

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.19 - phase: 0
(1287 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
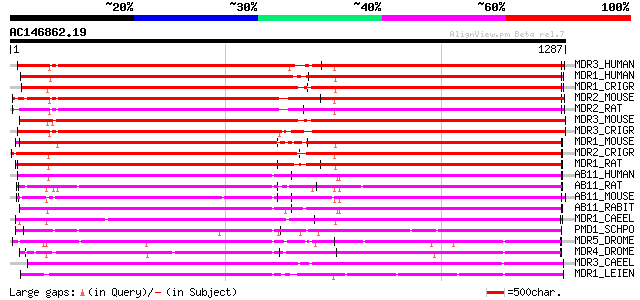
MDR3_HUMAN (P21439) Multidrug resistance protein 3 (P-glycoprote... 963 0.0
MDR1_HUMAN (P08183) Multidrug resistance protein 1 (P-glycoprote... 963 0.0
MDR1_CRIGR (P21448) Multidrug resistance protein 1 (P-glycoprote... 956 0.0
MDR2_MOUSE (P21440) Multidrug resistance protein 2 (P-glycoprote... 956 0.0
MDR2_RAT (Q08201) Multidrug resistance protein 2 (P-glycoprotein... 947 0.0
MDR3_MOUSE (P21447) Multidrug resistance protein 3 (P-glycoprote... 946 0.0
MDR3_CRIGR (P23174) Multidrug resistance protein 3 (P-glycoprote... 941 0.0
MDR1_MOUSE (P06795) Multidrug resistance protein 1 (P-glycoprote... 920 0.0
MDR2_CRIGR (P21449) Multidrug resistance protein 2 (P-glycoprote... 910 0.0
MDR1_RAT (P43245) Multidrug resistance protein 1 (P-glycoprotein 1) 903 0.0
AB11_HUMAN (O95342) Bile salt export pump (ATP-binding cassette,... 868 0.0
AB11_RAT (O70127) Bile salt export pump (ATP-binding cassette, s... 855 0.0
AB11_MOUSE (Q9QY30) Bile salt export pump (ATP-binding cassette,... 847 0.0
AB11_RABIT (Q9N0V3) Bile salt export pump (ATP-binding cassette,... 835 0.0
MDR1_CAEEL (P34712) Multidrug resistance protein 1 (P-glycoprote... 814 0.0
PMD1_SCHPO (P36619) Leptomycin B resistance protein pmd1 795 0.0
MDR5_DROME (Q00748) Multidrug resistance protein homolog 65 (P-g... 760 0.0
MDR4_DROME (Q00449) Multidrug resistance protein homolog 49 (P-g... 754 0.0
MDR3_CAEEL (P34713) Multidrug resistance protein 3 (P-glycoprote... 739 0.0
MDR1_LEIEN (Q06034) Multidrug resistance protein 1 (P-glycoprote... 667 0.0
>MDR3_HUMAN (P21439) Multidrug resistance protein 3 (P-glycoprotein 3)
Length = 1279
Score = 963 bits (2489), Expect = 0.0
Identities = 523/1288 (40%), Positives = 796/1288 (61%), Gaps = 61/1288 (4%)
Query: 19 DHDSNQDSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILIFG 78
D + S++ + K T K + + LF ++D D+L M +GT+ AI +G +PLM+++FG
Sbjct: 20 DFELGISSKQKRKKTKTVKMIGVLTLFRYSDWQDKLFMSLGTIMAIAHGSGLPLMMIVFG 79
Query: 79 TMINAFGDSTNS---------------KVVDEVSETTTYCDQVSLKFVYLAAGTFVASFL 123
M + F D+ + K+++E E T Y S L AG VA+++
Sbjct: 80 EMTDKFVDTAGNFSFPVNFSLSLLNPGKILEE--EMTRYAYYYS----GLGAGVLVAAYI 133
Query: 124 QLTCWMITGERQSARIRGLYLKTILRQDVSFFDKETNTGEVVGRMSGDTVLIKDAMGEKV 183
Q++ W + RQ +IR + ILRQ++ +FD T E+ R++ D I + +G+KV
Sbjct: 134 QVSFWTLAAGRQIRKIRQKFFHAILRQEIGWFDINDTT-ELNTRLTDDISKISEGIGDKV 192
Query: 184 GQFIQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKS 243
G F Q ++TF GF++ F +GW LT+V+++ P+L LS ++ + +++ S AAY+K+
Sbjct: 193 GMFFQAVATFFAGFIVGFIRGWKLTLVIMAISPILGLSAAVWAKILSAFSDKELAAYAKA 252
Query: 244 AGVVEQTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGVGFGTLFFVFICSY 303
V E+ +G+IRTV +F G+ + Y + L + +++A+++ + G F + SY
Sbjct: 253 GAVAEEALGAIRTVIAFGGQNKELERYQKHLENAKEIGIKKAISANISMGIAFLLIYASY 312
Query: 304 GLAVWFGGKMIIEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETIN 363
LA W+G ++I K YT G+ MTV F++LIG+ +GQ +P + AFA + AA+ +F+ I+
Sbjct: 313 ALAFWYGSTLVISKEYTIGNAMTVFFSILIGAFSVGQAAPCIDAFANARGAAYVIFDIID 372
Query: 364 RKPEIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSG 423
P+ID++ G K D I+G++E DV FSYP+R + I G +L + SG T ALVG SG
Sbjct: 373 NNPKIDSFSERGHKPDSIKGNLEFNDVHFSYPSRANVKILKGLNLKVQSGQTVALVGSSG 432
Query: 424 SGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAY 483
GKST V LI+R YDP +G + IDG +++ F + ++R+ IG+VSQEPVLF+ +I ENI Y
Sbjct: 433 CGKSTTVQLIQRLYDPDEGTINIDGQDIRNFNVNYLREIIGVVSQEPVLFSTTIAENICY 492
Query: 484 GKDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPR 543
G+ T +EI+ A + ANA +FI KLPQ DT+VGE G QLSGGQKQR+AIARA++++P+
Sbjct: 493 GRGNVTMDEIKKAVKEANAYEFIMKLPQKFDTLVGERGAQLSGGQKQRIAIARALVRNPK 552
Query: 544 ILLLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERG 603
ILLLDEATSALD ESE VQ AL++ RTTIV+AHRLST+RN D IA G IVE+G
Sbjct: 553 ILLLDEATSALDTESEAEVQAALDKAREGRTTIVIAHRLSTVRNADVIAGFEDGVIVEQG 612
Query: 604 SHAELTNDPNGAYSQLIRLQ---EMKRSEQNDANDKNKPNSIVHSG------RQSSQRSF 654
SH+EL G Y +L+ +Q +SE+ + ND+ + +G R S+Q++
Sbjct: 613 SHSELMK-KEGVYFKLVNMQTSGSQIQSEEFELNDEKAATRMAPNGWKSRLFRHSTQKNL 671
Query: 655 SLRSISQGSAGNSGRHSFSASYVAPTTDGFLETEDGGPQASPSKNSSPPEVPLYRLAYFN 714
+ Q S L+ E G +A + P V ++ N
Sbjct: 672 KNSQMCQKS---------------------LDVETDGLEA------NVPPVSFLKVLKLN 704
Query: 715 KPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYKPADELRHDS-KVWAIVFVAVAVA 773
K E P ++GT+ A+ +G + P ++ S++I+ F D ++ +++++F+ + +
Sbjct: 705 KTEWPYFVVGTVCAIANGGLQPAFSVIFSEIIAIFGPGDDAVKQQKCNIFSLIFLFLGII 764
Query: 774 SLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASV 833
S + + FG AG L +R+R + F+ ++ ++SWFDD ++S+GAL RL+TDAA V
Sbjct: 765 SFFTFFLQGFTFGKAGEILTRRLRSMAFKAMLRQDMSWFDDHKNSTGALSTRLATDAAQV 824
Query: 834 RALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSAD 893
+ G L L+ QNIA + G++I+F WQL ++LA+ P++ ++G V++K+L G +
Sbjct: 825 QGATGTRLALIAQNIANLGTGIIISFIYGWQLTLLLLAVVPIIAVSGIVEMKLLAGNAKR 884
Query: 894 AKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSS 953
KK E A ++A +A+ +IRTV S E K +Y +K GP + V++ I G+ F S
Sbjct: 885 DKKELEAAGKIATEAIENIRTVVSLTQERKFESMYVEKLYGPYRNSVQKAHIYGITFSIS 944
Query: 954 FFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAA 1013
+Y A F GA L+ +G F DV LVF A+ A+ + + + PD AK +A
Sbjct: 945 QAFMYFSYAGCFRFGAYLIVNGHMRFRDVILVFSAIVFGAVALGHASSFAPDYAKAKLSA 1004
Query: 1014 ASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKT 1073
A +F + +++ IDS E G+ ++ +G+I FN V F YPTR +V + L L ++ G+T
Sbjct: 1005 AHLFMLFERQPLIDSYSEEGLKPDKFEGNITFNEVVFNYPTRANVPVLQGLSLEVKKGQT 1064
Query: 1074 VALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFND 1133
+ALVG SG GKSTV+ LL+RFYDP +G + LDG E +++ V+WLR Q+G+VSQEPILF+
Sbjct: 1065 LALVGSSGCGKSTVVQLLERFYDPLAGTVLLDGQEAKKLNVQWLRAQLGIVSQEPILFDC 1124
Query: 1134 TVRANIAYGKGGD-ATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVA 1192
++ NIAYG ++ EIV+AA+ AN H FI +L Y+T VG++G QLSGGQKQR+A
Sbjct: 1125 SIAENIAYGDNSRVVSQDEIVSAAKAANIHPFIETLPHKYETRVGDKGTQLSGGQKQRIA 1184
Query: 1193 IARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAV 1252
IARA+++ P+ILLLDEATSALD ESEKVVQ+ALD+ RT I++AHRLSTI+ ADLI V
Sbjct: 1185 IARALIRQPQILLLDEATSALDTESEKVVQEALDKAREGRTCIVIAHRLSTIQNADLIVV 1244
Query: 1253 VKNGVIAEKGKHEALLHKGGDYASLVAL 1280
+NG + E G H+ LL + G Y S+V++
Sbjct: 1245 FQNGRVKEHGTHQQLLAQKGIYFSMVSV 1272
Score = 440 bits (1132), Expect = e-122
Identities = 235/580 (40%), Positives = 353/580 (60%), Gaps = 20/580 (3%)
Query: 723 MGTITAVLHGAIMPVIGLLVSKMISTFY-----------------KPADELRHDSKVWAI 765
+GTI A+ HG+ +P++ ++ +M F P L + +A
Sbjct: 59 LGTIMAIAHGSGLPLMMIVFGEMTDKFVDTAGNFSFPVNFSLSLLNPGKILEEEMTRYAY 118
Query: 766 VFVAVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGAR 825
+ + L+ + F+ +A G+ I++IR+ F ++ E+ WFD + + L R
Sbjct: 119 YYSGLGAGVLVAAYIQVSFWTLAAGRQIRKIRQKFFHAILRQEIGWFDI--NDTTELNTR 176
Query: 826 LSTDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVK 885
L+ D + + +GD +G+ Q +AT G ++ F W+L +++A++P+LGL+ V K
Sbjct: 177 LTDDISKISEGIGDKVGMFFQAVATFFAGFIVGFIRGWKLTLVIMAISPILGLSAAVWAK 236
Query: 886 VLKGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGII 945
+L FS Y +A VA +A+G+IRTV +F + K +E Y++ E + G+++ I
Sbjct: 237 ILSAFSDKELAAYAKAGAVAEEALGAIRTVIAFGGQNKELERYQKHLENAKEIGIKKAIS 296
Query: 946 SGLGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPD 1005
+ + G +F ++YA A F+ G+ LV + T + VFF++ + A V Q+ +
Sbjct: 297 ANISMGIAFLLIYASYALAFWYGSTLVISKEYTIGNAMTVFFSILIGAFSVGQAAPCIDA 356
Query: 1006 STNAKSAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLC 1065
NA+ AA IF I+D +IDS E G + +KG++EFN V F YP+R +V+I L
Sbjct: 357 FANARGAAYVIFDIIDNNPKIDSFSERGHKPDSIKGNLEFNDVHFSYPSRANVKILKGLN 416
Query: 1066 LNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVS 1125
L ++SG+TVALVG SG GKST + L+QR YDPD G I +DG +I+ V +LR+ +G+VS
Sbjct: 417 LKVQSGQTVALVGSSGCGKSTTVQLIQRLYDPDEGTINIDGQDIRNFNVNYLREIIGVVS 476
Query: 1126 QEPILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSG 1185
QEP+LF+ T+ NI YG+G + T EI A + ANA++FI L + +DT+VGERG QLSG
Sbjct: 477 QEPVLFSTTIAENICYGRG-NVTMDEIKKAVKEANAYEFIMKLPQKFDTLVGERGAQLSG 535
Query: 1186 GQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIK 1245
GQKQR+AIARA+V+NPKILLLDEATSALD ESE VQ ALD+ RTTI++AHRLST++
Sbjct: 536 GQKQRIAIARALVRNPKILLLDEATSALDTESEAEVQAALDKAREGRTTIVIAHRLSTVR 595
Query: 1246 GADLIAVVKNGVIAEKGKHEALLHKGGDYASLVALHTSDS 1285
AD+IA ++GVI E+G H L+ K G Y LV + TS S
Sbjct: 596 NADVIAGFEDGVIVEQGSHSELMKKEGVYFKLVNMQTSGS 635
>MDR1_HUMAN (P08183) Multidrug resistance protein 1 (P-glycoprotein 1)
(CD243 antigen)
Length = 1280
Score = 963 bits (2489), Expect = 0.0
Identities = 521/1273 (40%), Positives = 791/1273 (61%), Gaps = 36/1273 (2%)
Query: 25 DSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILIFGTMINAF 84
+++ KDK TV ++ +F +++ D+L M++GTL AI +G +PLM+L+FG M + F
Sbjct: 20 NNKSEKDKKEKKPTVSVFSMFRYSNWLDKLYMVVGTLAAIIHGAGLPLMMLVFGEMTDIF 79
Query: 85 GDSTNSKVV-------DEVSETTTYC----DQVSLKFVY--LAAGTFVASFLQLTCWMIT 131
++ N + + ++++T + D + Y + AG VA+++Q++ W +
Sbjct: 80 ANAGNLEDLMSNITNRSDINDTGFFMNLEEDMTRYAYYYSGIGAGVLVAAYIQVSFWCLA 139
Query: 132 GERQSARIRGLYLKTILRQDVSFFDKETNTGEVVGRMSGDTVLIKDAMGEKVGQFIQFMS 191
RQ +IR + I+RQ++ +FD + GE+ R++ D I + +G+K+G F Q M+
Sbjct: 140 AGRQIHKIRKQFFHAIMRQEIGWFDVH-DVGELNTRLTDDVSKINEGIGDKIGMFFQSMA 198
Query: 192 TFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKSAGVVEQTI 251
TF GF++ FT+GW LT+V+L+ P+L LS ++ + +++ + AY+K+ V E+ +
Sbjct: 199 TFFTGFIVGFTRGWKLTLVILAISPVLGLSAAVWAKILSSFTDKELLAYAKAGAVAEEVL 258
Query: 252 GSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGVGFGTLFFVFICSYGLAVWFGG 311
+IRTV +F G+K+ YN++L + + +++A+ + + G F + SY LA W+G
Sbjct: 259 AAIRTVIAFGGQKKELERYNKNLEEAKRIGIKKAITANISIGAAFLLIYASYALAFWYGT 318
Query: 312 KMIIEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETINRKPEIDAY 371
+++ Y+ G V+TV F+VLIG+ +GQ SPS+ AFA + AA+++F+ I+ KP ID+Y
Sbjct: 319 TLVLSGEYSIGQVLTVFFSVLIGAFSVGQASPSIEAFANARGAAYEIFKIIDNKPSIDSY 378
Query: 372 DTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVS 431
SG K D+I+G++E R+V FSYP+R + I G +L + SG T ALVG SG GKST V
Sbjct: 379 SKSGHKPDNIKGNLEFRNVHFSYPSRKEVKILKGLNLKVQSGQTVALVGNSGCGKSTTVQ 438
Query: 432 LIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKDCATDE 491
L++R YDPT+G V +DG +++ ++++R+ IG+VSQEPVLF +I ENI YG++ T +
Sbjct: 439 LMQRLYDPTEGMVSVDGQDIRTINVRFLREIIGVVSQEPVLFATTIAENIRYGRENVTMD 498
Query: 492 EIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLDEAT 551
EI A + ANA FI KLP DT+VGE G QLSGGQKQR+AIARA++++P+ILLLDEAT
Sbjct: 499 EIEKAVKEANAYDFIMKLPHKFDTLVGERGAQLSGGQKQRIAIARALVRNPKILLLDEAT 558
Query: 552 SALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAELTND 611
SALD ESE +VQ AL++ RTTIV+AHRLST+RN D IA G IVE+G+H EL +
Sbjct: 559 SALDTESEAVVQVALDKARKGRTTIVIAHRLSTVRNADVIAGFDDGVIVEKGNHDELMKE 618
Query: 612 PNGAYSQLIRLQEM-KRSEQNDANDKNKPNSIVHSGRQSSQRSFSLRSISQGSAGNSGRH 670
G Y +L+ +Q E +A D++K + RS +R S R
Sbjct: 619 -KGIYFKLVTMQTAGNEVELENAADESKSEIDALEMSSNDSRSSLIRK-------RSTRR 670
Query: 671 SFSASYVAPTTDGFLETEDGGPQASPSKNSSPPEVPLYRLAYFNKPEIPVLLMGTITAVL 730
S S + +D + + S P V +R+ N E P ++G A++
Sbjct: 671 SVRGS----------QAQDRKLSTKEALDESIPPVSFWRIMKLNLTEWPYFVVGVFCAII 720
Query: 731 HGAIMPVIGLLVSKMISTFYKPAD--ELRHDSKVWAIVFVAVAVASLLIIPCRFYFFGVA 788
+G + P ++ SK+I F + D R +S +++++F+A+ + S + + + FG A
Sbjct: 721 NGGLQPAFAIIFSKIIGVFTRIDDPETKRQNSNLFSLLFLALGIISFITFFLQGFTFGKA 780
Query: 789 GGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQNI 848
G L +R+R + F ++ +VSWFDD ++++GAL RL+ DAA V+ +G L ++ QNI
Sbjct: 781 GEILTKRLRYMVFRSMLRQDVSWFDDPKNTTGALTTRLANDAAQVKGAIGSRLAVITQNI 840
Query: 849 ATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDA 908
A + G++I+F WQL ++LA+ P++ + G V++K+L G + KK E A ++A +A
Sbjct: 841 ANLGTGIIISFIYGWQLTLLLLAIVPIIAIAGVVEMKMLSGQALKDKKELEGAGKIATEA 900
Query: 909 VGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDACVFYAG 968
+ + RTV S E+K +Y Q + P + +R+ I G+ F + M+Y A F G
Sbjct: 901 IENFRTVVSLTQEQKFEHMYAQSLQVPYRNSLRKAHIFGITFSFTQAMMYFSYAGCFRFG 960
Query: 969 ARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDS 1028
A LV +F DV LVF A+ AM V Q + PD AK +AA I I+++ IDS
Sbjct: 961 AYLVAHKLMSFEDVLLVFSAVVFGAMAVGQVSSFAPDYAKAKISAAHIIMIIEKTPLIDS 1020
Query: 1029 SDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVI 1088
G+ ++G++ F V F YPTR D+ + L L ++ G+T+ALVG SG GKSTV+
Sbjct: 1021 YSTEGLMPNTLEGNVTFGEVVFNYPTRPDIPVLQGLSLEVKKGQTLALVGSSGCGKSTVV 1080
Query: 1089 SLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGD-A 1147
LL+RFYDP +G + LDG EI+R+ V+WLR +G+VSQEPILF+ ++ NIAYG
Sbjct: 1081 QLLERFYDPLAGKVLLDGKEIKRLNVQWLRAHLGIVSQEPILFDCSIAENIAYGDNSRVV 1140
Query: 1148 TEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLD 1207
++ EIV AA+ AN H FI SL Y T VG++G QLSGGQKQR+AIARA+V+ P ILLLD
Sbjct: 1141 SQEEIVRAAKEANIHAFIESLPNKYSTKVGDKGTQLSGGQKQRIAIARALVRQPHILLLD 1200
Query: 1208 EATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEAL 1267
EATSALD ESEKVVQ+ALD+ RT I++AHRLSTI+ ADLI V +NG + E G H+ L
Sbjct: 1201 EATSALDTESEKVVQEALDKAREGRTCIVIAHRLSTIQNADLIVVFQNGRVKEHGTHQQL 1260
Query: 1268 LHKGGDYASLVAL 1280
L + G Y S+V++
Sbjct: 1261 LAQKGIYFSMVSV 1273
Score = 444 bits (1143), Expect = e-124
Identities = 239/613 (38%), Positives = 365/613 (58%), Gaps = 25/613 (4%)
Query: 693 QASPSKNSSPPEVPLYRL-AYFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYK 751
++ K P V ++ + Y N + +++GT+ A++HGA +P++ L+ +M F
Sbjct: 22 KSEKDKKEKKPTVSVFSMFRYSNWLDKLYMVVGTLAAIIHGAGLPLMMLVFGEMTDIFAN 81
Query: 752 PAD---------------------ELRHDSKVWAIVFVAVAVASLLIIPCRFYFFGVAGG 790
+ L D +A + + L+ + F+ +A G
Sbjct: 82 AGNLEDLMSNITNRSDINDTGFFMNLEEDMTRYAYYYSGIGAGVLVAAYIQVSFWCLAAG 141
Query: 791 KLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQNIAT 850
+ I +IRK F ++ E+ WFD H G L RL+ D + + +GD +G+ Q++AT
Sbjct: 142 RQIHKIRKQFFHAIMRQEIGWFD--VHDVGELNTRLTDDVSKINEGIGDKIGMFFQSMAT 199
Query: 851 IIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDAVG 910
