Y G D+WSCGVILY
Sbjct: 151 MNAKIADFGLSNM----MSDGEFLRTSCGSPNYAAPEVISGRLYAGPEVDIWSCGVILYA 206
Query: 120 VLTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDL 179
+L G LPFDD ++ L++K+ G F P++L+ ++ +L +P R T+ +I+E
Sbjct: 207 LLCGTLPFDDEHVPTLFKKIRGGVFYIPEYLNRSVATLLMHMLQVDPLKRATIKDIREHE 266
Query: 180 WFKEGY-NQAYHEDEE-DGVYVDDQAFNIHELSHEEEKSSSQ 219
WFK+ + + ED D +DD+A + E+ + E + S+
Sbjct: 267 WFKQDLPSYLFPEDPSYDANVIDDEA--VKEVCEKFECTESE 306
>MELK_HUMAN (Q14680) Maternal embryonic leucine zipper kinase (EC
2.7.1.37) (hMELK) (Protein kinase PK38) (hPK38)
Length = 651
Score = 181 bits (459), Expect = 2e-45
Identities = 90/225 (40%), Positives = 137/225 (60%), Gaps = 2/225 (0%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
MVLEY G ELFD I S+ +L E E R +F+Q++ V+Y HS+G HRDLK EN+L D
Sbjct: 84 MVLEYCPGGELFDYIISQDRLSEEETRVVFRQIVSAVAYVHSQGYAHRDLKPENLLFDEY 143
Query: 61 GNLKITDFGLSALPQQFRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYVV 120
LK+ DFGL A P+ + D L T CGS Y APE++ + Y G+ +DVWS G++LYV+
Sbjct: 144 HKLKLIDFGLCAKPKGNK-DYHLQTCCGSLAYAAPELIQGKSYLGSEADVWSMGILLYVL 202
Query: 121 LTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDLW 180
+ GFLPFDD N+ LY+K+++G + PKWLS + +++++L +PK RI+M + W
Sbjct: 203 MCGFLPFDDDNVMALYKKIMRGKYDVPKWLSPSSILLLQQMLQVDPKKRISMKNLLNHPW 262
Query: 181 FKEGYNQAYH-EDEEDGVYVDDQAFNIHELSHEEEKSSSQSPIRI 224
+ YN + + +++DD + H + + + I +
Sbjct: 263 IMQDYNYPVEWQSKNPFIHLDDDCVTELSVHHRNNRQTMEDLISL 307
>SNF1_CANGA (Q00372) Carbon catabolite derepressing protein kinase
(EC 2.7.1.-)
Length = 612
Score = 181 bits (458), Expect = 3e-45
Identities = 93/183 (50%), Positives = 122/183 (65%), Gaps = 6/183 (3%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
MV+EY G ELFD I + K+ E E R+ FQQ+I V YCH + HRDLK EN+L+D
Sbjct: 114 MVIEYA-GNELFDYIVQRNKMSEQEARRFFQQIISAVEYCHRHKIVHRDLKPENLLLDEH 172
Query: 61 GNLKITDFGLSALPQQFRADG-LLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYV 119
N+KI DFGLS + DG L T+CGS NY APE+++ + Y G DVWSCGVILYV
Sbjct: 173 LNVKIADFGLSNI----MTDGNFLKTSCGSPNYAAPEVISGKLYAGPEVDVWSCGVILYV 228
Query: 120 VLTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDL 179
+L LPFDD ++ VL++ + G + PK+LS GA ++IKR+L NP RI++ EI +D
Sbjct: 229 MLCRRLPFDDESIPVLFKNISNGVYTLPKFLSPGASDLIKRMLIVNPLNRISIHEIMQDE 288
Query: 180 WFK 182
WFK
Sbjct: 289 WFK 291
>KKK1_YEAST (P34244) Probable serine/threonine-protein kinase
YKL101W (EC 2.7.1.37)
Length = 1518
Score = 181 bits (458), Expect = 3e-45
Identities = 123/338 (36%), Positives = 180/338 (52%), Gaps = 28/338 (8%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
+VLEYV G ELFD + SKGKLPE E F+Q+++GVSYCHS + HRDLK EN+L+D K
Sbjct: 191 LVLEYVDGGELFDYLVSKGKLPEREAIHYFKQIVEGVSYCHSFNICHRDLKPENLLLDKK 250