G ++ F W+L ++LA++P+LGL+ V K+L F+ Y +A VA + +
Sbjct: 200 FFTGFIVGFTRGWKLTLVILAISPVLGLSAAVWAKILSSFTDKELLAYAKAGAVAEEVLA 259
Query: 911 SIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDACVFYAGAR 970
+IRTV +F ++K +E Y + E + G+++ I + + G++F ++YA A F+ G
Sbjct: 260 AIRTVIAFGGQKKELERYNKNLEEAKRIGIKKAITANISIGAAFLLIYASYALAFWYGTT 319
Query: 971 LVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDSSD 1030
LV G+ + V VFF++ + A V Q+ + NA+ AA IF I+D K IDS
Sbjct: 320 LVLSGEYSIGQVLTVFFSVLIGAFSVGQASPSIEAFANARGAAYEIFKIIDNKPSIDSYS 379
Query: 1031 ESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISL 1090
+SG + +KG++EF +V F YP+R +V+I L L ++SG+TVALVG SG GKST + L
Sbjct: 380 KSGHKPDNIKGNLEFRNVHFSYPSRKEVKILKGLNLKVQSGQTVALVGNSGCGKSTTVQL 439
Query: 1091 LQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEA 1150
+QR YDP G +++DG +I+ + V++LR+ +G+VSQEP+LF T+ NI YG+ + T
Sbjct: 440 MQRLYDPTEGMVSVDGQDIRTINVRFLREIIGVVSQEPVLFATTIAENIRYGRE-NVTMD 498
Query: 1151 EIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEAT 1210
EI A + ANA+ FI L +DT+VGERG QLSGGQKQR+AIARA+V+NPKILLLDEAT
Sbjct: 499 EIEKAVKEANAYDFIMKLPHKFDTLVGERGAQLSGGQKQRIAIARALVRNPKILLLDEAT 558
Query: 1211 SALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLHK 1270
SALD ESE VVQ ALD+ RTTI++AHRLST++ AD+IA +GVI EKG H+ L+ +
Sbjct: 559 SALDTESEAVVQVALDKARKGRTTIVIAHRLSTVRNADVIAGFDDGVIVEKGNHDELMKE 618
Query: 1271 GGDYASLVALHTS 1283
G Y LV + T+
Sbjct: 619 KGIYFKLVTMQTA 631
>MDR1_CRIGR (P21448) Multidrug resistance protein 1 (P-glycoprotein 1)
Length = 1276
Score = 956 bits (2471), Expect = 0.0
Identities = 512/1271 (40%), Positives = 790/1271 (61%), Gaps = 38/1271 (2%)
Query: 27 EKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILIFGTMINAFG- 85
+ K+K V ++ +F +A DRL ML+GTL AI +G+++PLM+L+FG M ++F
Sbjct: 21 KSKKEKKEKKPVVSVFTMFRYAGWLDRLYMLVGTLAAIIHGVALPLMMLVFGDMTDSFAS 80
Query: 86 ------DSTNSKVVDEVSETTTYCDQVSLKFVY----LAAGTFVASFLQLTCWMITGERQ 135
++TN+ S+ ++ + Y + AG + +++Q++ W + RQ
Sbjct: 81 VGNIPTNATNNATQVNASDIFGKLEEEMTTYAYYYTGIGAGVLIVAYIQVSFWCLAAGRQ 140
Query: 136 SARIRGLYLKTILRQDVSFFDKETNTGEVVGRMSGDTVLIKDAMGEKVGQFIQFMSTFIG 195
+IR + I+ Q++ +FD + GE+ R++ D I + +G+K+G F Q M+TF G
Sbjct: 141 IHKIRQKFFHAIMNQEIGWFDVH-DVGELNTRLTDDVSKINEGIGDKIGMFFQAMATFFG 199
Query: 196 GFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKSAGVVEQTIGSIR 255
GF+I FT+GW LT+V+L+ P+L LS + + +++ + AY+K+ V E+ + +IR
Sbjct: 200 GFIIGFTRGWKLTLVILAISPVLGLSAGIWAKILSSFTDKELQAYAKAGAVAEEVLAAIR 259
Query: 256 TVASFTGEKQATANYNRSLIKVYKTAVQEALASGVGFGTLFFVFICSYGLAVWFGGKMII 315
TV +F G+K+ YN +L + + +++A+ + + G F + SY LA W+G ++I
Sbjct: 260 TVIAFGGQKKELERYNNNLEEAKRLGIKKAITANISMGAAFLLIYASYALAFWYGTSLVI 319
Query: 316 EKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETINRKPEIDAYDTSG 375
K Y+ G V+TV FAVLIG+ +GQ SP++ AFA + AA+++F I+ KP ID++ +G
Sbjct: 320 SKEYSIGQVLTVFFAVLIGAFSIGQASPNIEAFANARGAAYEIFNIIDNKPSIDSFSKNG 379
Query: 376 KKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIER 435
K D+I+G++E +++ FSYP+R D I G +L + SG T ALVG SG GKST V L++R
Sbjct: 380 YKPDNIKGNLEFKNIHFSYPSRKDVQILKGLNLKVQSGQTVALVGNSGCGKSTTVQLLQR 439
Query: 436 FYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKDCATDEEIRV 495
YDPT+G V IDG +++ ++++R+ IG+VSQEPVLF +I ENI YG++ T +EI
Sbjct: 440 LYDPTEGVVSIDGQDIRTINVRYLREIIGVVSQEPVLFATTIAENIRYGRENVTMDEIEK 499
Query: 496 AAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLDEATSALD 555
A + ANA FI KLP DT+VGE G QLSGGQKQR+AIARA++++P+ILLLDEATSALD
Sbjct: 500 AVKEANAYDFIMKLPHKFDTLVGERGAQLSGGQKQRIAIARALVRNPKILLLDEATSALD 559
Query: 556 AESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAELTNDPNGA 615
ESE +VQ AL++ RTTIV+AHRLST+RN D IA G IVE+G+H EL + G
Sbjct: 560 TESEAVVQAALDKAREGRTTIVIAHRLSTVRNADIIAGFDGGVIVEQGNHEELMRE-KGI 618
Query: 616 YSQLIRLQEMKRSEQ--NDAND-KNKPNSIVHSGRQSSQRSFSLRSISQGSAGNSGRHSF 672
Y +L+ Q + N+ + KN+ +++ S + S+ RS + G +
Sbjct: 619 YFKLVMTQTAGNEIELGNEVGESKNEIDNLDMSSKDSASSLIRRRSTRRSIRGPHDQ--- 675
Query: 673 SASYVAPTTDGFLETEDGGPQASPSKNSSPPEVPLYRLAYFNKPEIPVLLMGTITAVLHG 732
D L T++ + + P + +R+ N E P ++G A+++G
Sbjct: 676 ---------DRKLSTKE-------ALDEDVPPISFWRILKLNSSEWPYFVVGIFCAIVNG 719
Query: 733 AIMPVIGLLVSKMISTFYKPADE--LRHDSKVWAIVFVAVAVASLLIIPCRFYFFGVAGG 790
A+ P ++ SK++ F + D+ RHDS +++++F+ + V S + + + FG AG
Sbjct: 720 ALQPAFSIIFSKVVGVFTRNTDDETKRHDSNLFSLLFLILGVISFITFFLQGFTFGKAGE 779
Query: 791 KLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQNIAT 850
L +R+R + F+ ++ +VSWFD+ ++++GAL RL+ DA V+ G L ++ QNIA
Sbjct: 780 ILTKRLRYMVFKSMLRQDVSWFDNPKNTTGALTTRLANDAGQVKGATGARLAVITQNIAN 839
Query: 851 IIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDAVG 910
+ G++I+ WQL ++LA+ P++ + G V++K+L G + KK E + ++A +A+
Sbjct: 840 LGTGIIISLIYGWQLTLLLLAIVPIIAIAGVVEMKMLSGQALKDKKELEGSGKIATEAIE 899
Query: 911 SIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDACVFYAGAR 970
+ RTV S E+K +Y Q + P + +++ + G+ F + M+Y A F GA
Sbjct: 900 NFRTVVSLTREQKFENMYAQSLQIPYRNALKKAHVFGITFSFTQAMMYFSYAACFRFGAY 959
Query: 971 LVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDSSD 1030
LV TF +V LVF A+ AM V Q + PD AK +A+ I I+++ IDS
Sbjct: 960 LVARELMTFENVLLVFSAIVFGAMAVGQVSSFAPDYAKAKVSASHIIMIIEKVPSIDSYS 1019
Query: 1031 ESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISL 1090
G+ ++G+++FN V F YPTR D+ + L L ++ G+T+ALVG SG GKSTV+ L
Sbjct: 1020 TGGLKPNTLEGNVKFNEVVFNYPTRPDIPVLQGLNLEVKKGQTLALVGSSGCGKSTVVQL 1079
Query: 1091 LQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGD-ATE 1149
L+RFYDP +G + LDG E+ ++ V+WLR +G+VSQEPILF+ ++ NIAYG ++
Sbjct: 1080 LERFYDPMAGTVFLDGKEVNQLNVQWLRAHLGIVSQEPILFDCSIAENIAYGDNSRVVSQ 1139
Query: 1150 AEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEA 1209
EI AA+ AN HQFI SL Y+T VG++G QLSGGQKQR+AIARA+V+ P ILLLDEA
Sbjct: 1140 DEIERAAKEANIHQFIESLPDKYNTRVGDKGTQLSGGQKQRIAIARALVRQPHILLLDEA 1199
Query: 1210 TSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLH 1269
TSALD ESEKVVQ+ALD+ RT I++AHRLSTI+ ADLI V++NG + E G H+ LL
Sbjct: 1200 TSALDTESEKVVQEALDKAREGRTCIVIAHRLSTIQNADLIVVIQNGKVKEHGTHQQLLA 1259
Query: 1270 KGGDYASLVAL 1280
+ G Y S+V++
Sbjct: 1260 QKGIYFSMVSV 1270
Score = 443 bits (1140), Expect = e-123
Identities = 237/613 (38%), Positives = 363/613 (58%), Gaps = 23/613 (3%)
Query: 691 GPQASPSKNSSPPEVPLYRL-AYFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTF 749
G ++ K P V ++ + Y + +L+GT+ A++HG +P++ L+ M +F
Sbjct: 19 GRKSKKEKKEKKPVVSVFTMFRYAGWLDRLYMLVGTLAAIIHGVALPLMMLVFGDMTDSF 78
Query: 750 YKPAD-------------------ELRHDSKVWAIVFVAVAVASLLIIPCRFYFFGVAGG 790
+ +L + +A + + L++ + F+ +A G
Sbjct: 79 ASVGNIPTNATNNATQVNASDIFGKLEEEMTTYAYYYTGIGAGVLIVAYIQVSFWCLAAG 138
Query: 791 KLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQNIAT 850
+ I +IR+ F +++ E+ WFD H G L RL+ D + + +GD +G+ Q +AT
Sbjct: 139 RQIHKIRQKFFHAIMNQEIGWFD--VHDVGELNTRLTDDVSKINEGIGDKIGMFFQAMAT 196
Query: 851 IIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDAVG 910
G +I F W+L ++LA++P+LGL+ + K+L F+ + Y +A VA + +
Sbjct: 197 FFGGFIIGFTRGWKLTLVILAISPVLGLSAGIWAKILSSFTDKELQAYAKAGAVAEEVLA 256
Query: 911 SIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDACVFYAGAR 970
+IRTV +F ++K +E Y E + G+++ I + + G++F ++YA A F+ G
Sbjct: 257 AIRTVIAFGGQKKELERYNNNLEEAKRLGIKKAITANISMGAAFLLIYASYALAFWYGTS 316
Query: 971 LVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDSSD 1030
LV + + V VFFA+ + A + Q+ + NA+ AA IF I+D K IDS
Sbjct: 317 LVISKEYSIGQVLTVFFAVLIGAFSIGQASPNIEAFANARGAAYEIFNIIDNKPSIDSFS 376
Query: 1031 ESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISL 1090
++G + +KG++EF ++ F YP+R DVQI L L ++SG+TVALVG SG GKST + L
Sbjct: 377 KNGYKPDNIKGNLEFKNIHFSYPSRKDVQILKGLNLKVQSGQTVALVGNSGCGKSTTVQL 436
Query: 1091 LQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEA 1150
LQR YDP G +++DG +I+ + V++LR+ +G+VSQEP+LF T+ NI YG+ + T
Sbjct: 437 LQRLYDPTEGVVSIDGQDIRTINVRYLREIIGVVSQEPVLFATTIAENIRYGRE-NVTMD 495
Query: 1151 EIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEAT 1210
EI A + ANA+ FI L +DT+VGERG QLSGGQKQR+AIARA+V+NPKILLLDEAT
Sbjct: 496 EIEKAVKEANAYDFIMKLPHKFDTLVGERGAQLSGGQKQRIAIARALVRNPKILLLDEAT 555
Query: 1211 SALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLHK 1270
SALD ESE VVQ ALD+ RTTI++AHRLST++ AD+IA GVI E+G HE L+ +
Sbjct: 556 SALDTESEAVVQAALDKAREGRTTIVIAHRLSTVRNADIIAGFDGGVIVEQGNHEELMRE 615
Query: 1271 GGDYASLVALHTS 1283
G Y LV T+
Sbjct: 616 KGIYFKLVMTQTA 628
>MDR2_MOUSE (P21440) Multidrug resistance protein 2 (P-glycoprotein 2)
Length = 1276
Score = 956 bits (2470), Expect = 0.0
Identities = 524/1292 (40%), Positives = 795/1292 (60%), Gaps = 56/1292 (4%)
Query: 6 RLEGDFVSVQPVEDHDSNQDSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIG 65
RL+GDF SNQ EK K ++ + L LF ++D D+L M +GTL AI
Sbjct: 13 RLDGDFEL-----GSISNQGREKKKKVNL----IGLLTLFRYSDWQDKLFMFLGTLMAIA 63
Query: 66 NGLSIPLMILIFGTMINAFGDSTNS---------------KVVDEVSETTTYCDQVSLKF 110
+G +PLM+++FG M + F D+T + ++++E E T Y S
Sbjct: 64 HGSGLPLMMIVFGEMTDKFVDNTGNFSLPVNFSLSMLNPGRILEE--EMTRYAYYYS--- 118
Query: 111 VYLAAGTFVASFLQLTCWMITGERQSARIRGLYLKTILRQDVSFFDKETNTGEVVGRMSG 170
L G VA+++Q++ W + RQ +IR + ILRQ++ +FD + T E+ R++
Sbjct: 119 -GLGGGVLVAAYIQVSFWTLAAGRQIKKIRQKFFHAILRQEMGWFDIKGTT-ELNTRLTD 176
Query: 171 DTVLIKDAMGEKVGQFIQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIA 230
D I + +G+KVG F Q ++TF GF++ F +GW LT+V+++ P+L LS ++ + +++
Sbjct: 177 DVSKISEGIGDKVGMFFQAIATFFAGFIVGFIRGWKLTLVIMAISPILGLSTAVWAKILS 236
Query: 231 KASSTGQAAYSKSAGVVEQTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGV 290
S AAY+K+ V E+ G+IRTV +F G+ + Y + L K +++A+++ +
Sbjct: 237 TFSDKELAAYAKAGAVAEEAPGAIRTVIAFGGQNKELERYQKHLENAKKIGIKKAISANI 296
Query: 291 GFGTLFFVFICSYGLAVWFGGKMIIEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAA 350
G F + SY LA W+G ++I K YT G+ MTV F++LIG+ +GQ +P + AFA
Sbjct: 297 SMGIAFLLIYASYALAFWYGSTLVISKEYTIGNAMTVFFSILIGAFSVGQAAPCIDAFAN 356
Query: 351 GQAAAFKMFETINRKPEIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSL 410
+ AA+ +F+ I+ P+ID++ G K D+I+G++E DV FSYP+R + I G +L +
Sbjct: 357 ARGAAYVIFDIIDNNPKIDSFSERGHKPDNIKGNLEFSDVHFSYPSRANIKILKGLNLKV 416
Query: 411 PSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEP 470
SG T ALVG SG GKST V L++R YDPT+G++ IDG +++ F ++ +R+ IG+VSQEP
Sbjct: 417 KSGQTVALVGNSGCGKSTTVQLLQRLYDPTEGKISIDGQDIRNFNVRCLREIIGVVSQEP 476
Query: 471 VLFTCSIKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQ 530
VLF+ +I ENI YG+ T +EI A + ANA FI KLPQ DT+VG+ G QLSGGQKQ
Sbjct: 477 VLFSTTIAENIRYGRGNVTMDEIEKAVKEANAYDFIMKLPQKFDTLVGDRGAQLSGGQKQ 536
Query: 531 RVAIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDT 590
R+AIARA++++P+ILLLDEATSALD ESE VQ AL++ RTTIV+AHRLSTIRN D
Sbjct: 537 RIAIARALVRNPKILLLDEATSALDTESEAEVQAALDKAREGRTTIVIAHRLSTIRNADV 596
Query: 591 IAVIHQGKIVERGSHAELTNDPNGAYSQLIRLQEMKRSEQNDANDKNKPNSIVHSGRQSS 650
IA G IVE+GSH+EL G Y +L+ +Q +G Q
Sbjct: 597 IAGFEDGVIVEQGSHSELMK-KEGIYFRLVNMQT--------------------AGSQIL 635
Query: 651 QRSFSLRSISQGSAGNSGRHSFSASYVAPTTDGFLET--EDGGPQASPSKNSSPPEVPLY 708
F + + +AG+ + + A +T L++ ++ + + +++ P V
Sbjct: 636 SEEFEVELSDEKAAGDVAPNGWKARIFRNSTKKSLKSPHQNRLDEETNELDANVPPVSFL 695
Query: 709 RLAYFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYKPADELRHDS-KVWAIVF 767
++ NK E P ++GT+ A+ +GA+ P +++S+MI+ F D ++ ++++VF
Sbjct: 696 KVLKLNKTEWPYFVVGTVCAIANGALQPAFSIILSEMIAIFGPGDDAVKQQKCNMFSLVF 755
Query: 768 VAVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLS 827
+ + V S + + FG AG L R+R + F+ ++ ++SWFDD ++S+GAL RL+
Sbjct: 756 LGLGVLSFFTFFLQGFTFGKAGEILTTRLRSMAFKAMLRQDMSWFDDHKNSTGALSTRLA 815
Query: 828 TDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVL 887
TDAA V+ G L L+ QN A + G++I+F WQL ++L++ P + + G V++K+L
Sbjct: 816 TDAAQVQGATGTKLALIAQNTANLGTGIIISFIYGWQLTLLLLSVVPFIAVAGIVEMKML 875
Query: 888 KGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISG 947
G + KK E A ++A +A+ +IRTV S E K +Y +K GP + VR+ I G
Sbjct: 876 AGNAKRDKKEMEAAGKIATEAIENIRTVVSLTQERKFESMYVEKLHGPYRNSVRKAHIYG 935
Query: 948 LGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDST 1007
+ F S +Y A F G+ L+ +G F DV LVF A+ + A+ + + + PD
Sbjct: 936 ITFSISQAFMYFSYAGCFRFGSYLIVNGHMRFKDVILVFSAIVLGAVALGHASSFAPDYA 995
Query: 1008 NAKSAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLN 1067
AK +AA +F++ +++ IDS G+ ++ +G + FN V F YPTR +V + L L
Sbjct: 996 KAKLSAAYLFSLFERQPLIDSYSGEGLWPDKFEGSVTFNEVVFNYPTRANVPVLQGLSLE 1055
Query: 1068 IRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQE 1127
++ G+T+ALVG SG GKSTV+ LL+RFYDP +G + LDG E +++ V+WLR Q+G+VSQE
Sbjct: 1056 VKKGQTLALVGSSGCGKSTVVQLLERFYDPMAGSVLLDGQEAKKLNVQWLRAQLGIVSQE 1115
Query: 1128 PILFNDTVRANIAYGKGGDAT-EAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGG 1186
PILF+ ++ NIAYG EIV AA+ AN H FI +L + Y+T VG++G QLSGG
Sbjct: 1116 PILFDCSIAENIAYGDNSRVVPHDEIVRAAKEANIHPFIETLPQKYNTRVGDKGTQLSGG 1175
Query: 1187 QKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKG 1246
QKQR+AIARA+++ P++LLLDEATSALD ESEKVVQ+ALD+ RT I++AHRLSTI+
Sbjct: 1176 QKQRIAIARALIRQPRVLLLDEATSALDTESEKVVQEALDKAREGRTCIVIAHRLSTIQN 1235
Query: 1247 ADLIAVVKNGVIAEKGKHEALLHKGGDYASLV 1278
ADLI V++NG + E G H+ LL + G Y S+V
Sbjct: 1236 ADLIVVIENGKVKEHGTHQQLLAQKGIYFSMV 1267
Score = 429 bits (1104), Expect = e-119
Identities = 233/582 (40%), Positives = 351/582 (60%), Gaps = 20/582 (3%)
Query: 721 LLMGTITAVLHGAIMPVIGLLVSKMISTFY-----------------KPADELRHDSKVW 763
+ +GT+ A+ HG+ +P++ ++ +M F P L + +
Sbjct: 54 MFLGTLMAIAHGSGLPLMMIVFGEMTDKFVDNTGNFSLPVNFSLSMLNPGRILEEEMTRY 113
Query: 764 AIVFVAVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALG 823
A + + L+ + F+ +A G+ I++IR+ F ++ E+ WFD + L
Sbjct: 114 AYYYSGLGGGVLVAAYIQVSFWTLAAGRQIKKIRQKFFHAILRQEMGWFDI--KGTTELN 171
Query: 824 ARLSTDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQ 883
RL+ D + + +GD +G+ Q IAT G ++ F W+L +++A++P+LGL+ V
Sbjct: 172 TRLTDDVSKISEGIGDKVGMFFQAIATFFAGFIVGFIRGWKLTLVIMAISPILGLSTAVW 231
Query: 884 VKVLKGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRG 943