Query: 61 GN-LKITDFGLSALPQQFRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYV 119
+KI DFG++AL + LL T+CGS +Y +PEI+ R Y+G SDVWSCG++L+
Sbjct: 251 NRRIKIADFGMAALELPNK---LLKTSCGSPHYASPEIVMGRPYHGGPSDVWSCGIVLFA 307
Query: 120 VLTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDL 179
+LTG LPF+D N+ L KV G +Q P LS+ A+++I +IL +P+ RIT EI +
Sbjct: 308 LLTGHLPFNDDNIKKLLLKVQSGKYQMPSNLSSEARDLISKILVIDPEKRITTQEILKHP 367
Query: 180 WFKE----GYNQAYHEDEEDGVYVDDQAFNIHELSH--------EEEKSSSQSPIRINAF 227
K+ N+ + +D + ++H L++ + +S +R
Sbjct: 368 LIKKYDDLPVNKVLRKMRKDNMARGKSNSDLHLLNNVSPSIVTLHSKGEIDESILRSLQI 427
Query: 228 QLIGMSSCLDLSGFFEKEDVSER---------KIRFTANLSAKVLMEK--IEDSVTEMEF 276
G+S L + +K E+ K R + +LS+ +K E SV E
Sbjct: 428 LWHGVSRELITAKLLQKPMSEEKLFYSLLLQYKQRHSISLSSSSENKKSATESSVNEPRI 487
Query: 277 RVQKKN-GKMKVMQENKDHKTLGSLSVIIEVMDIQNES 313
K + EN D KTL SL + E N++
Sbjct: 488 EYASKTANNTGLRSENNDVKTLHSLEIHSEDTSTVNQN 525
>SNF1_YEAST (P06782) Carbon catabolite derepressing protein kinase
(EC 2.7.1.-)
Length = 633
Score = 180 bits (456), Expect = 5e-45
Identities = 108/283 (38%), Positives = 155/283 (54%), Gaps = 23/283 (8%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
MV+EY G ELFD I + K+ E E R+ FQQ+I V YCH + HRDLK EN+L+D
Sbjct: 130 MVIEYA-GNELFDYIVQRDKMSEQEARRFFQQIISAVEYCHRHKIVHRDLKPENLLLDEH 188
Query: 61 GNLKITDFGLSALPQQFRADG-LLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYV 119
N+KI DFGLS + DG L T+CGS NY APE+++ + Y G DVWSCGVILYV
Sbjct: 189 LNVKIADFGLSNI----MTDGNFLKTSCGSPNYAAPEVISGKLYAGPEVDVWSCGVILYV 244
Query: 120 VLTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDL 179
+L LPFDD ++ VL++ + G + PK+LS GA +IKR+L NP RI++ EI +D
Sbjct: 245 MLCRRLPFDDESIPVLFKNISNGVYTLPKFLSPGAAGLIKRMLIVNPLNRISIHEIMQDD 304
Query: 180 WFKEGYNQAY-------HEDEEDGVYVDDQAFNIHELSHEEEKSSSQSPIRINAFQLIGM 232
WFK + H +EE N + S ++ S I N ++
Sbjct: 305 WFKVDLPEYLLPPDLKPHPEEE----------NENNDSKKDGSSPDNDEIDDNLVNILSS 354
Query: 233 SSCLDLSGFFEKEDVSERKIRFTANLSAKVLMEKIEDSVTEME 275
+ + +E + SE F A +L+++ + + +M+
Sbjct: 355 TMGYEKDEIYESLESSEDTPAFNEIRDAYMLIKENKSLIKDMK 397
>SNF1_CANAL (P52497) Carbon catabolite derepressing protein kinase
(EC 2.7.1.-)
Length = 620
Score = 180 bits (456), Expect = 5e-45
Identities = 94/185 (50%), Positives = 127/185 (67%), Gaps = 7/185 (3%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
MV+E+ G+ELFD I +GK+PE E R+ FQQ+I V YCH + HRDLK EN+L+D +
Sbjct: 128 MVIEFA-GKELFDYIVQRGKMPEDEARRFFQQIIAAVEYCHRHKIVHRDLKPENLLLDDQ 186
Query: 61 GNLKITDFGLSALPQQFRADG-LLHTTCGSANYV-APEILANRGYNGASSDVWSCGVILY 118
N+KI DFGLS + DG L T+CGS NY+ APE+++ + Y G DVWS GVILY
Sbjct: 187 LNVKIADFGLSNI----MTDGNFLKTSCGSPNYMPAPEVISGKLYAGPEVDVWSAGVILY 242