K+L FS Y +A VA +A G+IRTV +F + K +E Y++ E K G+++
Sbjct: 232 AKILSTFSDKELAAYAKAGAVAEEAPGAIRTVIAFGGQNKELERYQKHLENAKKIGIKKA 291
Query: 944 IISGLGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLV 1003
I + + G +F ++YA A F+ G+ LV + T + VFF++ + A V Q+ +
Sbjct: 292 ISANISMGIAFLLIYASYALAFWYGSTLVISKEYTIGNAMTVFFSILIGAFSVGQAAPCI 351
Query: 1004 PDSTNAKSAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFND 1063
NA+ AA IF I+D +IDS E G + +KG++EF+ V F YP+R +++I
Sbjct: 352 DAFANARGAAYVIFDIIDNNPKIDSFSERGHKPDNIKGNLEFSDVHFSYPSRANIKILKG 411
Query: 1064 LCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGL 1123
L L ++SG+TVALVG SG GKST + LLQR YDP G I++DG +I+ V+ LR+ +G+
Sbjct: 412 LNLKVKSGQTVALVGNSGCGKSTTVQLLQRLYDPTEGKISIDGQDIRNFNVRCLREIIGV 471
Query: 1124 VSQEPILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQL 1183
VSQEP+LF+ T+ NI YG+G + T EI A + ANA+ FI L + +DT+VG+RG QL
Sbjct: 472 VSQEPVLFSTTIAENIRYGRG-NVTMDEIEKAVKEANAYDFIMKLPQKFDTLVGDRGAQL 530
Query: 1184 SGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLST 1243
SGGQKQR+AIARA+V+NPKILLLDEATSALD ESE VQ ALD+ RTTI++AHRLST
Sbjct: 531 SGGQKQRIAIARALVRNPKILLLDEATSALDTESEAEVQAALDKAREGRTTIVIAHRLST 590
Query: 1244 IKGADLIAVVKNGVIAEKGKHEALLHKGGDYASLVALHTSDS 1285
I+ AD+IA ++GVI E+G H L+ K G Y LV + T+ S
Sbjct: 591 IRNADVIAGFEDGVIVEQGSHSELMKKEGIYFRLVNMQTAGS 632
>MDR2_RAT (Q08201) Multidrug resistance protein 2 (P-glycoprotein 2)
(P-glycoprotein 3)
Length = 1278
Score = 947 bits (2448), Expect = 0.0
Identities = 517/1294 (39%), Positives = 787/1294 (59%), Gaps = 58/1294 (4%)
Query: 6 RLEGDFVSVQPVEDHDSNQDSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIG 65
RL+GDF + S +S++K + LF ++D D+L ML+GT AI
Sbjct: 13 RLDGDF---------ELGSISNQSREKKKKVNLIGPLTLFRYSDWQDKLFMLLGTAMAIA 63
Query: 66 NGLSIPLMILIFGTMINAFGDSTNS---------------KVVDEVSETTTYCDQVSLKF 110
+G +PLM+++FG M + F D+ + ++++E E T Y S
Sbjct: 64 HGSGLPLMMIVFGEMTDKFVDNAGNFSLPVNFSLSMLNPGRILEE--EMTRYAYYYS--- 118
Query: 111 VYLAAGTFVASFLQLTCWMITGERQSARIRGLYLKTILRQDVSFFDKETNTGEVVGRMSG 170
L G +A+++Q++ W + RQ +IR + ILRQ++ +FD + T E+ R++
Sbjct: 119 -GLGGGVLLAAYIQVSFWTLAAGRQIRKIRQKFFHAILRQEMGWFDIKGTT-ELNTRLTD 176
Query: 171 DTVLIKDAMGEKVGQFIQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIA 230
D I + +G+KVG F Q ++TF GF++ F +GW LT+V+++ +L LS ++ + +++
Sbjct: 177 DISKISEGIGDKVGMFFQAIATFFAGFIVGFIRGWKLTLVIMAITAILGLSTAVWAKILS 236
Query: 231 KASSTGQAAYSKSAGVVEQTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGV 290
S AAY+K+ V E+ +G+IRTV +F G+ + Y + L K +++A+++ +
Sbjct: 237 TFSDKELAAYAKAGAVAEEALGAIRTVIAFGGQNKELERYQKHLENAKKIGIKKAISANI 296
Query: 291 GFGTLFFVFICSYGLAVWFGGKMIIEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAA 350
G F + SY LA W+G ++I K YT G+ MTV F++LIG+ +GQ +P + AF
Sbjct: 297 SMGIAFLLIYASYALAFWYGSTLVISKEYTIGNAMTVFFSILIGAFSVGQAAPCIDAFPN 356
Query: 351 GQAAAFKMFETINRKPEIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSL 410
+ AA+ +F+ I+ P+ID++ G K D I+G++E DV FSYP+R + I G +L +
Sbjct: 357 ARGAAYVIFDIIDNNPKIDSFSERGHKPDSIKGNLEFSDVHFSYPSRANIKILKGLNLKV 416
Query: 411 PSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEP 470
SG T ALVG SG GKST V L++R YDPT+G + IDG +++ F ++ +R+ IG+VSQEP
Sbjct: 417 KSGQTVALVGNSGCGKSTTVQLLQRLYDPTEGTISIDGQDIRNFNVRCLREFIGVVSQEP 476
Query: 471 VLFTCSIKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQ 530
VLF+ +I ENI YG+ T +EI+ A + ANA FI KLPQ DT+VG+ G QLSGGQKQ
Sbjct: 477 VLFSTTIAENIRYGRGNVTMDEIKKAVKEANAYDFIMKLPQKFDTLVGDRGAQLSGGQKQ 536
Query: 531 RVAIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDT 590
R+AIARA++++P+ILLLDEATSALD ESE VQ AL++ RTTIV+AHRLST+RN D
Sbjct: 537 RIAIARALVRNPKILLLDEATSALDTESEAEVQAALDKAREGRTTIVIAHRLSTVRNADV 596
Query: 591 IAVIHQGKIVERGSHAELTNDPNGAYSQLIRLQEMKRSEQNDANDKNKPNSIVHSGRQSS 650
IA G IVE+GSH+EL G Y +L+ +Q SG Q
Sbjct: 597 IAGFEDGVIVEQGSHSELIK-KEGIYFRLVNMQT--------------------SGSQIL 635
Query: 651 QRSFSLRSISQGSAGNSGRHSFSASYVAPTTDGFLETEDGGPQ----ASPSKNSSPPEVP 706
F + + +AG + + A +T L++ + +++ P V
Sbjct: 636 SEEFEVELSDEKAAGGVAPNGWKARIFRNSTKKSLKSSRAHQNRLDVETNELDANVPPVS 695
Query: 707 LYRLAYFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYKPADELRHDS-KVWAI 765
++ NK E P ++GT+ A+ +GA+ P +++S+MI+ F D ++ ++++
Sbjct: 696 FLKVLRLNKTEWPYFVVGTLCAIANGALQPAFSIILSEMIAIFGPGDDTVKQQKCNMFSL 755
Query: 766 VFVAVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGAR 825
VF+ + V S + + FG AG L R+R + F+ ++ ++SWFDD ++S+GAL R
Sbjct: 756 VFLGLGVHSFFTFFLQGFTFGKAGEILTTRLRSMAFKAMLRQDMSWFDDHKNSTGALSTR 815
Query: 826 LSTDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVK 885
L+TDAA V+ G L L+ QN A + G++I+F WQL ++L++ P + + G V++K
Sbjct: 816 LATDAAQVQGATGTRLALIAQNTANLGTGIIISFIYGWQLTLLLLSVVPFIAVAGIVEMK 875
Query: 886 VLKGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGII 945
+L G + KK E A ++A +A+ +IRTV S E K +Y +K GP + VR+ I
Sbjct: 876 MLAGNAKRDKKEMEAAGKIATEAIENIRTVVSLTQERKFESMYVEKLHGPYRNSVRKAHI 935
Query: 946 SGLGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPD 1005
G+ F S +Y A F G+ L+ +G F DV LVF A+ + A+ + + + PD
Sbjct: 936 YGITFSISQAFMYFSYAGCFRFGSYLIVNGHMRFKDVILVFSAIVLGAVALGHASSFAPD 995
Query: 1006 STNAKSAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLC 1065
AK +AA +F++ +++ IDS GM ++ +G + FN V F YPTR +V + L
Sbjct: 996 YAKAKLSAAYLFSLFERQPLIDSYSREGMWPDKFEGSVTFNEVVFNYPTRANVPVLQGLS 1055
Query: 1066 LNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVS 1125
L ++ G+T+ALVG SG GKSTV+ LL+RFYDP +G + LDG E +++ V+WLR Q+G+VS
Sbjct: 1056 LEVKKGQTLALVGSSGCGKSTVVQLLERFYDPMAGTVLLDGQEAKKLNVQWLRAQLGIVS 1115
Query: 1126 QEPILFNDTVRANIAYGKGGD-ATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLS 1184
QEPILF+ ++ NIAYG ++ EIV AA+ AN H FI +L + Y+T VG++G QLS
Sbjct: 1116 QEPILFDCSIAKNIAYGDNSRVVSQDEIVRAAKEANIHPFIETLPQKYETRVGDKGTQLS 1175
Query: 1185 GGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTI 1244
GGQKQR+AIARA+++ P++LLLDEATSALD ESEKVVQ+ALD+ RT I++AHRLSTI
Sbjct: 1176 GGQKQRIAIARALIRQPRVLLLDEATSALDTESEKVVQEALDKAREGRTCIVIAHRLSTI 1235
Query: 1245 KGADLIAVVKNGVIAEKGKHEALLHKGGDYASLV 1278
+ ADLI V+ NG + E G H+ LL + G Y S+V
Sbjct: 1236 QNADLIVVIDNGKVKEHGTHQQLLAQKGIYFSMV 1269
Score = 430 bits (1105), Expect = e-119
Identities = 242/621 (38%), Positives = 361/621 (57%), Gaps = 20/621 (3%)
Query: 682 DGFLETEDGGPQASPSKNSSPPEVPLYRLAYFNKPEIPVLLMGTITAVLHGAIMPVIGLL 741
DG E Q+ K PL Y + + +L+GT A+ HG+ +P++ ++
Sbjct: 15 DGDFELGSISNQSREKKKKVNLIGPLTLFRYSDWQDKLFMLLGTAMAIAHGSGLPLMMIV 74
Query: 742 VSKMISTFY-----------------KPADELRHDSKVWAIVFVAVAVASLLIIPCRFYF 784
+M F P L + +A + + LL + F
Sbjct: 75 FGEMTDKFVDNAGNFSLPVNFSLSMLNPGRILEEEMTRYAYYYSGLGGGVLLAAYIQVSF 134
Query: 785 FGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALGLL 844
+ +A G+ I++IR+ F ++ E+ WFD + L RL+ D + + +GD +G+
Sbjct: 135 WTLAAGRQIRKIRQKFFHAILRQEMGWFDI--KGTTELNTRLTDDISKISEGIGDKVGMF 192
Query: 845 VQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQV 904
Q IAT G ++ F W+L +++A+ +LGL+ V K+L FS Y +A V
Sbjct: 193 FQAIATFFAGFIVGFIRGWKLTLVIMAITAILGLSTAVWAKILSTFSDKELAAYAKAGAV 252
Query: 905 ANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDACV 964
A +A+G+IRTV +F + K +E Y++ E K G+++ I + + G +F ++YA A
Sbjct: 253 AEEALGAIRTVIAFGGQNKELERYQKHLENAKKIGIKKAISANISMGIAFLLIYASYALA 312
Query: 965 FYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKS 1024
F+ G+ LV + T + VFF++ + A V Q+ + NA+ AA IF I+D
Sbjct: 313 FWYGSTLVISKEYTIGNAMTVFFSILIGAFSVGQAAPCIDAFPNARGAAYVIFDIIDNNP 372
Query: 1025 QIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGK 1084
+IDS E G + +KG++EF+ V F YP+R +++I L L ++SG+TVALVG SG GK
Sbjct: 373 KIDSFSERGHKPDSIKGNLEFSDVHFSYPSRANIKILKGLNLKVKSGQTVALVGNSGCGK 432
Query: 1085 STVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKG 1144
ST + LLQR YDP G I++DG +I+ V+ LR+ +G+VSQEP+LF+ T+ NI YG+G
Sbjct: 433 STTVQLLQRLYDPTEGTISIDGQDIRNFNVRCLREFIGVVSQEPVLFSTTIAENIRYGRG 492
Query: 1145 GDATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKIL 1204
+ T EI A + ANA+ FI L + +DT+VG+RG QLSGGQKQR+AIARA+V+NPKIL
Sbjct: 493 -NVTMDEIKKAVKEANAYDFIMKLPQKFDTLVGDRGAQLSGGQKQRIAIARALVRNPKIL 551
Query: 1205 LLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKH 1264
LLDEATSALD ESE VQ ALD+ RTTI++AHRLST++ AD+IA ++GVI E+G H
Sbjct: 552 LLDEATSALDTESEAEVQAALDKAREGRTTIVIAHRLSTVRNADVIAGFEDGVIVEQGSH 611
Query: 1265 EALLHKGGDYASLVALHTSDS 1285
L+ K G Y LV + TS S
Sbjct: 612 SELIKKEGIYFRLVNMQTSGS 632
>MDR3_MOUSE (P21447) Multidrug resistance protein 3 (P-glycoprotein 3)
(MDR1A)
Length = 1276
Score = 946 bits (2445), Expect = 0.0
Identities = 515/1286 (40%), Positives = 788/1286 (61%), Gaps = 45/1286 (3%)
Query: 22 SNQDSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILIFGTMI 81
S + K+K V + +F +A DRL ML+GTL AI +G+++PLM+LIFG M
Sbjct: 16 SKMGKKSKKEKKEKKPAVSVLTMFRYAGWLDRLYMLVGTLAAIIHGVALPLMMLIFGDMT 75
Query: 82 NAFG-------DSTNSKVVDEVS-------ETTTYCDQVSLKFVYLAAGTFVASFLQLTC 127
++F +STN D+ + E TTY + + + AG + +++Q++
Sbjct: 76 DSFASVGNVSKNSTNMSEADKRAMFAKLEEEMTTY----AYYYTGIGAGVLIVAYIQVSF 131
Query: 128 WMITGERQSARIRGLYLKTILRQDVSFFDKETNTGEVVGRMSGDTVLIKDAMGEKVGQFI 187
W + RQ +IR + I+ Q++ +FD + GE+ R++ D I + +G+K+G F
Sbjct: 132 WCLAAGRQIHKIRQKFFHAIMNQEIGWFDVH-DVGELNTRLTDDVSKINEGIGDKIGMFF 190
Query: 188 QFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKSAGVV 247
Q M+TF GGF+I FT+GW LT+V+L+ P+L LS + + +++ + AY+K+ V
Sbjct: 191 QAMATFFGGFIIGFTRGWKLTLVILAISPVLGLSAGIWAKILSSFTDKELHAYAKAGAVA 250
Query: 248 EQTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGVGFGTLFFVFICSYGLAV 307
E+ + +IRTV +F G+K+ YN +L + + +++A+ + + G F + SY LA
Sbjct: 251 EEVLAAIRTVIAFGGQKKELERYNNNLEEAKRLGIKKAITANISMGAAFLLIYASYALAF 310
Query: 308 WFGGKMIIEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETINRKPE 367
W+G ++I K Y+ G V+TV F+VLIG+ +GQ SP++ AFA + AA+++F+ I+ KP
Sbjct: 311 WYGTSLVISKEYSIGQVLTVFFSVLIGAFSVGQASPNIEAFANARGAAYEVFKIIDNKPS 370
Query: 368 IDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGKS 427
ID++ SG K D+I+G++E +++ FSYP+R + I G +L + SG T ALVG SG GKS
Sbjct: 371 IDSFSKSGHKPDNIQGNLEFKNIHFSYPSRKEVQILKGLNLKVKSGQTVALVGNSGCGKS 430
Query: 428 TVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKDC 487
T V L++R YDP DG V IDG +++ ++++R+ IG+VSQEPVLF +I ENI YG++
Sbjct: 431 TTVQLMQRLYDPLDGMVSIDGQDIRTINVRYLREIIGVVSQEPVLFATTIAENIRYGRED 490
Query: 488 ATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLL 547
T +EI A + ANA FI KLP DT+VGE G QLSGGQKQR+AIARA++++P+ILLL
Sbjct: 491 VTMDEIEKAVKEANAYDFIMKLPHQFDTLVGERGAQLSGGQKQRIAIARALVRNPKILLL 550
Query: 548 DEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAE 607
DEATSALD ESE +VQ AL++ RTTIV+AHRLST+RN D IA G IVE+G+H E
Sbjct: 551 DEATSALDTESEAVVQAALDKAREGRTTIVIAHRLSTVRNADVIAGFDGGVIVEQGNHDE 610
Query: 608 LTNDPNGAYSQLIRLQEMKRSEQ--NDA-NDKNKPNSIVHSGRQSSQRSFSLRSISQGSA 664
L + G Y +L+ Q + N+A K++ +++ S + S RS +
Sbjct: 611 LMRE-KGIYFKLVMTQTAGNEIELGNEACKSKDEIDNLDMSSKDSGSSLIRRRSTRKSIC 669
Query: 665 GNSGRHSFSASYVAPTTDGFLETEDGGPQASPSKNSSPPEVPLYRLAYFNKPEIPVLLMG 724
G + D L T++ + + P +R+ N E P ++G
Sbjct: 670 GPHDQ------------DRKLSTKE-------ALDEDVPPASFWRILKLNSTEWPYFVVG 710
Query: 725 TITAVLHGAIMPVIGLLVSKMISTFYK--PADELRHDSKVWAIVFVAVAVASLLIIPCRF 782
A+++G + P ++ SK++ F P + R +S +++++F+ + + S + +
Sbjct: 711 IFCAIINGGLQPAFSVIFSKVVGVFTNGGPPETQRQNSNLFSLLFLILGIISFITFFLQG 770
Query: 783 YFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALG 842
+ FG AG L +R+R + F+ ++ +VSWFDD ++++GAL RL+ DAA V+ G L
Sbjct: 771 FTFGKAGEILTKRLRYMVFKSMLRQDVSWFDDPKNTTGALTTRLANDAAQVKGATGSRLA 830
Query: 843 LLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEAS 902
++ QNIA + G++I+ WQL ++LA+ P++ + G V++K+L G + KK E +
Sbjct: 831 VIFQNIANLGTGIIISLIYGWQLTLLLLAIVPIIAIAGVVEMKMLSGQALKDKKELEGSG 890
Query: 903 QVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDA 962
++A +A+ + RTV S E+K +Y Q + P + +++ + G+ F + M+Y A
Sbjct: 891 KIATEAIENFRTVVSLTREQKFETMYAQSLQIPYRNAMKKAHVFGITFFFTQAMMYFSYA 950
Query: 963 CVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQ 1022
F GA LV TF +V LVF A+ AM V Q + PD A +A+ I I+++
Sbjct: 951 ACFRFGAYLVTQQLMTFENVLLVFSAIVFGAMAVGQVSSFAPDYAKATVSASHIIRIIEK 1010
Query: 1023 KSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGS 1082
+IDS G+ ++G+++F+ F YPTR + + L L ++ G+T+ALVG SG
Sbjct: 1011 TPEIDSYSTQGLKPNMLEGNVQFSGFVFNYPTRPSIPVLQGLSLEVKKGQTLALVGSSGC 1070
Query: 1083 GKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYG 1142
GKSTV+ LL+RFYDP +G + LDG EI+++ V+WLR Q+G+VSQEPILF+ ++ NIAYG
Sbjct: 1071 GKSTVVQLLERFYDPMAGSVFLDGKEIKQLNVQWLRAQLGIVSQEPILFDCSIAENIAYG 1130
Query: 1143 KGGDATE-AEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNP 1201
EIV AA+ AN HQFI SL Y+T VG++G QLSGGQKQR+AIARA+V+ P
Sbjct: 1131 DNSRVVSYEEIVRAAKEANIHQFIDSLPDKYNTRVGDKGTQLSGGQKQRIAIARALVRQP 1190
Query: 1202 KILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEK 1261
ILLLDEATSALD ESEKVVQ+ALD+ RT I++AHRLSTI+ ADLI V++NG + E
Sbjct: 1191 HILLLDEATSALDTESEKVVQEALDKAREGRTCIVIAHRLSTIQNADLIVVIQNGKVKEH 1250
Query: 1262 GKHEALLHKGGDYASLVALHTSDSTS 1287
G H+ LL + G Y S+V++ S
Sbjct: 1251 GTHQQLLAQKGIYFSMVSVQAGAKRS 1276
>MDR3_CRIGR (P23174) Multidrug resistance protein 3 (P-glycoprotein 3)
Length = 1281
Score = 941 bits (2432), Expect = 0.0
Identities = 513/1297 (39%), Positives = 789/1297 (60%), Gaps = 63/1297 (4%)
Query: 19 DHDSNQDSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILIFG 78
D + S + ++K + LF ++D D+L ML+GT+ AI +G +PLM+++FG
Sbjct: 20 DFELGSISNQGRNKKKKVNLIGPLTLFRYSDWQDKLFMLLGTIMAIAHGSGLPLMMIVFG 79
Query: 79 TMINAFGDSTNS---------------KVVDEVSETTTYCDQVSLKFVYLAAGTFVASFL 123
M + F ++ + ++++E E T Y S L G VA+++
Sbjct: 80 EMTDKFVNNAGNFSLPVNFSLSMINPGRILEE--EMTRYAYYYS----GLGGGVLVAAYI 133
Query: 124 QLTCWMITGERQSARIRGLYLKTILRQDVSFFDKETNTGEVVGRMSGDTVLIKDAMGEKV 183
Q++ W + RQ +IR + ILRQ++ +FD + T E+ R++ D I + +G+KV
Sbjct: 134 QVSFWTLAAGRQIKKIRQNFFHAILRQEMGWFDIKGTT-ELNTRLTDDISKISEGIGDKV 192
Query: 184 GQFIQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKS 243
G F Q ++TF GF++ F +GW LT+V+++ P+L LS ++ + +++ S AAY+K+