Query: 119 VVLTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKED 178
V+L G LPFDD + L++K+ G + P +LSAGA++++ R+L NP RIT+ EI ED
Sbjct: 243 VMLCGRLPFDDEFIPALFKKISNGVYTLPNYLSAGAKHLLTRMLVVNPLNRITIHEIMED 302
Query: 179 LWFKE 183
WFK+
Sbjct: 303 DWFKQ 307
>MELK_MOUSE (Q61846) Maternal embryonic leucine zipper kinase (EC
2.7.1.37) (Protein kinase PK38) (mPK38)
Length = 643
Score = 176 bits (447), Expect = 5e-44
Identities = 85/186 (45%), Positives = 124/186 (65%), Gaps = 1/186 (0%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
MVLEY G ELFD I S+ +L E E R +F+Q++ V+Y HS+G HRDLK EN+L D
Sbjct: 84 MVLEYCPGGELFDYIISQDRLSEEETRVVFRQILSAVAYVHSQGYAHRDLKPENLLFDEN 143
Query: 61 GNLKITDFGLSALPQQFRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYVV 120
LK+ DFGL A P+ + D L T CGS Y APE++ + Y G+ +DVWS G++LYV+
Sbjct: 144 HKLKLIDFGLCAKPKGNK-DYHLQTCCGSLAYAAPELIQGKSYLGSEADVWSMGILLYVL 202
Query: 121 LTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDLW 180
+ GFLPFDD N+ LY+K+++G ++ PKWLS + +++++L +PK RI+M + W
Sbjct: 203 MCGFLPFDDDNVMALYKKIMRGKYEVPKWLSPSSILLLQQMLQVDPKKRISMRNLLNHPW 262
Query: 181 FKEGYN 186
+ Y+
Sbjct: 263 VMQDYS 268
>MRK4_HUMAN (Q96L34) MAP/microtubule affinity-regulating kinase 4
(EC 2.7.1.37) (MAP/microtubule affinity-regulating
kinase like 1)
Length = 752
Score = 175 bits (444), Expect = 1e-43
Identities = 85/199 (42%), Positives = 123/199 (61%), Gaps = 7/199 (3%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
+V+EY + E+FD + S G++ E E R F+Q++ V YCH K + HRDLK EN+L+DA+
Sbjct: 133 LVMEYASAGEVFDYLVSHGRMKEKEARAKFRQIVSAVHYCHQKNIVHRDLKAENLLLDAE 192
Query: 61 GNLKITDFGLSALPQQFRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYVV 120
N+KI DFG S +F L T CGS Y APE+ + Y+G D+WS GVILY +
Sbjct: 193 ANIKIADFGFS---NEFTLGSKLDTFCGSPPYAAPELFQGKKYDGPEVDIWSLGVILYTL 249
Query: 121 LTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDLW 180
++G LPFD NL L ++VL+G ++ P ++S ++I++R L NP R T+ +I +D W
Sbjct: 250 VSGSLPFDGHNLKELRERVLRGKYRVPFYMSTDCESILRRFLVLNPAKRCTLEQIMKDKW 309
Query: 181 FKEGYN----QAYHEDEED 195
GY + Y E EED
Sbjct: 310 INIGYEGEELKPYTEPEED 328
>ST29_HUMAN (Q8IWQ3) Serine/threonine-protein kinase 29 (EC
2.7.1.37) (BRSK2) (HUSSY-12)
Length = 736
Score = 174 bits (441), Expect = 3e-43
Identities = 80/187 (42%), Positives = 123/187 (64%), Gaps = 3/187 (1%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
+VLE+V+G ELFD + KG+L E RK F+Q+I + +CHS + HRDLK EN+L+D K
Sbjct: 93 LVLEHVSGGELFDYLVKKGRLTPKEARKFFRQIISALDFCHSHSICHRDLKPENLLLDEK 152
Query: 61 GNLKITDFGLSALPQQFRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYVV 120
N++I DFG+++L D LL T+CGS +Y PE++ Y+G +DVWSCGVIL+ +
Sbjct: 153 NNIRIADFGMASLQV---GDSLLETSCGSPHYACPEVIRGEKYDGRKADVWSCGVILFAL 209