Sbjct: 193 GMFFQAVATFFAGFIVGFIRGWKLTLVIMAISPILGLSAAVWAKILSTFSDKELAAYAKA 252
Query: 244 AGVVEQTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGVGFGTLFFVFICSY 303
V E+ +G+IRTV +F G+ + Y + L K +++A+++ + G F + SY
Sbjct: 253 GAVAEEALGAIRTVIAFGGQNKELERYQKHLENAKKIGIKKAISANISMGIAFLLIYASY 312
Query: 304 GLAVWFGGKMIIEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETIN 363
LA W+G ++I K YT G+ MTV F++LIG+ +GQ +P + AFA + AA+ +F+ I+
Sbjct: 313 ALAFWYGSTLVISKEYTIGNAMTVFFSILIGAFSVGQAAPCIDAFANARGAAYVIFDIID 372
Query: 364 RKPEIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSG 423
P+ID++ G K D I+G+++ DV FSYP+R + I G +L + SG T ALVG SG
Sbjct: 373 NNPKIDSFSERGHKPDSIKGNLDFSDVHFSYPSRANIKILKGLNLKVQSGQTVALVGNSG 432
Query: 424 SGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAY 483
GK+T + L++R YDPT+G + IDG +++ F ++++R+ IG+VSQEPVLF+ +I ENI Y
Sbjct: 433 CGKTTTLQLLQRLYDPTEGTISIDGQDIRNFNVRYLREIIGVVSQEPVLFSTTIAENIRY 492
Query: 484 GKDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPR 543
G+ T EEI+ A + ANA +FI KLPQ DT+VGE G QLSGGQKQR+AIARA++++P+
Sbjct: 493 GRGNVTMEEIKKAVKEANAYEFIMKLPQKFDTLVGERGAQLSGGQKQRIAIARALVRNPK 552
Query: 544 ILLLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERG 603
ILLLDEATSALD ESE VQ AL++ RTTIV+AHRLST+RN D IA G IVE+G
Sbjct: 553 ILLLDEATSALDTESEAEVQAALDKAREGRTTIVIAHRLSTVRNADVIAGFEDGVIVEQG 612
Query: 604 SHAELTNDPNGAYSQLIRLQ-----------EMKRSEQNDANDKNKPNSIVHSGRQSSQR 652
SH+EL G Y +L+ +Q E++ SE+ A D PN +
Sbjct: 613 SHSELMQK-EGVYFKLVNMQTSGSQILSQEFEVELSEEKAA-DGMTPNG---------WK 661
Query: 653 SFSLRSISQGSAGNSGRHSFSASYVAPTTDGFLETEDGGPQASPSKNSSPPEVPLYRLAY 712
S R+ ++ S +S H A D ++ P V ++
Sbjct: 662 SHIFRNSTKKSLKSSRAHHHRLDVDADELD-----------------ANVPPVSFLKVLK 704
Query: 713 FNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYKPADELRHDS-KVWAIVFVAVA 771
NK E P ++GT+ A+++GA+ P I +++S+MI+ F D ++ ++++VF+ +
Sbjct: 705 LNKTEWPYFVVGTVCAIVNGALQPAISIILSEMIAIFGPGDDAVKQQKCNLFSLVFLGLG 764
Query: 772 VASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAA 831
V S + + FG AG L R+R + F+ ++ ++SWFDD ++S+GAL RL+TD A
Sbjct: 765 VLSFFTFFLQGFTFGKAGEILTTRLRSMAFKAMLRQDMSWFDDYKNSTGALSTRLATDRA 824
Query: 832 SVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFS 891
V+ G L L+ QN A + G++I+F WQL ++L++ P + ++G V++K+L G +
Sbjct: 825 QVQGATGTRLALIAQNTANLGTGIIISFIYGWQLTLLLLSVVPFIAVSGIVEMKMLAGNA 884
Query: 892 ADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFG 951
KK E A ++A +A+ +IRTV S E K +Y +K P + V+ I G+ F
Sbjct: 885 KRDKKALEAAGKIATEAIENIRTVVSLTQERKFESMYVEKLHEPYRNSVQMAHIYGITFS 944
Query: 952 SSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKS 1011
S +Y A F GA L+ +G F DV LVF A+ A+ + + + PD AK
Sbjct: 945 ISQAFMYFSYAGCFRFGAYLIVNGHMRFRDVILVFSAIVFGAVALGHASSFAPDYAKAKL 1004
Query: 1012 AAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSG 1071
+AA +F++ +++ IDS G+ ++ +G + FN V F YPTR ++ + L L ++ G
Sbjct: 1005 SAAHLFSLFERQPLIDSYSGEGLWPDKFEGSVTFNEVVFNYPTRANMPVLQGLSLEVKKG 1064
Query: 1072 KTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILF 1131
+T+ALVG SG GKSTV+ LL+RFYDP +G + LDG E +++ ++WLR Q+G+VSQEP+LF
Sbjct: 1065 QTLALVGSSGCGKSTVVQLLERFYDPMAGTVLLDGQEAKKLNIQWLRAQLGIVSQEPVLF 1124
Query: 1132 NDTVRANIAYGKGGD-ATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQR 1190
+ ++ NIAYG ++ EIV AA+ AN H FI +L + Y T VG++G QLSGGQKQR
Sbjct: 1125 DCSIAENIAYGDNSRVVSQDEIVRAAKAANIHPFIETLPQKYKTRVGDKGTQLSGGQKQR 1184
Query: 1191 VAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLI 1250
+AI RA+++ P++LLLDEATSALD ESEKVVQ+ALD+ RT I++AHRLSTI+ ADLI
Sbjct: 1185 LAIRRALIRQPRVLLLDEATSALDTESEKVVQEALDKAREGRTCIVIAHRLSTIQNADLI 1244
Query: 1251 AVVKNGVIAEKGKHEALLHKGGDYASLVALHTSDSTS 1287
V++NG + E G H+ LL + G Y S+V + S
Sbjct: 1245 VVIQNGKVKEHGTHQQLLAQKGIYFSMVNIQAGAQNS 1281
>MDR1_MOUSE (P06795) Multidrug resistance protein 1 (P-glycoprotein 1)
Length = 1276
Score = 920 bits (2378), Expect = 0.0
Identities = 497/1273 (39%), Positives = 782/1273 (61%), Gaps = 35/1273 (2%)
Query: 22 SNQDSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILIFGTMI 81
S + K+K V ++ +F +AD D+L M++GTL AI +G +PL++L+FG M
Sbjct: 16 SKMGKKSKKEKKEKKPAVGVFGMFRYADWLDKLCMILGTLAAIIHGTLLPLLMLVFGNMT 75
Query: 82 NAFGDSTNSKVVDEVSET----TTYCDQVSLK---------FVYLAAGTFVASFLQLTCW 128
++F + S + +++ T SL+ + + AG + +++Q++ W
Sbjct: 76 DSFTKAEASILPSITNQSGPNSTLIISNSSLEEEMAIYAYYYTGIGAGVLIVAYIQVSLW 135
Query: 129 MITGERQSARIRGLYLKTILRQDVSFFDKETNTGEVVGRMSGDTVLIKDAMGEKVGQFIQ 188
+ RQ +IR + I+ Q++ +FD + GE+ R++ D I D +G+K+G F Q
Sbjct: 136 CLAAGRQIHKIRQKFFHAIMNQEIGWFDVH-DVGELNTRLTDDVSKINDGIGDKIGMFFQ 194
Query: 189 FMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKSAGVVE 248
++TF+ GF+I F GW LT+V+L+ PL+ LS ++ + V+ ++ AY+K+ V E
Sbjct: 195 SITTFLAGFIIGFISGWKLTLVILAVSPLIGLSSALWAKVLTSFTNKELQAYAKAGAVAE 254
Query: 249 QTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGVGFGTLFFVFICSYGLAVW 308
+ + +IRTV +F G+++ YN++L + +++A+ + + G + + SY LA W
Sbjct: 255 EVLAAIRTVIAFGGQQKELERYNKNLEEAKNVGIKKAITASISIGIAYLLVYASYALAFW 314
Query: 309 FGGKMIIEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETINRKPEI 368
+G +++ Y+ G+V+TV F++L+G+ +G +P++ AFA + AAF++F+ I+ +P I
Sbjct: 315 YGTSLVLSNEYSIGEVLTVFFSILLGTFSIGHLAPNIEAFANARGAAFEIFKIIDNEPSI 374
Query: 369 DAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGKST 428
D++ T G K D I G++E ++V F+YP+R + I G +L + SG T ALVG SG GKST
Sbjct: 375 DSFSTKGYKPDSIMGNLEFKNVHFNYPSRSEVQILKGLNLKVKSGQTVALVGNSGCGKST 434
Query: 429 VVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKDCA 488
V L++R YDP +G V IDG +++ ++++R+ IG+VSQEPVLF +I ENI YG++
Sbjct: 435 TVQLMQRLYDPLEGVVSIDGQDIRTINVRYLREIIGVVSQEPVLFATTIAENIRYGREDV 494
Query: 489 TDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLD 548
T +EI A + ANA FI KLP DT+VGE G QLSGGQKQR+AIARA++++P+ILLLD
Sbjct: 495 TMDEIEKAVKEANAYDFIMKLPHQFDTLVGERGAQLSGGQKQRIAIARALVRNPKILLLD 554
Query: 549 EATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAEL 608
EATSALD ESE +VQ AL++ RTTIV+AHRLST+RN D IA G IVE+G+H EL
Sbjct: 555 EATSALDTESEAVVQAALDKAREGRTTIVIAHRLSTVRNADVIAGFDGGVIVEQGNHDEL 614
Query: 609 TNDPNGAYSQLIRLQEMKRSEQNDANDKNKPNSIVHSGRQSSQRSFSLRSISQGSAGNSG 668
+ G Y +L+ M ++ N+ N G QS + L S+ S
Sbjct: 615 MRE-KGIYFKLV----MTQTRGNEIEPGNNA-----YGSQSDTDASEL--TSEESKSPLI 662
Query: 669 RHSFSASYVAPTTDGFLETEDGGPQASPSKNSSPPEVPLYRLAYFNKPEIPVLLMGTITA 728
R S S + ++ + + P V +R+ N E P LL+G + A
Sbjct: 663 RRSIYRSVHRK------QDQERRLSMKEAVDEDVPLVSFWRILNLNLSEWPYLLVGVLCA 716
Query: 729 VLHGAIMPVIGLLVSKMISTFYKPADE--LRHDSKVWAIVFVAVAVASLLIIPCRFYFFG 786
V++G I PV ++ S+++ F + D R + ++++ F+ + + S + + + FG
Sbjct: 717 VINGCIQPVFAIVFSRIVGVFSRDDDHETKRQNCNLFSLFFLVMGLISFVTYFFQGFTFG 776
Query: 787 VAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQ 846
AG L +R+R + F+ ++ ++SWFDD ++S+G+L RL++DA+SV+ +G L ++ Q
Sbjct: 777 KAGEILTKRVRYMVFKSMLRQDISWFDDHKNSTGSLTTRLASDASSVKGAMGARLAVVTQ 836
Query: 847 NIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVAN 906
N+A + G++++ WQL +++ + PL+ L G +++K+L G + KK E + ++A
Sbjct: 837 NVANLGTGVILSLVYGWQLTLLLVVIIPLIVLGGIIEMKLLSGQALKDKKQLEISGKIAT 896
Query: 907 DAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDACVFY 966
+A+ + RT+ S E+K +Y Q + P + +++ + G+ F + M+Y A F
Sbjct: 897 EAIENFRTIVSLTREQKFETMYAQSLQVPYRNAMKKAHVFGITFSFTQAMMYFSYAACFR 956
Query: 967 AGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQI 1026
GA LV TF +V LVF A+ AM + + PD AK +A+ I I+++ +I
Sbjct: 957 FGAYLVAQQLMTFENVMLVFSAVVFGAMAAGNTSSFAPDYAKAKVSASHIIRIIEKTPEI 1016
Query: 1027 DSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKST 1086
DS G+ ++G+++FN V F YPTR ++ + L L ++ G+T+ALVG SG GKST
Sbjct: 1017 DSYSTEGLKPTLLEGNVKFNGVQFNYPTRPNIPVLQGLSLEVKKGQTLALVGSSGCGKST 1076
Query: 1087 VISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGD 1146
V+ LL+RFYDP +G + LDG EI+++ V+WLR +G+VSQEPILF+ ++ NIAYG
Sbjct: 1077 VVQLLERFYDPMAGSVFLDGKEIKQLNVQWLRAHLGIVSQEPILFDCSIAENIAYGDNSR 1136
Query: 1147 A-TEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILL 1205
A + EIV AA+ AN HQFI SL Y+T VG++G QLSGGQKQR+AIARA+V+ P ILL
Sbjct: 1137 AVSHEEIVRAAKEANIHQFIDSLPDKYNTRVGDKGTQLSGGQKQRIAIARALVRQPHILL 1196
Query: 1206 LDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHE 1265
LDEATSALD ESEKVVQ+ALD+ RT I++AHRLSTI+ ADLI V++NG + E G H+
Sbjct: 1197 LDEATSALDTESEKVVQEALDKAREGRTCIVIAHRLSTIQNADLIVVIENGKVKEHGTHQ 1256
Query: 1266 ALLHKGGDYASLV 1278
LL + G Y S+V
Sbjct: 1257 QLLAQKGIYFSMV 1269
Score = 433 bits (1113), Expect = e-120
Identities = 230/614 (37%), Positives = 359/614 (58%), Gaps = 25/614 (4%)
Query: 691 GPQASPSKNSSPPEVPLYRL-AYFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTF 749
G ++ K P V ++ + Y + + +++GT+ A++HG ++P++ L+ M +F
Sbjct: 19 GKKSKKEKKEKKPAVGVFGMFRYADWLDKLCMILGTLAAIIHGTLLPLLMLVFGNMTDSF 78
Query: 750 YKPA---------------------DELRHDSKVWAIVFVAVAVASLLIIPCRFYFFGVA 788
K L + ++A + + L++ + + +A
Sbjct: 79 TKAEASILPSITNQSGPNSTLIISNSSLEEEMAIYAYYYTGIGAGVLIVAYIQVSLWCLA 138
Query: 789 GGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQNI 848
G+ I +IR+ F +++ E+ WFD H G L RL+ D + + +GD +G+ Q+I
Sbjct: 139 AGRQIHKIRQKFFHAIMNQEIGWFD--VHDVGELNTRLTDDVSKINDGIGDKIGMFFQSI 196
Query: 849 ATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDA 908
T + G +I F + W+L ++LA++PL+GL+ + KVL F+ + Y +A VA +
Sbjct: 197 TTFLAGFIIGFISGWKLTLVILAVSPLIGLSSALWAKVLTSFTNKELQAYAKAGAVAEEV 256
Query: 909 VGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDACVFYAG 968
+ +IRTV +F ++K +E Y + E G+++ I + + G ++ ++YA A F+ G
Sbjct: 257 LAAIRTVIAFGGQQKELERYNKNLEEAKNVGIKKAITASISIGIAYLLVYASYALAFWYG 316
Query: 969 ARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDS 1028
LV + + +V VFF++ + + + NA+ AA IF I+D + IDS
Sbjct: 317 TSLVLSNEYSIGEVLTVFFSILLGTFSIGHLAPNIEAFANARGAAFEIFKIIDNEPSIDS 376
Query: 1029 SDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVI 1088
G + + G++EF +V F YP+R +VQI L L ++SG+TVALVG SG GKST +
Sbjct: 377 FSTKGYKPDSIMGNLEFKNVHFNYPSRSEVQILKGLNLKVKSGQTVALVGNSGCGKSTTV 436
Query: 1089 SLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDAT 1148
L+QR YDP G +++DG +I+ + V++LR+ +G+VSQEP+LF T+ NI YG+ D T
Sbjct: 437 QLMQRLYDPLEGVVSIDGQDIRTINVRYLREIIGVVSQEPVLFATTIAENIRYGR-EDVT 495
Query: 1149 EAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDE 1208
EI A + ANA+ FI L +DT+VGERG QLSGGQKQR+AIARA+V+NPKILLLDE
Sbjct: 496 MDEIEKAVKEANAYDFIMKLPHQFDTLVGERGAQLSGGQKQRIAIARALVRNPKILLLDE 555
Query: 1209 ATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALL 1268
ATSALD ESE VVQ ALD+ RTTI++AHRLST++ AD+IA GVI E+G H+ L+
Sbjct: 556 ATSALDTESEAVVQAALDKAREGRTTIVIAHRLSTVRNADVIAGFDGGVIVEQGNHDELM 615
Query: 1269 HKGGDYASLVALHT 1282
+ G Y LV T
Sbjct: 616 REKGIYFKLVMTQT 629
Score = 399 bits (1024), Expect = e-110
Identities = 230/612 (37%), Positives = 352/612 (56%), Gaps = 13/612 (2%)
Query: 13 SVQPVEDHDSNQDSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPL 72
SV +D + +++ D+DV V +++ + + S+ +L+G L A+ NG P+
Sbjct: 669 SVHRKQDQERRLSMKEAVDEDVPL--VSFWRILNL-NLSEWPYLLVGVLCAVINGCIQPV 725
Query: 73 MILIFGTMINAFGDSTNSKVVDEVSETTTYCDQVSLKFVYLAAGTFVASFLQLTCWMITG 132
++F ++ F D+ C+ SL F+ + +FV F Q + G
Sbjct: 726 FAIVFSRIVGVFSRD------DDHETKRQNCNLFSLFFLVMGLISFVTYFFQGFTFGKAG 779
Query: 133 ERQSARIRGLYLKTILRQDVSFFDKETN-TGEVVGRMSGDTVLIKDAMGEKVGQFIQFMS 191
E + R+R + K++LRQD+S+FD N TG + R++ D +K AMG ++ Q ++
Sbjct: 780 EILTKRVRYMVFKSMLRQDISWFDDHKNSTGSLTTRLASDASSVKGAMGARLAVVTQNVA 839
Query: 192 TFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKSAGVVEQTI 251
G +++ GW LT++++ IPL++L G + +++ + + S + + I
Sbjct: 840 NLGTGVILSLVYGWQLTLLLVVIIPLIVLGGIIEMKLLSGQALKDKKQLEISGKIATEAI 899
Query: 252 GSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGVGFGTLFFVFICSYGLAVWFGG 311
+ RT+ S T E++ Y +SL Y+ A+++A G+ F + SY FG
Sbjct: 900 ENFRTIVSLTREQKFETMYAQSLQVPYRNAMKKAHVFGITFSFTQAMMYFSYAACFRFGA 959
Query: 312 KMIIEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETINRKPEIDAY 371
++ ++ T +VM V AV+ G+ G TS +A + +A + I + PEID+Y
Sbjct: 960 YLVAQQLMTFENVMLVFSAVVFGAMAAGNTSSFAPDYAKAKVSASHIIRIIEKTPEIDSY 1019
Query: 372 DTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVS 431
T G K + G+++ V F+YPTRP+ + G SL + G T ALVG SG GKSTVV
Sbjct: 1020 STEGLKPTLLEGNVKFNGVQFNYPTRPNIPVLQGLSLEVKKGQTLALVGSSGCGKSTVVQ 1079
Query: 432 LIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKD--CAT 489
L+ERFYDP G V +DG +K+ ++W+R +G+VSQEP+LF CSI ENIAYG + +
Sbjct: 1080 LLERFYDPMAGSVFLDGKEIKQLNVQWLRAHLGIVSQEPILFDCSIAENIAYGDNSRAVS 1139
Query: 490 DEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLDE 549
EEI AA+ AN +FID LP +T VG+ GTQLSGGQKQR+AIARA+++ P ILLLDE
Sbjct: 1140 HEEIVRAAKEANIHQFIDSLPDKYNTRVGDKGTQLSGGQKQRIAIARALVRQPHILLLDE 1199
Query: 550 ATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAELT 609
ATSALD ESE++VQEAL++ RT IV+AHRLSTI+N D I VI GK+ E G+H +L
Sbjct: 1200 ATSALDTESEKVVQEALDKAREGRTCIVIAHRLSTIQNADLIVVIENGKVKEHGTHQQLL 1259
Query: 610 NDPNGAYSQLIR 621
G Y +++
Sbjct: 1260 AQ-KGIYFSMVQ 1270
>MDR2_CRIGR (P21449) Multidrug resistance protein 2 (P-glycoprotein 2)
Length = 1276
Score = 910 bits (2353), Expect = 0.0
Identities = 496/1290 (38%), Positives = 782/1290 (60%), Gaps = 37/1290 (2%)
Query: 5 IRLEGDFVSVQPVEDHDSNQDSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAI 64
+ E DF S + +D K + K+ V ++ +F +AD D+L M++GTL A+
Sbjct: 1 MEFEEDF-SARADKDFLKMGRKSKKEKKEKENPNVGIFGMFRYADWLDKLYMVLGTLAAV 59
Query: 65 GNGLSIPLMILIFGTMINAFGDS--------TNSKVVD--EVSETTTYCDQVSLKFVY-- 112
+G S+PL++L+FG M ++F + TN ++ EV + D + + Y
Sbjct: 60 LHGTSLPLLMLVFGNMTDSFTKAETSIWPNMTNQSEINNTEVISGSLEEDMATYAYYYTG 119
Query: 113 LAAGTFVASFLQLTCWMITGERQSARIRGLYLKTILRQDVSFFDKETNTGEVVGRMSGDT 172
+ AG + +++Q++ W + RQ +IR + I+ Q++ +FD + GE+ R++ D
Sbjct: 120 IGAGVLIVAYIQVSFWCLAAGRQINKIRQKFFHAIMNQEIGWFDVH-DIGELNTRLTDDV 178
Query: 173 VLIKDAMGEKVGQFIQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIAKA 232
I D +G+K+G F Q ++TF+ F++ F GW LT+V+L+ PL+ LS +M + V+
Sbjct: 179 SKINDGIGDKIGMFFQSIATFLAAFIVGFISGWKLTLVILAVSPLIGLSSAMWAKVLTSF 238
Query: 233 SSTGQAAYSKSAGVVEQTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGVGF 292
++ AY+K+ V E+ + +IRTV +F G+ + YN++L + +++A+ + +
Sbjct: 239 TNKELQAYAKAGAVAEEVLAAIRTVIAFGGQNKELERYNKNLEEAKNVGIKKAVTANISI 298
Query: 293 GTLFFVFICSYGLAVWFGGKMIIEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQ 352
G + + SY LA W+G +++ Y+ G V+TV F++L G+ +G +P++ FA +
Sbjct: 299 GIAYLLVYASYALAFWYGTSLVLSNEYSVGQVLTVFFSILFGTFSIGHIAPNIEVFANAR 358
Query: 353 AAAFKMFETINRKPEIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPS 412
AA+++F+ I+ +P ID++ T G K D + G++E ++V FSYP+R I G +L + S
Sbjct: 359 GAAYEIFKIIDNEPSIDSFSTQGHKPDSVMGNLEFKNVHFSYPSRSGIKILKGLNLKVQS 418
Query: 413 GTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVL 472
G T ALVG+SG GKST V L++R YDPT+G V IDG +++ ++++R+ IG+VSQEPVL
Sbjct: 419 GQTVALVGKSGCGKSTTVQLLQRLYDPTEGVVSIDGQDIRTINVRYLREIIGVVSQEPVL 478
Query: 473 FTCSIKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRV 532
F +I ENI YG++ T +EI A + ANA FI KLP DT+VGE G QLSGGQKQR+
Sbjct: 479 FATTIAENIRYGRENVTMDEIEKAVKEANAYDFIMKLPHKFDTLVGERGAQLSGGQKQRI 538
Query: 533 AIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIA 592
AIARA++++P+ILLLDEATSALD ESE +VQ AL++ RTTIV+AHRLST+RN D IA
Sbjct: 539 AIARALVRNPKILLLDEATSALDTESEAVVQAALDKAREGRTTIVIAHRLSTVRNADVIA 598
Query: 593 VIHQGKIVERGSHAELTNDPNGAYSQLIRLQEM-KRSEQNDANDKNKPNSIVHSGRQSSQ 651
G IVE+G+H EL + G Y +L+ +Q E D ++ ++I
Sbjct: 599 GFDGGVIVEQGNHEELMKE-KGIYCRLVMMQTRGNEVELGSEADGSQSDTIASELTSEEF 657
Query: 652 RSFSLRSISQGSAGNSGRHSFSASYVAPTTDGFLETEDGGPQASPSKNSSPPEVPLYRLA 711
+S S+R + S S + ++ +++ P V + +
Sbjct: 658 KSPSVRKSTCRSICGS------------------QDQERRVSVKEAQDEDVPLVSFWGIL 699
Query: 712 YFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYKPAD--ELRHDSKVWAIVFVA 769
N E P L++G + AV++G + PV ++ S +I F + D + + ++++ F+
Sbjct: 700 KLNITEWPYLVVGVLCAVINGCMQPVFSIVFSGIIGVFTRDDDPKTKQQNCNLFSLFFLV 759
Query: 770 VAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTD 829
+ + + + + FG AG L +R+R + F+ ++ ++SWFDD +S+GAL RL++D
Sbjct: 760 MGMICFVTYFFQGFTFGKAGEILTKRLRYMVFKSMLRQDISWFDDHRNSTGALTTRLASD 819
Query: 830 AASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKG 889
AA+V+ + L + QN+A + G++I+ WQL +++ +APL+ L+G +++KVL G
Sbjct: 820 AANVKGAMSSRLAGITQNVANLGTGIIISLVYGWQLTLLLVVIAPLIILSGMMEMKVLSG 879
Query: 890 FSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLG 949
+ KK E + ++A +A+ + RTV S E+K +Y Q + P + +++ + G+
Sbjct: 880 QALKDKKELEVSGKIATEAIENFRTVVSLTREQKFENMYAQSLQIPYRNALKKAHVFGIT 939
Query: 950 FGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNA 1009
F + M+Y A F GA LV TF +V LVF A+ A+ + + PD A
Sbjct: 940 FSFTQAMMYFSYAACFRFGAYLVAHQIMTFENVMLVFSAVVFGAIAAGNASSFAPDYAKA 999
Query: 1010 KSAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIR 1069
K +A+ I I+++ IDS G+ ++G+++FN V F YPTR D+ + L L ++
Sbjct: 1000 KVSASHIIRIMEKIPSIDSYSTRGLKPNWLEGNVKFNEVVFNYPTRPDIPVLQGLSLEVK 1059
Query: 1070 SGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPI 1129
G+T+ALVG SG GKSTV+ LL+RFYDP +G + LDG EI+++ V+WLR +G+VSQEPI
Sbjct: 1060 KGQTLALVGSSGCGKSTVVQLLERFYDPMAGTVFLDGKEIKQLNVQWLRAHLGIVSQEPI 1119
Query: 1130 LFNDTVRANIAYGKGGD-ATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQK 1188
LF+ ++ NIAYG ++ EI AA+ AN HQFI SL Y+T VG++G QLSGGQK
Sbjct: 1120 LFDCSIAENIAYGDNSRVVSQDEIERAAKEANIHQFIESLPDKYNTRVGDKGTQLSGGQK 1179
Query: 1189 QRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGAD 1248
QR+AIARA+V+ P ILLLDEATSALD ESEKVVQ+ALD+ RT I++AHRLSTI+ AD
Sbjct: 1180 QRIAIARALVRQPHILLLDEATSALDTESEKVVQEALDKAREGRTCIVIAHRLSTIQNAD 1239
Query: 1249 LIAVVKNGVIAEKGKHEALLHKGGDYASLV 1278
LI V++NG + E G H+ LL + G Y S+V
Sbjct: 1240 LIVVIQNGKVKEHGTHQQLLAQKGIYFSMV 1269
Score = 429 bits (1104), Expect = e-119
Identities = 236/632 (37%), Positives = 361/632 (56%), Gaps = 26/632 (4%)
Query: 672 FSASYVAPTTDGFLETEDGGPQASPSKNSSPPEVPLYRL-AYFNKPEIPVLLMGTITAVL 730
F + A FL+ G K P V ++ + Y + + +++GT+ AVL
Sbjct: 3 FEEDFSARADKDFLKM--GRKSKKEKKEKENPNVGIFGMFRYADWLDKLYMVLGTLAAVL 60
Query: 731 HGAIMPVIGLLVSKMISTFYKP--------------------ADELRHDSKVWAIVFVAV 770
HG +P++ L+ M +F K + L D +A + +
Sbjct: 61 HGTSLPLLMLVFGNMTDSFTKAETSIWPNMTNQSEINNTEVISGSLEEDMATYAYYYTGI 120
Query: 771 AVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDA 830
L++ + F+ +A G+ I +IR+ F +++ E+ WFD H G L RL+ D
Sbjct: 121 GAGVLIVAYIQVSFWCLAAGRQINKIRQKFFHAIMNQEIGWFD--VHDIGELNTRLTDDV 178
Query: 831 ASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGF 890
+ + +GD +G+ Q+IAT + ++ F + W+L ++LA++PL+GL+ + KVL F
Sbjct: 179 SKINDGIGDKIGMFFQSIATFLAAFIVGFISGWKLTLVILAVSPLIGLSSAMWAKVLTSF 238
Query: 891 SADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGF 950
+ + Y +A VA + + +IRTV +F + K +E Y + E G+++ + + +
Sbjct: 239 TNKELQAYAKAGAVAEEVLAAIRTVIAFGGQNKELERYNKNLEEAKNVGIKKAVTANISI 298
Query: 951 GSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAK 1010
G ++ ++YA A F+ G LV + + V VFF++ + + NA+
Sbjct: 299 GIAYLLVYASYALAFWYGTSLVLSNEYSVGQVLTVFFSILFGTFSIGHIAPNIEVFANAR 358
Query: 1011 SAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRS 1070
AA IF I+D + IDS G + V G++EF +V F YP+R ++I L L ++S
Sbjct: 359 GAAYEIFKIIDNEPSIDSFSTQGHKPDSVMGNLEFKNVHFSYPSRSGIKILKGLNLKVQS 418
Query: 1071 GKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPIL 1130
G+TVALVG+SG GKST + LLQR YDP G +++DG +I+ + V++LR+ +G+VSQEP+L
Sbjct: 419 GQTVALVGKSGCGKSTTVQLLQRLYDPTEGVVSIDGQDIRTINVRYLREIIGVVSQEPVL 478
Query: 1131 FNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQR 1190
F T+ NI YG+ + T EI A + ANA+ FI L +DT+VGERG QLSGGQKQR
Sbjct: 479 FATTIAENIRYGRE-NVTMDEIEKAVKEANAYDFIMKLPHKFDTLVGERGAQLSGGQKQR 537
Query: 1191 VAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLI 1250
+AIARA+V+NPKILLLDEATSALD ESE VVQ ALD+ RTTI++AHRLST++ AD+I
Sbjct: 538 IAIARALVRNPKILLLDEATSALDTESEAVVQAALDKAREGRTTIVIAHRLSTVRNADVI 597
Query: 1251 AVVKNGVIAEKGKHEALLHKGGDYASLVALHT 1282
A GVI E+G HE L+ + G Y LV + T
Sbjct: 598 AGFDGGVIVEQGNHEELMKEKGIYCRLVMMQT 629
>MDR1_RAT (P43245) Multidrug resistance protein 1 (P-glycoprotein 1)
Length = 1277
Score = 903 bits (2333), Expect = 0.0
Identities = 493/1282 (38%), Positives = 787/1282 (60%), Gaps = 45/1282 (3%)
Query: 19 DHDSNQDSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILIFG 78
D + ++ +KSK + V ++ +F +AD D+L M +GTL AI +G +PL++L+FG
Sbjct: 12 DKNFSKMGKKSKKEKEKKPAVGIFGMFRYADWLDKLCMALGTLAAIIHGTLLPLLMLVFG 71
Query: 79 TMINAFGDS---------TNSKVVDE---VSETTTYCDQVSLKFVY--LAAGTFVASFLQ 124
M ++F S TN ++ VS+T+ D + Y + AG + +++Q
Sbjct: 72 YMTDSFTPSRDPHSDRAITNQSEINSTHTVSDTSLEEDMAMYAYYYTGIGAGVLIVAYIQ 131
Query: 125 LTCWMITGERQSARIRGLYLKTILRQDVSFFDKETNTGEVVGRMSGDTVLIKDAMGEKVG 184
++ W + RQ +IR + I+ Q++ +FD + GE+ R++ D I D +G+K+G
Sbjct: 132 VSLWCLAAGRQIHKIRQKFFHAIMNQEIGWFDVN-DAGELNTRLTDDVSKINDGIGDKLG 190
Query: 185 QFIQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKSA 244
F Q ++TF GF+I F GW LT+V+L+ PL+ LS +M + V+ ++ AY+K+
Sbjct: 191 MFFQSITTFSAGFIIGFISGWKLTLVILAVSPLIGLSSAMWAKVLTSFTNKELQAYAKAG 250
Query: 245 GVVEQTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGVGFGTLFFVFICSYG 304
V E+ + +IRTV +F G+K+ YN++L + + +++A+ + + G + + SY
Sbjct: 251 AVAEEVLAAIRTVIAFGGQKKELERYNKNLEEAKRVGIKKAITANISIGIAYLLVYASYA 310
Query: 305 LAVWFGGKMIIEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETINR 364
LA W+G +++ Y+ G V+TV F++L+G+ +G +P++ AFA + AA+++F+ I+
Sbjct: 311 LAFWYGTSLVLSNEYSIGQVLTVFFSILLGTFSIGHLAPNIEAFANARGAAYEIFKIIDN 370
Query: 365 KPEIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGS 424
+P ID++ T G K D I G++E ++V F+YP+R + I G +L + SG T ALVG SG
Sbjct: 371 EPSIDSFSTKGHKPDSIMGNLEFKNVYFNYPSRSEVKILKGLNLKVKSGQTVALVGNSGC 430
Query: 425 GKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYG 484
GKST V L++R YDP +GEV IDG +++ ++++R+ IG+VSQEPVLF +I ENI YG
Sbjct: 431 GKSTTVQLLQRLYDPIEGEVSIDGQDIRTINVRYLREIIGVVSQEPVLFATTIAENIRYG 490
Query: 485 KDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRI 544
++ T +EI A + ANA FI KLP DT+VGE G QLSGGQKQR+AIARA++++P+I
Sbjct: 491 RENVTMDEIEKAVKEANAYDFIMKLPHKFDTLVGERGAQLSGGQKQRIAIARALVRNPKI 550
Query: 545 LLLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGS 604
LLLDEATSALD ESE +VQ AL++ RTTIV+AHRLST+RN D IA G IVE+G+
Sbjct: 551 LLLDEATSALDTESEAVVQAALDKAREGRTTIVIAHRLSTVRNADVIAGFDGGVIVEQGN 610
Query: 605 HAELTNDPNGAYSQLIRLQEMKRSEQNDANDKNKPNSIVHSGRQSSQRSFS---LRSISQ 661
H EL + G Y +L+ + + + +E N+ + S + +S+ S S RSI
Sbjct: 611 HEELMKE-KGIYFKLV-MTQTRGNEIEPGNNAYESQSDTGASELTSEESKSPLIRRSI-- 666
Query: 662 GSAGNSGRHSFSASYVAPTTDGFLETEDGGPQASPSKNSSPPEVPLYRLAYFNKPEIPVL 721
R S + ++ + + P V +++ N E P L
Sbjct: 667 -------RRSIHRR----------QDQERRLSSKEDVDEDVPMVSFWQILKLNISEWPYL 709
Query: 722 LMGTITAVLHGAIMPVIGLLVSKMISTFYKPADE--LRHDSKVWAIVFVAVAVASLLIIP 779
++G + AV++G I PV ++ SK++ F + D + + +++++F+ + + S +
Sbjct: 710 VVGVLCAVINGCIQPVFAIVFSKIVGVFSRDDDHETKQRNCNLFSLLFLVMGMISFVTYF 769
Query: 780 CRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGD 839
+ + FG AG L +R+R + F+ ++ ++SWFDD ++++G+L RL++DA++V+ +G
Sbjct: 770 FQGFTFGKAGEILTKRLRYMVFKSMLRQDISWFDDHKNTTGSLTTRLASDASNVKGAMGS 829
Query: 840 ALGLLVQNIATIIVGMVIAFQA--SWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKL 897
L ++ QN+A + G++++ WQL +++ + PL+ L G +++K+L G + KK
Sbjct: 830 RLAVVTQNVANLGTGIILSLVLVYGWQLTLLLVVIIPLIVLGGIIEMKLLSGQALKDKKE 889
Query: 898 YEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSSFFML 957
E + ++A +A+ + RTV S E+K +Y Q + P + +++ + G+ F + M+
Sbjct: 890 LEISGKIATEAIENFRTVVSLTREQKFETMYAQSLQIPYRNALKKAHVFGITFAFTQAMI 949
Query: 958 YAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIF 1017
Y A F GA LV TF +V LVF A+ AM + + PD AK +A+ I
Sbjct: 950 YFSYAACFRFGAYLVARELMTFENVMLVFSAVVFGAMAAGNTSSFAPDYAKAKVSASHII 1009
Query: 1018 AILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALV 1077
I+++ +IDS G+ ++G+++FN V F YPTR ++ + L ++ G+T+ LV
Sbjct: 1010 GIIEKIPEIDSYSTEGLKPNWLEGNVKFNGVKFNYPTRPNIPVLQGLSFEVKKGQTLRLV 1069
Query: 1078 GESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRA 1137
G SG GKSTV+ LL+RFY+P +G + LDG EI+++ V+ +R +G+VSQEPILF+ ++
Sbjct: 1070 GSSGCGKSTVVQLLERFYNPMAGTVFLDGKEIKQLNVQCVR-ALGIVSQEPILFDCSIAE 1128
Query: 1138 NIAYGKGGD-ATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARA 1196
NIAYG + EIV AA AN HQFI SL + Y+T VG++G QLSGGQKQR+AIARA
Sbjct: 1129 NIAYGDNSRVVSHEEIVRAAREANIHQFIDSLPEKYNTRVGDKGTQLSGGQKQRIAIARA 1188
Query: 1197 IVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNG 1256
+V+ P ILLLDEATSALD ESEKVVQ+ALD+ RT +++AHRLSTI+ ADLI V++NG
Sbjct: 1189 LVRQPHILLLDEATSALDTESEKVVQEALDKAREGRTCVVIAHRLSTIQNADLIVVIQNG 1248
Query: 1257 VIAEKGKHEALLHKGGDYASLV 1278
+ E G H+ LL + G Y S+V
Sbjct: 1249 QVKEHGTHQQLLAQKGIYFSMV 1270
Score = 426 bits (1095), Expect = e-118
Identities = 226/584 (38%), Positives = 346/584 (58%), Gaps = 25/584 (4%)
Query: 721 LLMGTITAVLHGAIMPVIGLLVSKMISTFYKPAD----------------------ELRH 758
+ +GT+ A++HG ++P++ L+ M +F D L
Sbjct: 49 MALGTLAAIIHGTLLPLLMLVFGYMTDSFTPSRDPHSDRAITNQSEINSTHTVSDTSLEE 108
Query: 759 DSKVWAIVFVAVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHS 818
D ++A + + L++ + + +A G+ I +IR+ F +++ E+ WFD +
Sbjct: 109 DMAMYAYYYTGIGAGVLIVAYIQVSLWCLAAGRQIHKIRQKFFHAIMNQEIGWFD--VND 166
Query: 819 SGALGARLSTDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGL 878
+G L RL+ D + + +GD LG+ Q+I T G +I F + W+L ++LA++PL+GL
Sbjct: 167 AGELNTRLTDDVSKINDGIGDKLGMFFQSITTFSAGFIIGFISGWKLTLVILAVSPLIGL 226
Query: 879 NGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKK 938
+ + KVL F+ + Y +A VA + + +IRTV +F ++K +E Y + E +
Sbjct: 227 SSAMWAKVLTSFTNKELQAYAKAGAVAEEVLAAIRTVIAFGGQKKELERYNKNLEEAKRV 286
Query: 939 GVRRGIISGLGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQ 998
G+++ I + + G ++ ++YA A F+ G LV + + V VFF++ + +
Sbjct: 287 GIKKAITANISIGIAYLLVYASYALAFWYGTSLVLSNEYSIGQVLTVFFSILLGTFSIGH 346
Query: 999 SGTLVPDSTNAKSAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDV 1058
+ NA+ AA IF I+D + IDS G + + G++EF +V F YP+R +V
Sbjct: 347 LAPNIEAFANARGAAYEIFKIIDNEPSIDSFSTKGHKPDSIMGNLEFKNVYFNYPSRSEV 406
Query: 1059 QIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLR 1118
+I L L ++SG+TVALVG SG GKST + LLQR YDP G +++DG +I+ + V++LR
Sbjct: 407 KILKGLNLKVKSGQTVALVGNSGCGKSTTVQLLQRLYDPIEGEVSIDGQDIRTINVRYLR 466
Query: 1119 QQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQKGYDTIVGE 1178
+ +G+VSQEP+LF T+ NI YG+ + T EI A + ANA+ FI L +DT+VGE
Sbjct: 467 EIIGVVSQEPVLFATTIAENIRYGR-ENVTMDEIEKAVKEANAYDFIMKLPHKFDTLVGE 525
Query: 1179 RGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVA 1238
RG QLSGGQKQR+AIARA+V+NPKILLLDEATSALD ESE VVQ ALD+ RTTI++A
Sbjct: 526 RGAQLSGGQKQRIAIARALVRNPKILLLDEATSALDTESEAVVQAALDKAREGRTTIVIA 585
Query: 1239 HRLSTIKGADLIAVVKNGVIAEKGKHEALLHKGGDYASLVALHT 1282
HRLST++ AD+IA GVI E+G HE L+ + G Y LV T
Sbjct: 586 HRLSTVRNADVIAGFDGGVIVEQGNHEELMKEKGIYFKLVMTQT 629
Score = 375 bits (962), Expect = e-103
Identities = 224/614 (36%), Positives = 348/614 (56%), Gaps = 16/614 (2%)
Query: 13 SVQPVEDHDSNQDSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPL 72
S+ +D + S++ D+DV V +++ + S+ +++G L A+ NG P+
Sbjct: 669 SIHRRQDQERRLSSKEDVDEDVPM--VSFWQILKL-NISEWPYLVVGVLCAVINGCIQPV 725
Query: 73 MILIFGTMINAFGDSTNSKVVDEVSETTTYCDQVSLKFVYLAAGTFVASFLQLTCWMITG 132
++F ++ F D+ C+ SL F+ + +FV F Q + G
Sbjct: 726 FAIVFSKIVGVFSRD------DDHETKQRNCNLFSLLFLVMGMISFVTYFFQGFTFGKAG 779
Query: 133 ERQSARIRGLYLKTILRQDVSFFDKETNT-GEVVGRMSGDTVLIKDAMGEKVGQFIQFMS 191
E + R+R + K++LRQD+S+FD NT G + R++ D +K AMG ++ Q ++
Sbjct: 780 EILTKRLRYMVFKSMLRQDISWFDDHKNTTGSLTTRLASDASNVKGAMGSRLAVVTQNVA 839
Query: 192 TFIGGFVIAFTK--GWLLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKSAGVVEQ 249
G +++ GW LT++++ IPL++L G + +++ + + S + +
Sbjct: 840 NLGTGIILSLVLVYGWQLTLLLVVIIPLIVLGGIIEMKLLSGQALKDKKELEISGKIATE 899
Query: 250 TIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGVGFGTLFFVFICSYGLAVWF 309
I + RTV S T E++ Y +SL Y+ A+++A G+ F + SY F
Sbjct: 900 AIENFRTVVSLTREQKFETMYAQSLQIPYRNALKKAHVFGITFAFTQAMIYFSYAACFRF 959
Query: 310 GGKMIIEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETINRKPEID 369