Query: 121 LTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDLW 180
L G LPFDD NL L +KV +G F P ++ Q++++ + + + R+T+ I++ +W
Sbjct: 210 LVGALPFDDDNLRQLLEKVKRGVFHMPHFIPPDCQSLLRGMSEVDAARRLTLEHIQKHIW 269
Query: 181 FKEGYNQ 187
+ G N+
Sbjct: 270 YIGGKNE 276
>SNF1_SCHPO (O74536) SNF1-like protein kinase ssp2 (EC 2.7.1.-)
Length = 576
Score = 174 bits (441), Expect = 3e-43
Identities = 94/204 (46%), Positives = 126/204 (61%), Gaps = 7/204 (3%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
MV+EY GE LFD I K ++ E EGR+ FQQ+I + YCH + HRDLK EN+L+D
Sbjct: 109 MVIEYAGGE-LFDYIVEKKRMTEDEGRRFFQQIICAIEYCHRHKIVHRDLKPENLLLDDN 167
Query: 61 GNLKITDFGLSALPQQFRADG-LLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYV 119
N+KI DFGLS + DG L T+CGS NY APE++ + Y G DVWSCG++LYV
Sbjct: 168 LNVKIADFGLSNI----MTDGNFLKTSCGSPNYAAPEVINGKLYAGPEVDVWSCGIVLYV 223
Query: 120 VLTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDL 179
+L G LPFDD + L++KV + P +LS GAQ++I+R++ +P RIT+ EI+ D
Sbjct: 224 MLVGRLPFDDEFIPNLFKKVNSCVYVMPDFLSPGAQSLIRRMIVADPMQRITIQEIRRDP 283
Query: 180 WFKEGYNQAYHEDEE-DGVYVDDQ 202
WF EE G Y D +
Sbjct: 284 WFNVNLPDYLRPMEEVQGSYADSR 307
>KI11_HUMAN (Q8TDC3) Probable serine/threonine-protein kinase
KIAA1811 (EC 2.7.1.37)
Length = 794
Score = 172 bits (437), Expect = 8e-43
Identities = 77/187 (41%), Positives = 125/187 (66%), Gaps = 3/187 (1%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
+VLE+V+G ELFD + KG+L E RK F+Q++ + +CHS + HRDLK EN+L+D K
Sbjct: 124 LVLEHVSGGELFDYLVKKGRLTPKEARKFFRQIVSALDFCHSYSICHRDLKPENLLLDEK 183
Query: 61 GNLKITDFGLSALPQQFRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYVV 120
N++I DFG+++L D LL T+CGS +Y PE++ Y+G +D+WSCGVIL+ +
Sbjct: 184 NNIRIADFGMASLQV---GDSLLETSCGSPHYACPEVIKGEKYDGRRADMWSCGVILFAL 240
Query: 121 LTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDLW 180
L G LPFDD NL L +KV +G F P ++ Q++++ +++ P+ R+++ +I++ W
Sbjct: 241 LVGALPFDDDNLRQLLEKVKRGVFHMPHFIPPDCQSLLRGMIEVEPEKRLSLEQIQKHPW 300
Query: 181 FKEGYNQ 187
+ G ++
Sbjct: 301 YLGGKHE 307
>YNA3_CAEEL (P45894) Putative serine/threonine-protein kinase PAR2.3
(EC 2.7.1.37)
Length = 622
Score = 171 bits (433), Expect = 2e-42
Identities = 87/183 (47%), Positives = 116/183 (62%), Gaps = 5/183 (2%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
+V+E V+G ELF I KG LP E R+ FQQ+I GVSYCH+ + HRDLK EN+L+DA
Sbjct: 99 LVMELVSGGELFSYITRKGALPIRESRRYFQQIISGVSYCHNHMIVHRDLKPENLLLDAN 158
Query: 61 GNLKITDFGLSALPQQFRADG-LLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYV 119
N+KI DFGLS + DG LL T CGS NY APE+++N+ Y G D+WSCGVILY
Sbjct: 159 KNIKIADFGLS----NYMTDGDLLSTACGSPNYAAPELISNKLYVGPEVDLWSCGVILYA 214
Query: 120 VLTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDL 179