G ++ + T +VM V AV+ G+ G TS +A + +A + I + PEID
Sbjct: 960 GAYLVARELMTFENVMLVFSAVVFGAMAAGNTSSFAPDYAKAKVSASHIIGIIEKIPEID 1019
Query: 370 AYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGKSTV 429
+Y T G K + + G+++ V F+YPTRP+ + G S + G T LVG SG GKSTV
Sbjct: 1020 SYSTEGLKPNWLEGNVKFNGVKFNYPTRPNIPVLQGLSFEVKKGQTLRLVGSSGCGKSTV 1079
Query: 430 VSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKD--C 487
V L+ERFY+P G V +DG +K+ ++ +R +G+VSQEP+LF CSI ENIAYG +
Sbjct: 1080 VQLLERFYNPMAGTVFLDGKEIKQLNVQCVR-ALGIVSQEPILFDCSIAENIAYGDNSRV 1138
Query: 488 ATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLL 547
+ EEI AA AN +FID LP+ +T VG+ GTQLSGGQKQR+AIARA+++ P ILLL
Sbjct: 1139 VSHEEIVRAAREANIHQFIDSLPEKYNTRVGDKGTQLSGGQKQRIAIARALVRQPHILLL 1198
Query: 548 DEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAE 607
DEATSALD ESE++VQEAL++ RT +V+AHRLSTI+N D I VI G++ E G+H +
Sbjct: 1199 DEATSALDTESEKVVQEALDKAREGRTCVVIAHRLSTIQNADLIVVIQNGQVKEHGTHQQ 1258
Query: 608 LTNDPNGAYSQLIR 621
L G Y +++
Sbjct: 1259 LLAQ-KGIYFSMVQ 1271
>AB11_HUMAN (O95342) Bile salt export pump (ATP-binding cassette,
sub-family B, member 11)
Length = 1321
Score = 868 bits (2242), Expect = 0.0
Identities = 494/1295 (38%), Positives = 761/1295 (58%), Gaps = 38/1295 (2%)
Query: 18 EDHDSNQDSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILIF 77
+ +++++ S +K V ++LF F+ +D LM +G+L A +G++ P ++LIF
Sbjct: 24 KSYNNDKKSRLQDEKKGDGVRVGFFQLFRFSSSTDIWLMFVGSLCAFLHGIAQPGVLLIF 83
Query: 78 GTMINAFGDS---------------------TNSKVVDEVSETTTY----CDQVSLKFVY 112
GTM + F D TNS + ++ T + +KF
Sbjct: 84 GTMTDVFIDYDVELQELQIPGKACVNNTIVWTNSSLNQNMTNGTRCGLLNIESEMIKFAS 143
Query: 113 LAAGTFVA----SFLQLTCWMITGERQSARIRGLYLKTILRQDVSFFDKETNTGEVVGRM 168
AG VA ++Q+ W+I RQ ++R Y + I+R ++ +FD + GE+ R
Sbjct: 144 YYAGIAVAVLITGYIQICFWVIAAARQIQKMRKFYFRRIMRMEIGWFDCNS-VGELNTRF 202
Query: 169 SGDTVLIKDAMGEKVGQFIQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMV 228
S D I DA+ +++ FIQ M++ I GF++ F +GW LT+V++S PL+ + + +
Sbjct: 203 SDDINKINDAIADQMALFIQRMTSTICGFLLGFFRGWKLTLVIISVSPLIGIGAATIGLS 262
Query: 229 IAKASSTGQAAYSKSAGVVEQTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALAS 288
++K + AY+K+ V ++ I S+RTVA+F GEK+ Y ++L+ + +++ +
Sbjct: 263 VSKFTDYELKAYAKAGVVADEVISSMRTVAAFGGEKREVERYEKNLVFAQRWGIRKGIVM 322
Query: 289 GVGFGTLFFVFICSYGLAVWFGGKMIIEKG-YTGGDVMTVIFAVLIGSTCLGQTSPSLSA 347
G G ++ + Y +A W+G +++++G YT G ++ + +V++G+ LG SP L A
Sbjct: 323 GFFTGFVWCLIFLCYAVAFWYGSTLVLDEGEYTPGTLVQIFLSVIVGALNLGNASPCLEA 382
Query: 348 FAAGQAAAFKMFETINRKPEIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFS 407
FA G+AAA +FETI+RKP ID G KLD I+G+IE +V F YP+RP+ I N +
Sbjct: 383 FATGRAAATSIFETIDRKPIIDCMSEDGYKLDRIKGEIEFHNVTFHYPSRPEVKILNDLN 442
Query: 408 LSLPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVS 467
+ + G ALVG SG+GKST + LI+RFYDP +G V +DG +++ ++W+R +IG+V
Sbjct: 443 MVIKPGEMTALVGPSGAGKSTALQLIQRFYDPCEGMVTVDGHDIRSLNIQWLRDQIGIVE 502
Query: 468 QEPVLFTCSIKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGG 527
QEPVLF+ +I ENI YG++ AT E+I AA+ ANA FI LPQ DT+VGE G Q+SGG
Sbjct: 503 QEPVLFSTTIAENIRYGREDATMEDIVQAAKEANAYNFIMDLPQQFDTLVGEGGGQMSGG 562
Query: 528 QKQRVAIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRN 587
QKQRVAIARA++++P+ILLLD ATSALD ESE +VQE L++I T I VAHRLST+R
Sbjct: 563 QKQRVAIARALIRNPKILLLDMATSALDNESEAMVQEVLSKIQHGHTIISVAHRLSTVRA 622
Query: 588 VDTIAVIHQGKIVERGSHAELTNDPNGAYSQLIRLQEMKRSEQN--DANDKNKPNSIVHS 645
DTI G VERG+H EL + G Y L+ LQ N D D + + + +
Sbjct: 623 ADTIIGFEHGTAVERGTHEELL-ERKGVYFTLVTLQSQGNQALNEEDIKDATEDDMLART 681
Query: 646 GRQSSQRSFSLRSISQGSAGNSGRHSFSASYVAPTTDGFLETEDGGPQASPSKNSSPPEV 705
+ S + SI Q S + +A ED + P + P
Sbjct: 682 FSRGSYQDSLRASIRQRSKSQLS-YLVHEPPLAVVDHKSTYEEDRKDKDIPVQEEVEP-A 739
Query: 706 PLYRLAYFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYKP-ADELRHDSKVWA 764
P+ R+ F+ PE P +L+G++ A ++G + P+ L S+++ TF P +E R
Sbjct: 740 PVRRILKFSAPEWPYMLVGSVGAAVNGTVTPLYAFLFSQILGTFSIPDKEEQRSQINGVC 799
Query: 765 IVFVAVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGA 824
++FVA+ SL + Y F +G L +R+RK F ++ +++WFDD+ +S GAL
Sbjct: 800 LLFVAMGCVSLFTQFLQGYAFAKSGELLTKRLRKFGFRAMLGQDIAWFDDLRNSPGALTT 859
Query: 825 RLSTDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQV 884
RL+TDA+ V+ G +G++V + + V M+IAF SW+L+ ++L P L L+G Q
Sbjct: 860 RLATDASQVQGAAGSQIGMIVNSFTNVTVAMIIAFSFSWKLSLVILCFFPFLALSGATQT 919
Query: 885 KVLKGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGI 944
++L GF++ K+ E Q+ N+A+ +IRTV+ E + +E + + E P K +++
Sbjct: 920 RMLTGFASRDKQALEMVGQITNEALSNIRTVAGIGKERRFIEALETELEKPFKTAIQKAN 979
Query: 945 ISGLGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVP 1004
I G F + +++ ++ + G L+ + FS VF V A+ ++A + ++ + P
Sbjct: 980 IYGFCFAFAQCIMFIANSASYRYGGYLISNEGLHFSYVFRVISAVVLSATALGRAFSYTP 1039
Query: 1005 DSTNAKSAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDL 1064
AK +AA F +LD++ I + +G + +G I+F F YP+R D Q+ N L
Sbjct: 1040 SYAKAKISAARFFQLLDRQPPISVYNTAGEKWDNFQGKIDFVDCKFTYPSRPDSQVLNGL 1099
Query: 1065 CLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLV 1124
++I G+T+A VG SG GKST I LL+RFYDPD G + +DG + +++ V++LR +G+V
Sbjct: 1100 SVSISPGQTLAFVGSSGCGKSTSIQLLERFYDPDQGKVMIDGHDSKKVNVQFLRSNIGIV 1159
Query: 1125 SQEPILFNDTVRANIAYGKGGDATEAE-IVAAAELANAHQFIGSLQKGYDTIVGERGIQL 1183
SQEP+LF ++ NI YG E ++AAA+ A H F+ SL + Y+T VG +G QL
Sbjct: 1160 SQEPVLFACSIMDNIKYGDNTKEIPMERVIAAAKQAQLHDFVMSLPEKYETNVGSQGSQL 1219
Query: 1184 SGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLST 1243
S G+KQR+AIARAIV++PKILLLDEATSALD ESEK VQ ALD+ RT I++AHRLST
Sbjct: 1220 SRGEKQRIAIARAIVRDPKILLLDEATSALDTESEKTVQVALDKAREGRTCIVIAHRLST 1279
Query: 1244 IKGADLIAVVKNGVIAEKGKHEALLHKGGDYASLV 1278
I+ AD+IAV+ GV+ EKG HE L+ + G Y LV
Sbjct: 1280 IQNADIIAVMAQGVVIEKGTHEELMAQKGAYYKLV 1314
Score = 398 bits (1022), Expect = e-110
Identities = 236/669 (35%), Positives = 374/669 (55%), Gaps = 53/669 (7%)
Query: 653 SFSLRSISQGSAGNSGRHSFSASYVAPTTDGFLETEDGGPQASPSKNSSPPEVPLYRLAY 712
S LRSI + N G S SY D + K V ++L
Sbjct: 4 SVILRSIKKFGEENDGFES-DKSY----------NNDKKSRLQDEKKGDGVRVGFFQLFR 52
Query: 713 FNKP-EIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYKPADELR----------HDSK 761
F+ +I ++ +G++ A LHG P + L+ M F EL+ +++
Sbjct: 53 FSSSTDIWLMFVGSLCAFLHGIAQPGVLLIFGTMTDVFIDYDVELQELQIPGKACVNNTI 112
Query: 762 VW---------------------------AIVFVAVAVASLLIIPCRFYFFGVAGGKLIQ 794
VW A + +AVA L+ + F+ +A + IQ
Sbjct: 113 VWTNSSLNQNMTNGTRCGLLNIESEMIKFASYYAGIAVAVLITGYIQICFWVIAAARQIQ 172
Query: 795 RIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQNIATIIVG 854
++RK F +++ ME+ WFD +S G L R S D + + D + L +Q + + I G
Sbjct: 173 KMRKFYFRRIMRMEIGWFDC--NSVGELNTRFSDDINKINDAIADQMALFIQRMTSTICG 230
Query: 855 MVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRT 914
++ F W+L ++++++PL+G+ + F+ K Y +A VA++ + S+RT
Sbjct: 231 FLLGFFRGWKLTLVIISVSPLIGIGAATIGLSVSKFTDYELKAYAKAGVVADEVISSMRT 290
Query: 915 VSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDACVFYAGARLV-E 973
V++F E++ +E Y++ + G+R+GI+ G G + +++ A F+ G+ LV +
Sbjct: 291 VAAFGGEKREVERYEKNLVFAQRWGIRKGIVMGFFTGFVWCLIFLCYAVAFWYGSTLVLD 350
Query: 974 DGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDSSDESG 1033
+G+ T + +F ++ + A+ + + + ++AA SIF +D+K ID E G
Sbjct: 351 EGEYTPGTLVQIFLSVIVGALNLGNASPCLEAFATGRAAATSIFETIDRKPIIDCMSEDG 410
Query: 1034 MTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQR 1093
L+ +KG+IEF++V+F YP+R +V+I NDL + I+ G+ ALVG SG+GKST + L+QR
Sbjct: 411 YKLDRIKGEIEFHNVTFHYPSRPEVKILNDLNMVIKPGEMTALVGPSGAGKSTALQLIQR 470
Query: 1094 FYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIV 1153
FYDP G +T+DG +I+ + ++WLR Q+G+V QEP+LF+ T+ NI YG+ DAT +IV
Sbjct: 471 FYDPCEGMVTVDGHDIRSLNIQWLRDQIGIVEQEPVLFSTTIAENIRYGRE-DATMEDIV 529
Query: 1154 AAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSAL 1213
AA+ ANA+ FI L + +DT+VGE G Q+SGGQKQRVAIARA+++NPKILLLD ATSAL
Sbjct: 530 QAAKEANAYNFIMDLPQQFDTLVGEGGGQMSGGQKQRVAIARALIRNPKILLLDMATSAL 589
Query: 1214 DAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLHKGGD 1273
D ESE +VQ+ L ++ T I VAHRLST++ AD I ++G E+G HE LL + G
Sbjct: 590 DNESEAMVQEVLSKIQHGHTIISVAHRLSTVRAADTIIGFEHGTAVERGTHEELLERKGV 649
Query: 1274 YASLVALHT 1282
Y +LV L +
Sbjct: 650 YFTLVTLQS 658
>AB11_RAT (O70127) Bile salt export pump (ATP-binding cassette,
sub-family B, member 11) (Sister of P-glycoprotein)
Length = 1321
Score = 855 bits (2208), Expect = 0.0
Identities = 494/1303 (37%), Positives = 774/1303 (58%), Gaps = 58/1303 (4%)
Query: 20 HDSNQDS---EKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILI 76
H++++ S +K K+ D+ V ++LF F+ D LMLMG + A+ +G++ P +++I
Sbjct: 26 HNNDKKSRLQDKMKEGDIR---VGFFELFRFSSSKDIWLMLMGGVCALLHGMAQPGILII 82
Query: 77 FGTMINAF-------------GDSTNSKVVDEVSETT-------TYCDQVSL-----KFV 111
FG M + F G + + + ++ + T C V + KF
Sbjct: 83 FGIMTDIFIKYDIERQELEIPGKACVNNTIVWINSSFHQNMTNGTVCGLVDIESEMIKFS 142
Query: 112 YLAAGT----FVASFLQLTCWMITGERQSARIRGLYLKTILRQDVSFFDKETNTGEVVGR 167
+ AG + + Q+ W+ITG RQ R+R +Y + I+R ++ +FD T+ GE+ R
Sbjct: 143 GIYAGVGMTVLILGYFQIRLWVITGARQIRRMRKIYFRRIMRMEIGWFDC-TSVGELNSR 201
Query: 168 MSGDTVLIKDAMGEKVGQFIQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSM 227
+ D I DA+ +++ F+Q MST + G ++ F +GW LT+V+L+ PL+ + ++ +
Sbjct: 202 FADDIEKINDAIADQLAHFLQRMSTAMCGLLLGFYRGWKLTLVILAVSPLIGIGAAVIGL 261
Query: 228 VIAKASSTGQAAYSKSAGVVEQTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALA 287
IAK + AY+K+ + ++ + SIRTVA+F GE + Y ++L+ + + + +
Sbjct: 262 SIAKFTELELKAYAKAGSIADEVLSSIRTVAAFGGENKEVERYEKNLVFAQRWGIWKGMV 321
Query: 288 SGVGFGTLFFVFICSYGLAVWFGGKMII-EKGYTGGDVMTVIFAVLIGSTCLGQTSPSLS 346
G G ++ + Y LA W+G +++ E+ YT G ++ + V++ + +G S L
Sbjct: 322 MGFFTGYMWCLIFFCYALAFWYGSTLVLDEEEYTPGTLVQIFLCVILAAMNIGHASSCLE 381
Query: 347 AFAAGQAAAFKMFETINRKPEIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGF 406
F+ G +AA +F+TI+R+P ID G KLD I+G+IE +V F YP+RPD I +
Sbjct: 382 IFSTGCSAATNIFQTIDRQPVIDCMSGDGYKLDRIKGEIEFHNVTFHYPSRPDVKILDNL 441
Query: 407 SLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLV 466
S+ + G T ALVG SG+GKST + LI+RFYDP +G V +DG +++ ++W+R +IG+V
Sbjct: 442 SMVIKPGETTALVGSSGAGKSTALQLIQRFYDPCEGMVTLDGHDIRSLNIRWLRDQIGIV 501
Query: 467 SQEPVLFTCSIKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSG 526
QEPVLF+ +I ENI +G++ AT E+I AA+ ANA FI LPQ DT+VGE G Q+SG
Sbjct: 502 EQEPVLFSTTIAENIRFGREDATMEDIVQAAKDANAYNFIMALPQQFDTLVGEGGGQMSG 561
Query: 527 GQKQRVAIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIR 586
GQKQRVAIARA++++P+ILLLD ATSALD ESE VQEALN+I T I VAHRLST+R
Sbjct: 562 GQKQRVAIARALIRNPKILLLDMATSALDNESEARVQEALNKIQHGHTIISVAHRLSTVR 621
Query: 587 NVDTIAVIHQGKIVERGSHAELTNDPNGAYSQLIRLQEMKRSEQNDANDKNKPNSIVHSG 646
D I G VERG+H EL + G Y L+ L Q+ ++ +K SI+ G
Sbjct: 622 AADVIIGFEHGVAVERGTHEELL-ERKGVYFMLVTL-------QSQGDNAHKETSIM--G 671
Query: 647 RQSSQRSFSLRSISQGSAGNSGRHSF---SASYVAPTT-DGFLETEDGGPQASPSKNS-- 700
+ +++ R+ S+GS +S R S S S ++ T D L D SK++
Sbjct: 672 KDATEGGTLERTFSRGSYRDSLRASIRQRSKSQLSLLTHDPPLAVADHKSSYKDSKDNDV 731
Query: 701 ---SPPEVPLYRLAYFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTF-YKPADEL 756
P+ R+ +N PE +L+G+++A ++GA+ P+ LL S+++ TF ++
Sbjct: 732 LVEEVEPAPVRRILKYNIPEWHYILVGSLSAAINGAVTPIYSLLFSQLLGTFSLLDKEQQ 791
Query: 757 RHDSKVWAIVFVAVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVE 816
R + + FV + S+ + Y F +G L +R+RK F+ ++ ++ WFDD+
Sbjct: 792 RSEIHSMCLFFVILGCVSIFTQFLQGYTFAKSGELLTKRLRKFGFKAMLGQDIGWFDDLR 851
Query: 817 HSSGALGARLSTDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLL 876
++ G L RL+TDA+ V+ G +G++V + II ++IAF SW+L+ I+ P L
Sbjct: 852 NNPGVLTTRLATDASQVQGATGSQVGMMVNSFTNIIAALLIAFFFSWKLSLIITIFFPFL 911
Query: 877 GLNGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPI 936
L+G VQ K+L GF++ K+ E+A Q+ ++A+ +IRTV+ E + ++ ++ + +
Sbjct: 912 ALSGAVQTKMLTGFASQDKQALEKAGQITSEALSNIRTVAGIGVEGRFIKAFEVELQTSY 971
Query: 937 KKGVRRGIISGLGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGV 996
K VR+ I GL F S + + ++ + G L+ FS VF V +++++A V
Sbjct: 972 KTAVRKANIYGLCFAFSQGIAFLANSAAYRYGGYLIAYEGLGFSHVFRVVSSVALSATAV 1031
Query: 997 SQSGTLVPDSTNAKSAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRL 1056
++ + P AK +AA F +LD+K I+ E+G + +G I+F F YP+R
Sbjct: 1032 GRTFSYTPSYAKAKISAARFFQLLDRKPPINVYSEAGEKWDNFQGKIDFIDCKFTYPSRP 1091
Query: 1057 DVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKW 1116
D+Q+ N L +++ G+T+A VG SG GKST I LL+RFYDPD G + +DG + +++ +++
Sbjct: 1092 DIQVLNGLSVSVNPGQTLAFVGSSGCGKSTSIQLLERFYDPDQGTVMIDGHDSKKVNIQF 1151
Query: 1117 LRQQMGLVSQEPILFNDTVRANIAYGKGGDATEAE-IVAAAELANAHQFIGSLQKGYDTI 1175
LR +G+VSQEP+LF+ ++ NI YG E +AAA+ A H F+ SL + Y+T
Sbjct: 1152 LRSNIGIVSQEPVLFDCSIMDNIKYGDNTKEISVERAIAAAKQAQLHDFVMSLPEKYETN 1211
Query: 1176 VGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTI 1235
VG +G QLS G+KQR+AIARAIV++PKILLLDEATSALD ESEK VQ ALD+ RT I
Sbjct: 1212 VGIQGSQLSRGEKQRIAIARAIVRDPKILLLDEATSALDTESEKTVQTALDKAREGRTCI 1271
Query: 1236 IVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLHKGGDYASLV 1278
++AHRLSTI+ +D+IAVV GV+ EKG HE L+ + G Y LV
Sbjct: 1272 VIAHRLSTIQNSDIIAVVSQGVVIEKGTHEKLMAQKGAYYKLV 1314
Score = 394 bits (1012), Expect = e-109
Identities = 222/609 (36%), Positives = 358/609 (58%), Gaps = 41/609 (6%)
Query: 712 YFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYK----------PADELRHDSK 761
+ + +I ++LMG + A+LHG P I ++ M F K P +++
Sbjct: 53 FSSSKDIWLMLMGGVCALLHGMAQPGILIIFGIMTDIFIKYDIERQELEIPGKACVNNTI 112
Query: 762 VW---------------------------AIVFVAVAVASLLIIPCRFYFFGVAGGKLIQ 794
VW + ++ V + L++ + + + G + I+
Sbjct: 113 VWINSSFHQNMTNGTVCGLVDIESEMIKFSGIYAGVGMTVLILGYFQIRLWVITGARQIR 172
Query: 795 RIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQNIATIIVG 854
R+RK+ F +++ ME+ WFD S G L +R + D + + D L +Q ++T + G
Sbjct: 173 RMRKIYFRRIMRMEIGWFDCT--SVGELNSRFADDIEKINDAIADQLAHFLQRMSTAMCG 230
Query: 855 MVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRT 914
+++ F W+L ++LA++PL+G+ V + F+ K Y +A +A++ + SIRT
Sbjct: 231 LLLGFYRGWKLTLVILAVSPLIGIGAAVIGLSIAKFTELELKAYAKAGSIADEVLSSIRT 290
Query: 915 VSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDACVFYAGARLVED 974
V++F E K +E Y++ + G+ +G++ G G + +++ A F+ G+ LV D
Sbjct: 291 VAAFGGENKEVERYEKNLVFAQRWGIWKGMVMGFFTGYMWCLIFFCYALAFWYGSTLVLD 350
Query: 975 GKS-TFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDSSDESG 1033
+ T + +F + +AAM + + + + + SAA +IF +D++ ID G
Sbjct: 351 EEEYTPGTLVQIFLCVILAAMNIGHASSCLEIFSTGCSAATNIFQTIDRQPVIDCMSGDG 410
Query: 1034 MTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQR 1093
L+ +KG+IEF++V+F YP+R DV+I ++L + I+ G+T ALVG SG+GKST + L+QR
Sbjct: 411 YKLDRIKGEIEFHNVTFHYPSRPDVKILDNLSMVIKPGETTALVGSSGAGKSTALQLIQR 470
Query: 1094 FYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIV 1153
FYDP G +TLDG +I+ + ++WLR Q+G+V QEP+LF+ T+ NI +G+ DAT +IV
Sbjct: 471 FYDPCEGMVTLDGHDIRSLNIRWLRDQIGIVEQEPVLFSTTIAENIRFGRE-DATMEDIV 529
Query: 1154 AAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSAL 1213
AA+ ANA+ FI +L + +DT+VGE G Q+SGGQKQRVAIARA+++NPKILLLD ATSAL
Sbjct: 530 QAAKDANAYNFIMALPQQFDTLVGEGGGQMSGGQKQRVAIARALIRNPKILLLDMATSAL 589
Query: 1214 DAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLHKGGD 1273
D ESE VQ+AL+++ T I VAHRLST++ AD+I ++GV E+G HE LL + G
Sbjct: 590 DNESEARVQEALNKIQHGHTIISVAHRLSTVRAADVIIGFEHGVAVERGTHEELLERKGV 649
Query: 1274 YASLVALHT 1282
Y LV L +
Sbjct: 650 YFMLVTLQS 658
Score = 377 bits (968), Expect = e-103
Identities = 228/611 (37%), Positives = 350/611 (56%), Gaps = 19/611 (3%)
Query: 17 VEDHDSNQDSEKSKDKDVTTKTV---PLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLM 73
V DH S+ + SKD DV + V P+ ++ + P + L+G+L A NG P+
Sbjct: 716 VADHKSSY--KDSKDNDVLVEEVEPAPVRRILKYNIPEWHYI-LVGSLSAAINGAVTPIY 772
Query: 74 ILIFGTMINAFGDSTNSKVVDEVSETTTYCDQVSLKFVYLAAGTFVASFLQLTCWMITGE 133
L+F ++ F + + SE + C L FV L + FLQ + +GE
Sbjct: 773 SLLFSQLLGTFSLLDKEQ---QRSEIHSMC----LFFVILGCVSIFTQFLQGYTFAKSGE 825
Query: 134 RQSARIRGLYLKTILRQDVSFFDK-ETNTGEVVGRMSGDTVLIKDAMGEKVGQFIQFMST 192
+ R+R K +L QD+ +FD N G + R++ D ++ A G +VG + +
Sbjct: 826 LLTKRLRKFGFKAMLGQDIGWFDDLRNNPGVLTTRLATDASQVQGATGSQVGMMVNSFTN 885
Query: 193 FIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKSAGVVEQTIG 252
I +IAF W L++++ P L LSG++ + ++ +S + A K+ + + +
Sbjct: 886 IIAALLIAFFFSWKLSLIITIFFPFLALSGAVQTKMLTGFASQDKQALEKAGQITSEALS 945
Query: 253 SIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGVGFGTLFFVFICSYGLAVWFGGK 312
+IRTVA E + + L YKTAV++A G+ F + + A +GG
Sbjct: 946 NIRTVAGIGVEGRFIKAFEVELQTSYKTAVRKANIYGLCFAFSQGIAFLANSAAYRYGGY 1005
Query: 313 MIIEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETINRKPEIDAYD 372
+I +G V V+ +V + +T +G+T ++A + +A + F+ ++RKP I+ Y
Sbjct: 1006 LIAYEGLGFSHVFRVVSSVALSATAVGRTFSYTPSYAKAKISAARFFQLLDRKPPINVYS 1065
Query: 373 TSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSL 432
+G+K D+ +G I+ D F+YP+RPD + NG S+S+ G T A VG SG GKST + L
Sbjct: 1066 EAGEKWDNFQGKIDFIDCKFTYPSRPDIQVLNGLSVSVNPGQTLAFVGSSGCGKSTSIQL 1125
Query: 433 IERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYG---KDCAT 489
+ERFYDP G V+IDG + K+ ++++R IG+VSQEPVLF CSI +NI YG K+ +
Sbjct: 1126 LERFYDPDQGTVMIDGHDSKKVNIQFLRSNIGIVSQEPVLFDCSIMDNIKYGDNTKEISV 1185
Query: 490 DEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLDE 549
+ I AA+ A F+ LP+ +T VG G+QLS G+KQR+AIARAI++DP+ILLLDE
Sbjct: 1186 ERAI-AAAKQAQLHDFVMSLPEKYETNVGIQGSQLSRGEKQRIAIARAIVRDPKILLLDE 1244
Query: 550 ATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAELT 609
ATSALD ESE+ VQ AL++ RT IV+AHRLSTI+N D IAV+ QG ++E+G+H +L
Sbjct: 1245 ATSALDTESEKTVQTALDKAREGRTCIVIAHRLSTIQNSDIIAVVSQGVVIEKGTHEKLM 1304
Query: 610 NDPNGAYSQLI 620
GAY +L+
Sbjct: 1305 AQ-KGAYYKLV 1314
>AB11_MOUSE (Q9QY30) Bile salt export pump (ATP-binding cassette,
sub-family B, member 11) (Sister of P-glycoprotein)
Length = 1321
Score = 847 bits (2188), Expect = 0.0
Identities = 475/1292 (36%), Positives = 753/1292 (57%), Gaps = 36/1292 (2%)
Query: 20 HDSNQDSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILIFGT 79
H++++ S K V ++LF F+ D LM MG++ A+ +G++ P MI++FG
Sbjct: 26 HNNDKKSRLQDKKKGEGARVGFFELFRFSSSKDNWLMFMGSVCALLHGMAQPGMIIVFGI 85
Query: 80 MINAF-------------------------GDSTNSKVVDEVSETTTYCDQVSLKFVYLA 114
+ + F S N + + S + +KF +
Sbjct: 86 LTDIFVEYDIERQELSIPGKVCMNNTIVWINSSFNQNMTNGTSCGLVDINSEVIKFSGIY 145
Query: 115 AGTFVA----SFLQLTCWMITGERQSARIRGLYLKTILRQDVSFFDKETNTGEVVGRMSG 170
AG VA + Q+ W+ITG RQ ++R Y + I+R ++ +FD T+ GE+ R S
Sbjct: 146 AGVGVAVLILGYFQIRLWVITGARQIRKMRKFYFRRIMRMEIGWFDC-TSVGELNSRFSD 204
Query: 171 DTVLIKDAMGEKVGQFIQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIA 230
D I +A+ +++ F+Q +ST + G ++ F +GW LT+V+L+ PL+ + ++ + +A
Sbjct: 205 DINKIDEAIADQMALFLQRLSTALSGLLLGFYRGWKLTLVILAVSPLIGIGAAVIGLSVA 264
Query: 231 KASSTGQAAYSKSAGVVEQTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGV 290
K + AY+K+ + ++ + SIRTVA+F GE + Y ++L+ + + + + G
Sbjct: 265 KFTELELKAYAKAGSIADEVLSSIRTVAAFGGENKEVERYEKNLMFAQRWGIWKGMVMGF 324
Query: 291 GFGTLFFVFICSYGLAVWFGGKMIIEKG-YTGGDVMTVIFAVLIGSTCLGQTSPSLSAFA 349
G ++ + Y LA W+G ++++++G YT G ++ + V+I + +G S L F+
Sbjct: 325 FTGYMWCLIFFCYALAFWYGSRLVLDEGEYTPGTLIQIFLCVIIAAMNIGNASSCLEIFS 384
Query: 350 AGQAAAFKMFETINRKPEIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLS 409
G +AA +F+TI+R+P +D G KLD I+G+IE +V F YP+RP+ I N S+
Sbjct: 385 TGCSAASSIFQTIDRQPVMDCMSGDGYKLDRIKGEIEFHNVTFHYPSRPEVKILNNLSMV 444
Query: 410 LPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQE 469
+ G T A VG SG+GKST + LI+RFYDP +G V +DG +++ ++W+R +IG+V QE
Sbjct: 445 IKPGETTAFVGSSGAGKSTALQLIQRFYDPCEGMVTLDGHDIRSLNIRWLRDQIGIVEQE 504
Query: 470 PVLFTCSIKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQK 529
PVLF+ +I ENI G++ AT E+I AA+ ANA FI LPQ DT+VGE G Q+SGGQK
Sbjct: 505 PVLFSTTIAENIRLGREEATMEDIVQAAKDANAYNFIMALPQQFDTLVGEGGGQMSGGQK 564
Query: 530 QRVAIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVD 589
QRVAIARA+++ P+ILLLD ATSALD ESE VQ ALN+I T I VAHRLST+R+ D
Sbjct: 565 QRVAIARALIRKPKILLLDMATSALDNESEAKVQGALNKIQHGHTIISVAHRLSTVRSAD 624
Query: 590 TIAVIHQGKIVERGSHAELTNDPNGAYSQLIRLQEMKRSEQNDANDKNKPNSIVHSGRQS 649
I G VERG+H EL + G Y L+ LQ + + + K K + + ++
Sbjct: 625 VIIGFEHGTAVERGTHEELL-ERKGVYFMLVTLQSQEDNTHKETGIKGKDTTEGDTPERT 683
Query: 650 SQRSFSLRSISQGSAGNSGRHSFSASYVAPTTDGFLETEDGGPQASPSKNSSPPEVPLYR 709
R S+ S S+ P G ++ + + P+ R
Sbjct: 684 FSRGSYQDSLRASIRQRSKSQLSHLSHEPPLAIGDHKSSYEDRKDNDVLVEEVEPAPVRR 743
Query: 710 LAYFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYKPADELRHDSKVWA--IVF 767
+ +N E P +L+G + A ++GA+ P+ LL S+++ TF D+ + S++++ + F
Sbjct: 744 ILKYNISEWPYILVGALCAAINGAVTPIYSLLFSQILKTF-SLVDKEQQRSEIYSMCLFF 802
Query: 768 VAVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLS 827
V + SL + Y F +G L +R+RK F+ ++ ++ WFDD++++ G L RL+
Sbjct: 803 VILGCVSLFTQFLQGYNFAKSGELLTKRLRKFGFKAMLRQDIGWFDDLKNNPGVLTTRLA 862
Query: 828 TDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVL 887
TDA+ V+ G +G++V + I V ++IAF +W+L+ ++ P L L+G VQ K+L
Sbjct: 863 TDASQVQGATGSQVGMMVNSFTNIFVAVLIAFLFNWKLSLVISVFFPFLALSGAVQTKML 922
Query: 888 KGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISG 947
GF++ K++ E+A Q+ N+A+ +IRTV+ E + ++ ++ + E K +R+ + G
Sbjct: 923 TGFASQDKEILEKAGQITNEALSNIRTVAGIGVEGRFIKAFEVELEKSYKTAIRKANVYG 982
Query: 948 LGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDST 1007
L + S + + ++ + G L+ FS VF V +++M+A V ++ + P
Sbjct: 983 LCYAFSQGISFLANSAAYRYGGYLIVYEDLNFSYVFRVVSSIAMSATAVGRTFSYTPSYA 1042
Query: 1008 NAKSAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLN 1067
AK +AA F +LD+K ID +G + +G I+F F YP+R D+Q+ N L ++
Sbjct: 1043 KAKISAARFFQLLDRKPPIDVYSGAGEKWDNFQGKIDFIDCKFTYPSRPDIQVLNGLSVS 1102
Query: 1068 IRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQE 1127
+ G+T+A VG SG GKST I LL+RFYDPD G + +DG + +++ V++LR +G+VSQE
Sbjct: 1103 VDPGQTLAFVGSSGCGKSTSIQLLERFYDPDQGTVMIDGHDSKKVNVQFLRSNIGIVSQE 1162
Query: 1128 PILFNDTVRANIAYGKGGDATEAE-IVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGG 1186
P+LF+ ++ NI YG E +AAA+ A H F+ SL + Y+T VG +G QLS G
Sbjct: 1163 PVLFDCSIMDNIKYGDNTKEISVERAIAAAKQAQLHDFVMSLPEKYETNVGIQGSQLSRG 1222
Query: 1187 QKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKG 1246
+KQR+AIARAIV++PKILLLDEATSALD ESEK VQ ALD+ RT I++AHRLSTI+
Sbjct: 1223 EKQRIAIARAIVRDPKILLLDEATSALDTESEKTVQLALDKAREGRTCIVIAHRLSTIQN 1282
Query: 1247 ADLIAVVKNGVIAEKGKHEALLHKGGDYASLV 1278
+D+IAV+ GV+ EKG H+ L+ + G Y LV
Sbjct: 1283 SDIIAVMSQGVVIEKGTHKKLMDQKGAYYKLV 1314
Score = 392 bits (1006), Expect = e-108
Identities = 234/674 (34%), Positives = 379/674 (55%), Gaps = 53/674 (7%)
Query: 653 SFSLRSISQGSAGNSGRHSFSASYVAPTTDGFLETEDGGPQASPSKNSSPPEVPLYRLAY 712
S LRS+ + N H+F + DGF D + K V + L
Sbjct: 4 SVILRSVKKFGEEN---HAFES-------DGF-HNNDKKSRLQDKKKGEGARVGFFELFR 52
Query: 713 FNKPEIPVLL-MGTITAVLHGAIMP----VIGLLVSKMISTFYK------PADELRHDSK 761
F+ + L+ MG++ A+LHG P V G+L + + P +++
Sbjct: 53 FSSSKDNWLMFMGSVCALLHGMAQPGMIIVFGILTDIFVEYDIERQELSIPGKVCMNNTI 112
Query: 762 VW---------------------------AIVFVAVAVASLLIIPCRFYFFGVAGGKLIQ 794
VW + ++ V VA L++ + + + G + I+
Sbjct: 113 VWINSSFNQNMTNGTSCGLVDINSEVIKFSGIYAGVGVAVLILGYFQIRLWVITGARQIR 172
Query: 795 RIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQNIATIIVG 854
++RK F +++ ME+ WFD S G L +R S D + + D + L +Q ++T + G
Sbjct: 173 KMRKFYFRRIMRMEIGWFDCT--SVGELNSRFSDDINKIDEAIADQMALFLQRLSTALSG 230
Query: 855 MVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRT 914
+++ F W+L ++LA++PL+G+ V + F+ K Y +A +A++ + SIRT
Sbjct: 231 LLLGFYRGWKLTLVILAVSPLIGIGAAVIGLSVAKFTELELKAYAKAGSIADEVLSSIRT 290
Query: 915 VSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDACVFYAGARLV-E 973
V++F E K +E Y++ + G+ +G++ G G + +++ A F+ G+RLV +
Sbjct: 291 VAAFGGENKEVERYEKNLMFAQRWGIWKGMVMGFFTGYMWCLIFFCYALAFWYGSRLVLD 350
Query: 974 DGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDSSDESG 1033
+G+ T + +F + +AAM + + + + + SAA+SIF +D++ +D G
Sbjct: 351 EGEYTPGTLIQIFLCVIIAAMNIGNASSCLEIFSTGCSAASSIFQTIDRQPVMDCMSGDG 410
Query: 1034 MTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQR 1093
L+ +KG+IEF++V+F YP+R +V+I N+L + I+ G+T A VG SG+GKST + L+QR
Sbjct: 411 YKLDRIKGEIEFHNVTFHYPSRPEVKILNNLSMVIKPGETTAFVGSSGAGKSTALQLIQR 470
Query: 1094 FYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIV 1153
FYDP G +TLDG +I+ + ++WLR Q+G+V QEP+LF+ T+ NI G+ +AT +IV
Sbjct: 471 FYDPCEGMVTLDGHDIRSLNIRWLRDQIGIVEQEPVLFSTTIAENIRLGRE-EATMEDIV 529
Query: 1154 AAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSAL 1213
AA+ ANA+ FI +L + +DT+VGE G Q+SGGQKQRVAIARA+++ PKILLLD ATSAL
Sbjct: 530 QAAKDANAYNFIMALPQQFDTLVGEGGGQMSGGQKQRVAIARALIRKPKILLLDMATSAL 589
Query: 1214 DAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLHKGGD 1273
D ESE VQ AL+++ T I VAHRLST++ AD+I ++G E+G HE LL + G
Sbjct: 590 DNESEAKVQGALNKIQHGHTIISVAHRLSTVRSADVIIGFEHGTAVERGTHEELLERKGV 649
Query: 1274 YASLVALHTSDSTS 1287
Y LV L + + +
Sbjct: 650 YFMLVTLQSQEDNT 663
Score = 374 bits (961), Expect = e-103
Identities = 227/611 (37%), Positives = 352/611 (57%), Gaps = 19/611 (3%)
Query: 17 VEDHDSNQDSEKSKDKDVTTKTV---PLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLM 73
+ DH S+ E KD DV + V P+ ++ + + S+ +L+G L A NG P+
Sbjct: 716 IGDHKSSY--EDRKDNDVLVEEVEPAPVRRILKY-NISEWPYILVGALCAAINGAVTPIY 772
Query: 74 ILIFGTMINAFGDSTNSKVVDEVSETTTYCDQVSLKFVYLAAGTFVASFLQLTCWMITGE 133
L+F ++ F S V E + Y + L FV L + FLQ + +GE
Sbjct: 773 SLLFSQILKTF-----SLVDKEQQRSEIY--SMCLFFVILGCVSLFTQFLQGYNFAKSGE 825
Query: 134 RQSARIRGLYLKTILRQDVSFFDK-ETNTGEVVGRMSGDTVLIKDAMGEKVGQFIQFMST 192
+ R+R K +LRQD+ +FD + N G + R++ D ++ A G +VG + +
Sbjct: 826 LLTKRLRKFGFKAMLRQDIGWFDDLKNNPGVLTTRLATDASQVQGATGSQVGMMVNSFTN 885
Query: 193 FIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKSAGVVEQTIG 252
+IAF W L++V+ P L LSG++ + ++ +S + K+ + + +
Sbjct: 886 IFVAVLIAFLFNWKLSLVISVFFPFLALSGAVQTKMLTGFASQDKEILEKAGQITNEALS 945
Query: 253 SIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGVGFGTLFFVFICSYGLAVWFGGK 312
+IRTVA E + + L K YKTA+++A G+ + + + A +GG
Sbjct: 946 NIRTVAGIGVEGRFIKAFEVELEKSYKTAIRKANVYGLCYAFSQGISFLANSAAYRYGGY 1005
Query: 313 MIIEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETINRKPEIDAYD 372
+I+ + V V+ ++ + +T +G+T ++A + +A + F+ ++RKP ID Y
Sbjct: 1006 LIVYEDLNFSYVFRVVSSIAMSATAVGRTFSYTPSYAKAKISAARFFQLLDRKPPIDVYS 1065
Query: 373 TSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSL 432
+G+K D+ +G I+ D F+YP+RPD + NG S+S+ G T A VG SG GKST + L
Sbjct: 1066 GAGEKWDNFQGKIDFIDCKFTYPSRPDIQVLNGLSVSVDPGQTLAFVGSSGCGKSTSIQL 1125
Query: 433 IERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYG---KDCAT 489
+ERFYDP G V+IDG + K+ ++++R IG+VSQEPVLF CSI +NI YG K+ +
Sbjct: 1126 LERFYDPDQGTVMIDGHDSKKVNVQFLRSNIGIVSQEPVLFDCSIMDNIKYGDNTKEISV 1185
Query: 490 DEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLDE 549
+ I AA+ A F+ LP+ +T VG G+QLS G+KQR+AIARAI++DP+ILLLDE
Sbjct: 1186 ERAI-AAAKQAQLHDFVMSLPEKYETNVGIQGSQLSRGEKQRIAIARAIVRDPKILLLDE 1244
Query: 550 ATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAELT 609
ATSALD ESE+ VQ AL++ RT IV+AHRLSTI+N D IAV+ QG ++E+G+H +L
Sbjct: 1245 ATSALDTESEKTVQLALDKAREGRTCIVIAHRLSTIQNSDIIAVMSQGVVIEKGTHKKLM 1304
Query: 610 NDPNGAYSQLI 620
D GAY +L+
Sbjct: 1305 -DQKGAYYKLV 1314
>AB11_RABIT (Q9N0V3) Bile salt export pump (ATP-binding cassette,
sub-family B, member 11) (Sister of P-glycoprotein)
Length = 1321
Score = 835 bits (2156), Expect = 0.0
Identities = 481/1295 (37%), Positives = 747/1295 (57%), Gaps = 45/1295 (3%)
Query: 22 SNQDSEKSKDKDVTTKT-VPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILIFGTM 80
+N+ + +DK + + ++LF F+ +D LM MG+L A +G++ P ++LIFGTM
Sbjct: 27 NNEKKSRLQDKKKSDSVRIGFFQLFRFSSWTDIWLMCMGSLCACIHGIAQPGVLLIFGTM 86
Query: 81 INAFGD-------------------------STNSKVVDEVSETTTYCDQVSLKFVYLAA 115
+ F D S N V + + ++F A
Sbjct: 87 TDVFIDYDTELQELKIPGKACVNNTIVWINSSLNQNVTNGTRCGLLDIESEMIRFAGYYA 146
Query: 116 GTFVA----SFLQLTCWMITGERQSARIRGLYLKTILRQDVSFFDKETNTGEVVGRMSGD 171
G +A ++Q+ W I Q ++R Y + I+R + + D + G++ S D
Sbjct: 147 GIGIAVLTTGYIQICFWGIAAAHQIQKMRKSYFRKIMRMGIGWVDCNS-VGKLNTPFSVD 205