+L G LPFDD+N+ L+ K+ G + P + A ++I +L +P R + I
Sbjct: 215 MLCGTLPFDDQNVPTLFAKIKSGRYTVPYSMEKQAADLISTMLQVDPVKRADVKRIVNHS 274
Query: 180 WFK 182
WF+
Sbjct: 275 WFR 277
>MRK2_MOUSE (Q05512) MAP/microtubule affinity-regulating kinase 2
(EC 2.7.1.37) (Serine/threonine-protein kinase Emk)
Length = 774
Score = 168 bits (426), Expect = 1e-41
Identities = 85/200 (42%), Positives = 124/200 (61%), Gaps = 8/200 (4%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
+V+EY +G E+FD + + G++ E E R F+Q++ V YCH K + HRDLK EN+L+DA
Sbjct: 127 LVMEYASGGEVFDYLVAHGRMKEKEARAKFRQIVLHVQYCHQKFIVHRDLKAENLLLDAD 186
Query: 61 GNLKITDFGLSALPQQFRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYVV 120
N+KI DFG S +F L T CGS Y APE+ + +G DVWS GVILY +
Sbjct: 187 MNIKIADFGFS---NEFTFGNKLDTFCGSPPYAAPELFQGKKIDGPEVDVWSLGVILYTL 243
Query: 121 LTGFLPFDDRNLAVLYQKVLKGDFQKPKWLSAGAQNIIKRILDPNPKTRITMAEIKEDLW 180
++G LPFD +NL L ++VL+G ++ P ++S +N++K+ L NP R T+ +I +D W
Sbjct: 244 VSGSLPFDGQNLKELRERVLRGKYRIPFYMSTDCENLLKKFLILNPSKRGTLEQIMKDRW 303
Query: 181 FKEGYNQAYHEDEEDGVYVD 200
G HED+E YV+
Sbjct: 304 MNVG-----HEDDELKPYVE 318
>GIN4_YEAST (Q12263) Serine/threonine-protein kinase GIN4 (EC
2.7.1.37)
Length = 1142
Score = 168 bits (426), Expect = 1e-41
Identities = 89/179 (49%), Positives = 123/179 (67%), Gaps = 7/179 (3%)
Query: 1 MVLEYVTGEELFDKIASKGKLPEGEGRKLFQQLIDGVSYCHSKGVFHRDLKLENVLVDAK 60
+VLEY ELF+ + +G LPE E + F+Q+I GVSYCH+ G+ HRDLK EN+L+D K
Sbjct: 108 LVLEYAEKGELFNLLVERGPLPEHEAIRFFRQIIIGVSYCHALGIVHRDLKPENLLLDHK 167
Query: 61 GNLKITDFGLSALPQQFRADGLLHTTCGSANYVAPEILANRGYNGASSDVWSCGVILYVV 120
N+KI DFG++AL + + LL T+CGS +Y APEI++ Y G +SDVWSCGVIL+ +
Sbjct: 168 YNIKIADFGMAALETEGK---LLETSCGSPHYAAPEIVSGIPYQGFASDVWSCGVILFAL 224
Query: 121 LTGFLPFD--DRNLAVLYQKVLKGDFQKPK--WLSAGAQNIIKRILDPNPKTRITMAEI 175
LTG LPFD D N+ L KV KG+F+ P +S AQ++I++IL +P+ RI +I
Sbjct: 225 LTGRLPFDEEDGNIRTLLLKVQKGEFEMPSDDEISREAQDLIRKILTVDPERRIKTRDI 283
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.135 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,326,105
Number of Sequences: 164201
Number of extensions: 1605251
Number of successful extensions: 7984
Number of sequences better than 10.0: 1686
Number of HSP's better than 10.0 without gapping: 1512
Number of HSP's successfully gapped in prelim test: 174
Number of HSP's that attempted gapping in prelim test: 4445
Number of HSP's gapped (non-prelim): 1864
length of query: 320
length of database: 59,974,054
effective HSP length: 110
effective length of query: 210
effective length of database: 41,911,944
effective search space: 8801508240
effective search space used: 8801508240
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 66 (30.0 bits)
Medicago: description of AC146862.23