Query: 172 TVLIKDAMGEKVGQFIQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIAK 231
I D+ +++ FIQ M++ I GF++ F++ W LT+V++S PL+ L ++ + ++K
Sbjct: 206 FNKINDSSADQLAIFIQGMTSPIFGFLVGFSQWWKLTLVIISVSPLIGLGAAIIGLSVSK 265
Query: 232 ASSTGQAAYSKSAGVVEQTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGVG 291
+ AY+K+ V ++ I S+RTVA+F GEK+ Y ++L+ + +++ + G
Sbjct: 266 FTDYELKAYAKAGSVADEVISSMRTVAAFGGEKKEVERYEKNLVFAQRWGIRKGIVMGFF 325
Query: 292 FGTLFFVFICSYGLAVWFGGKMIIEKG-YTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAA 350
G ++ + Y LA W+G K+++E+G Y+ G ++ + +V+IG+ LG SP L AFAA
Sbjct: 326 TGYMWCLIFFCYALAFWYGSKLVLEEGEYSPGALVQIFLSVIIGALNLGNASPCLEAFAA 385
Query: 351 GQAAAFKMFETINRKPEIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSL 410
G+AAA +FETI+RKP ID G KL+ I+G+IE +V F YP+RP+ I N S+ +
Sbjct: 386 GRAAASSIFETIDRKPIIDCMSEDGYKLERIKGEIEFHNVTFHYPSRPEVKILNNLSMVI 445
Query: 411 PSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEP 470
G ALVG SG+GKST + LI RFY PT+G V ++ +++ ++W+R +IG+V QEP
Sbjct: 446 KPGEMTALVGPSGAGKSTALQLIHRFYGPTEGMVTVESHDIRSSHIQWLRNQIGIVEQEP 505
Query: 471 VLFTCSIKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQ 530
VLF +I E I YG++ AT E++ AA+ ANA FI LPQ DT+VGE G Q+SGGQKQ
Sbjct: 506 VLFFHTIAEKIRYGREDATMEDLIQAAKEANAYNFIMDLPQQFDTLVGEGGGQMSGGQKQ 565
Query: 531 RVAIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDT 590
RVAIARA++++P+ILLLD ATSALD ESE +VQEAL++ T + VAHR +TIR D
Sbjct: 566 RVAIARALIRNPKILLLDMATSALDNESEAMVQEALSKTQHGHTIVSVAHRPATIRTADV 625
Query: 591 IAVIHQGKIVERGSHAELTNDPNGAYSQLIRLQEMKRSEQNDANDKNK-----PNSIVHS 645
I G VERG+ EL + G Y L+ LQ + + N+K+ P
Sbjct: 626 IIGCEHGAAVERGTEEELL-ERKGVYFALVTLQSQRNQGDQEENEKDATEDDIPEKTFSR 684
Query: 646 GRQSSQRSFSLRSISQGSAGNSGRHSFSASYVAPTTDGFLETEDGGPQASPSKNSSPPEV 705
G SLR S+ A +T ED + P++ P
Sbjct: 685 GNYQDSLRASLRQRSKSQLSYLAHEPPMAVEDHKST----HEEDRKDKDLPAQEDIEP-A 739
Query: 706 PLYRLAYFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYKP-ADELRHDSKVWA 764
+ R+ N PE P +L+G++ A ++GA+ P+ L S+++ TF P +E R
Sbjct: 740 SVRRIMKLNAPEWPYMLLGSMGAAVNGAVTPLYAFLFSQILGTFSLPDKEEQRSQINGIC 799
Query: 765 IVFVAVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGA 824
++FV + S + Y F +G L +R+RK F ++ ++ WFDD+ +S GAL
Sbjct: 800 LLFVTLGCVSFFTQFLQGYTFAKSGELLTKRLRKFGFRAMLGQDIGWFDDLRNSPGALTT 859
Query: 825 RLSTDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQV 884
RL+TDA+ V+ G +G++V + + V M+IAF SW+L ++ P L L+G +Q
Sbjct: 860 RLATDASQVQGATGSQIGMMVNSFTNVTVAMIIAFLFSWKLTLGIVCFFPFLALSGALQT 919
Query: 885 KVLKGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGI 944
K+L GF++ K+ E+A Q+ ++A+ +IRTV+ E K +E ++ + E P K +++
Sbjct: 920 KMLTGFASRDKQALEKAGQITSEALSNIRTVAGIGKERKFIETFEAELEKPYKMAIKKAN 979
Query: 945 ISGLGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVP 1004
+ GL FG S + + ++ + G L+ + FS VF V A+ ++A + ++ + P
Sbjct: 980 VYGLCFGFSQCITFIANSASYRYGGYLISNEGLHFSYVFRVISAVVLSATALGRASSYTP 1039
Query: 1005 DSTNAKSAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDL 1064
AK +AA F +LD++ I+ +G + +G I+F F YP+R D+Q+ N L
Sbjct: 1040 SYAKAKISAARFFQLLDRQPPINVYSSAGEKWDNFQGKIDFVDCKFTYPSRPDIQVLNGL 1099
Query: 1065 CLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLV 1124
+++ +T+A VG SG GKST I LL+RFYDPD G + +DG + +++ +++LR +G+V
Sbjct: 1100 SVSMSPRQTLAFVGSSGCGKSTSIQLLERFYDPDHGKVMIDGHDSRKVNIQFLRSNIGIV 1159
Query: 1125 SQEPILFNDTVRANIAYGKGGDATEAE-IVAAAELANAHQFIGSLQKGYDTIVGERGIQL 1183
SQEP+LF +++ NI YG E I+AAA+ A H F+ SL + Y+T VG +G QL
Sbjct: 1160 SQEPVLFACSIKDNIKYGDNTQEIPMERIIAAAKKAQVHDFVMSLPEKYETNVGSQGSQL 1219
Query: 1184 SGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLST 1243
S G+KQR+AIARAIV++PKILLLDEATSALD ESEK VQ ALD+ RT I++AHRLST
Sbjct: 1220 SRGEKQRIAIARAIVRDPKILLLDEATSALDTESEKTVQVALDKAREGRTCIVIAHRLST 1279
Query: 1244 IKGADLIAVVKNGVIAEKGKHEALLHKGGDYASLV 1278
I+ +D+IAV+ G++ EKG HE L+ + G Y LV
Sbjct: 1280 IQNSDIIAVMSQGMVIEKGTHEELMVQKGAYYKLV 1314
Score = 371 bits (953), Expect = e-102
Identities = 225/669 (33%), Positives = 365/669 (53%), Gaps = 53/669 (7%)
Query: 653 SFSLRSISQGSAGNSGRHSFSASYVAPTTDGFLETEDGGPQASPSKNSSPPEVPLYRLAY 712
S LRS+ + N G S DG E + K S + ++L
Sbjct: 4 SVILRSVKKFGEENHGFES----------DGSYNNEKKS-RLQDKKKSDSVRIGFFQLFR 52
Query: 713 FNK-PEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYKPADELR----------HDSK 761
F+ +I ++ MG++ A +HG P + L+ M F EL+ +++
Sbjct: 53 FSSWTDIWLMCMGSLCACIHGIAQPGVLLIFGTMTDVFIDYDTELQELKIPGKACVNNTI 112
Query: 762 VW---------------------------AIVFVAVAVASLLIIPCRFYFFGVAGGKLIQ 794
VW A + + +A L + F+G+A IQ
Sbjct: 113 VWINSSLNQNVTNGTRCGLLDIESEMIRFAGYYAGIGIAVLTTGYIQICFWGIAAAHQIQ 172
Query: 795 RIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQNIATIIVG 854
++RK F K++ M + W D +S G L S D + D L + +Q + + I G
Sbjct: 173 KMRKSYFRKIMRMGIGWVDC--NSVGKLNTPFSVDFNKINDSSADQLAIFIQGMTSPIFG 230
Query: 855 MVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRT 914
++ F W+L ++++++PL+GL + + F+ K Y +A VA++ + S+RT
Sbjct: 231 FLVGFSQWWKLTLVIISVSPLIGLGAAIIGLSVSKFTDYELKAYAKAGSVADEVISSMRT 290
Query: 915 VSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDACVFYAGARLV-E 973
V++F E+K +E Y++ + G+R+GI+ G G + +++ A F+ G++LV E
Sbjct: 291 VAAFGGEKKEVERYEKNLVFAQRWGIRKGIVMGFFTGYMWCLIFFCYALAFWYGSKLVLE 350
Query: 974 DGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDSSDESG 1033
+G+ + + +F ++ + A+ + + + ++AA+SIF +D+K ID E G
Sbjct: 351 EGEYSPGALVQIFLSVIIGALNLGNASPCLEAFAAGRAAASSIFETIDRKPIIDCMSEDG 410
Query: 1034 MTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQR 1093
LE +KG+IEF++V+F YP+R +V+I N+L + I+ G+ ALVG SG+GKST + L+ R
Sbjct: 411 YKLERIKGEIEFHNVTFHYPSRPEVKILNNLSMVIKPGEMTALVGPSGAGKSTALQLIHR 470
Query: 1094 FYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIV 1153
FY P G +T++ +I+ ++WLR Q+G+V QEP+LF T+ I YG+ DAT +++
Sbjct: 471 FYGPTEGMVTVESHDIRSSHIQWLRNQIGIVEQEPVLFFHTIAEKIRYGRE-DATMEDLI 529
Query: 1154 AAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSAL 1213
AA+ ANA+ FI L + +DT+VGE G Q+SGGQKQRVAIARA+++NPKILLLD ATSAL
Sbjct: 530 QAAKEANAYNFIMDLPQQFDTLVGEGGGQMSGGQKQRVAIARALIRNPKILLLDMATSAL 589
Query: 1214 DAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLHKGGD 1273
D ESE +VQ+AL + T + VAHR +TI+ AD+I ++G E+G E LL + G
Sbjct: 590 DNESEAMVQEALSKTQHGHTIVSVAHRPATIRTADVIIGCEHGAAVERGTEEELLERKGV 649
Query: 1274 YASLVALHT 1282
Y +LV L +
Sbjct: 650 YFALVTLQS 658
>MDR1_CAEEL (P34712) Multidrug resistance protein 1 (P-glycoprotein A)
Length = 1321
Score = 814 bits (2102), Expect = 0.0
Identities = 475/1296 (36%), Positives = 749/1296 (57%), Gaps = 42/1296 (3%)
Query: 13 SVQPVEDH-----DSNQDSEKSKD-KDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGN 66
+++ VED+ DSN + + ++D K+ V + +L+ + ++LL+ +GTL A+
Sbjct: 28 AIKTVEDYEGDNIDSNGEIKITRDAKEEVVNKVSIPQLYRYTTTLEKLLLFIGTLVAVIT 87
Query: 67 GLSIPLMILIFGTMINAF---------GDSTNSKVVDEVSETTTYCDQVSLKFVYLA--A 115
G +PLM ++ G + AF ST ++T D +++ + Y A
Sbjct: 88 GAGLPLMSILQGKVSQAFINEQIVINNNGSTFLPTGQNYTKTDFEHDVMNVVWSYAAMTV 147
Query: 116 GTFVASFLQLTCWMITGERQSARIRGLYLKTILRQDVSFFDKETNTGEVVGRMSGDTVLI 175
G + A + +TC++ E+ + R+R ++K+ILRQ++S+FD ++G + ++ + +
Sbjct: 148 GMWAAGQITVTCYLYVAEQMNNRLRREFVKSILRQEISWFDTN-HSGTLATKLFDNLERV 206
Query: 176 KDAMGEKVGQFIQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIAKASST 235
K+ G+K+G Q++S FI GF++AFT W LT+VML+ P+ L G IAK+ ST
Sbjct: 207 KEGTGDKIGMAFQYLSQFITGFIVAFTHSWQLTLVMLAVTPIQALCG----FAIAKSMST 262
Query: 236 ----GQAAYSKSAGVVEQTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGVG 291
Y+K+ VVE+TI SIRTV S G + Y+ ++ + K V + L G+
Sbjct: 263 FAIRETLRYAKAGKVVEETISSIRTVVSLNGLRYELERYSTAVEEAKKAGVLKGLFLGIS 322
Query: 292 FGTLFFVFICSYGLAVWFGGKMIIEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAG 351
FG + S+ LA + G + + GD++T +V++GS LG P L+
Sbjct: 323 FGAMQASNFISFALAFYIGVGWVHDGSLNFGDMLTTFSSVMMGSMALGLAGPQLAVLGTA 382
Query: 352 QAAAFKMFETINRKPEIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLP 411
Q AA ++E ++RKP ID+ +G+K I+GDI + +V F+YP+RPD I G +L +
Sbjct: 383 QGAASGIYEVLDRKPVIDSSSKAGRKDMKIKGDITVENVHFTYPSRPDVPILRGMNLRVN 442
Query: 412 SGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPV 471
+G T ALVG SG GKST++SL+ R+YD G++ IDG+++++ L+++R+ + +VSQEP
Sbjct: 443 AGQTVALVGSSGCGKSTIISLLLRYYDVLKGKITIDGVDVRDINLEFLRKNVAVVSQEPA 502
Query: 472 LFTCSIKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQR 531
LF C+I+ENI+ GK+ T EE+ A ++ANA KFI LP G +T+VG+ GTQLSGGQKQR
Sbjct: 503 LFNCTIEENISLGKEGITREEMVAACKMANAEKFIKTLPNGYNTLVGDRGTQLSGGQKQR 562
Query: 532 VAIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTI 591
+AIARA++++P+ILLLDEATSALDAESE IVQ+AL++ RTTI++AHRLSTIRN D I
Sbjct: 563 IAIARALVRNPKILLLDEATSALDAESEGIVQQALDKAAKGRTTIIIAHRLSTIRNADLI 622
Query: 592 AVIHQGKIVERGSHAELTNDPNGAYSQLIRLQEMKRSEQNDANDK-NKPNSIVHSGRQSS 650
G++VE G H L G Y L+ Q + + A K ++ NS+ RQ+S
Sbjct: 623 ISCKNGQVVEVGDHRALMAQ-QGLYYDLVTAQTFTDAVDSAAEGKFSRENSV---ARQTS 678
Query: 651 QRSFSLRSISQ-GSAGNSGRHSFSASYVAPTTDGFLETEDGGPQASPSK----NSSPPEV 705
+ R S+ N R S S E G S K ++ +
Sbjct: 679 EHEGLSRQASEMDDIMNRVRSSTIGSITNGPVIDEKEERIGKDALSRLKQELEENNAQKT 738
Query: 706 PLYRLAYFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFY-KPADELRHDSKVWA 764
L+ + Y +P L +G TA + G I P + + ++ F PAD L WA
Sbjct: 739 NLFEILYHARPHALSLFIGMSTATIGGFIYPTYSVFFTSFMNVFAGNPADFL-SQGHFWA 797
Query: 765 IVFVAVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGA 824
++F+ +A A + +F G+A L + +R F V+ + +FD +++SG +
Sbjct: 798 LMFLVLAAAQGICSFLMTFFMGIASESLTRDLRNKLFRNVLSQHIGFFDSPQNASGKIST 857
Query: 825 RLSTDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQV 884
RL+TD ++R + ++ + +++ G+ +AF WQ+A +++A+ P++ Y++
Sbjct: 858 RLATDVPNLRTAIDFRFSTVITTLVSMVAGIGLAFFYGWQMALLIIAILPIVAFGQYLRG 917
Query: 885 KVLKGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGI 944
+ G + + + ++ ++A +A+ ++RTV + E+ E + +K + P K+ ++
Sbjct: 918 RRFTGKNVKSASEFADSGKIAIEAIENVRTVQALAREDTFYENFCEKLDIPHKEAIKEAF 977
Query: 945 ISGLGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSD--VFLVFFALSMAAMGVSQSGTL 1002
I GL +G + +LY ++ C + G L+ T V V +A++++ + + +
Sbjct: 978 IQGLSYGCASSVLYLLNTCAYRMGLALIITDPPTMQPMRVLRVMYAITISTSTLGFATSY 1037
Query: 1003 VPDSTNAKSAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFN 1062
P+ A A IF +L + S+IDS +G +++ G + F +V F YP R +++I
Sbjct: 1038 FPEYAKATFAGGIIFGMLRKISKIDSLSLAG-EKKKLYGKVIFKNVRFAYPERPEIEILK 1096
Query: 1063 DLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMG 1122
L ++ G+T+ALVG SG GKSTV++LL+RFYD G I +DG EI+ + + R Q+
Sbjct: 1097 GLSFSVEPGQTLALVGPSGCGKSTVVALLERFYDTLGGEIFIDGSEIKTLNPEHTRSQIA 1156
Query: 1123 LVSQEPILFNDTVRANIAYG-KGGDATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGI 1181
+VSQEP LF+ ++ NI YG T A++ AA LAN H FI L +G++T VG+RG
Sbjct: 1157 IVSQEPTLFDCSIAENIIYGLDPSSVTMAQVEEAARLANIHNFIAELPEGFETRVGDRGT 1216
Query: 1182 QLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRL 1241
QLSGGQKQR+AIARA+V+NPKILLLDEATSALD ESEKVVQ+ALDR RT I++AHRL
Sbjct: 1217 QLSGGQKQRIAIARALVRNPKILLLDEATSALDTESEKVVQEALDRAREGRTCIVIAHRL 1276
Query: 1242 STIKGADLIAVVKNGVIAEKGKHEALLHKGGDYASL 1277
+T+ AD IAVV NG I EKG H L+ + G Y L
Sbjct: 1277 NTVMNADCIAVVSNGTIIEKGTHTQLMSEKGAYYKL 1312
Score = 410 bits (1055), Expect = e-114
Identities = 239/599 (39%), Positives = 346/599 (56%), Gaps = 32/599 (5%)
Query: 707 LYRLAYFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYKPA------------- 753
LYR Y E +L +GT+ AV+ GA +P++ +L K+ F
Sbjct: 65 LYR--YTTTLEKLLLFIGTLVAVITGAGLPLMSILQGKVSQAFINEQIVINNNGSTFLPT 122
Query: 754 ------DELRHD--SKVWAIVFVAVAV--ASLLIIPCRFYFFGVAGGKLIQRIRKLCFEK 803
+ HD + VW+ + V + A + + C Y ++ R+R+ +
Sbjct: 123 GQNYTKTDFEHDVMNVVWSYAAMTVGMWAAGQITVTCYLY----VAEQMNNRLRREFVKS 178
Query: 804 VVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQNIATIIVGMVIAFQASW 863
++ E+SWFD + SG L +L + V+ GD +G+ Q ++ I G ++AF SW
Sbjct: 179 ILRQEISWFDT--NHSGTLATKLFDNLERVKEGTGDKIGMAFQYLSQFITGFIVAFTHSW 236
Query: 864 QLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEK 923
QL ++LA+ P+ L G+ K + F+ Y +A +V + + SIRTV S
Sbjct: 237 QLTLVMLAVTPIQALCGFAIAKSMSTFAIRETLRYAKAGKVVEETISSIRTVVSLNGLRY 296
Query: 924 VMELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVF 983
+E Y E K GV +G+ G+ FG+ + A FY G V DG F D+
Sbjct: 297 ELERYSTAVEEAKKAGVLKGLFLGISFGAMQASNFISFALAFYIGVGWVHDGSLNFGDML 356
Query: 984 LVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDSSDESGMTLEEVKGDI 1043
F ++ M +M + +G + A+ AA+ I+ +LD+K IDSS ++G ++KGDI
Sbjct: 357 TTFSSVMMGSMALGLAGPQLAVLGTAQGAASGIYEVLDRKPVIDSSSKAGRKDMKIKGDI 416
Query: 1044 EFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHIT 1103
+V F YP+R DV I + L + +G+TVALVG SG GKST+ISLL R+YD G IT
Sbjct: 417 TVENVHFTYPSRPDVPILRGMNLRVNAGQTVALVGSSGCGKSTIISLLLRYYDVLKGKIT 476
Query: 1104 LDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQ 1163
+DG++++ + +++LR+ + +VSQEP LFN T+ NI+ GK G T E+VAA ++ANA +
Sbjct: 477 IDGVDVRDINLEFLRKNVAVVSQEPALFNCTIEENISLGKEG-ITREEMVAACKMANAEK 535
Query: 1164 FIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQD 1223
FI +L GY+T+VG+RG QLSGGQKQR+AIARA+V+NPKILLLDEATSALDAESE +VQ
Sbjct: 536 FIKTLPNGYNTLVGDRGTQLSGGQKQRIAIARALVRNPKILLLDEATSALDAESEGIVQQ 595
Query: 1224 ALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLHKGGDYASLVALHT 1282
ALD+ RTTII+AHRLSTI+ ADLI KNG + E G H AL+ + G Y LV T
Sbjct: 596 ALDKAAKGRTTIIIAHRLSTIRNADLIISCKNGQVVEVGDHRALMAQQGLYYDLVTAQT 654
>PMD1_SCHPO (P36619) Leptomycin B resistance protein pmd1
Length = 1362
Score = 795 bits (2053), Expect = 0.0
Identities = 488/1297 (37%), Positives = 720/1297 (54%), Gaps = 65/1297 (5%)
Query: 33 DVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILIFGTMINAFGDSTNSKV 92
D K ++ S+AD D +L L GT+ IG GL +PLM L+ G + AF D + K
Sbjct: 72 DTPAKLSGYPRILSYADKWDIMLQLAGTITGIGAGLGMPLMSLVSGQLAQAFTDLASGKG 131
Query: 93 VDEVSETTTYCDQVSLKFVYLAAGTFVASFLQLTCWMITGERQSARIRGLYLKTILRQDV 152
T D L F+Y+A G F S++ ++I GER + RIR YL IL Q++
Sbjct: 132 ASSFQHTV---DHFCLYFIYIAIGVFGCSYIYTVTFIIAGERIARRIRQDYLHAILSQNI 188
Query: 153 SFFDKETNTGEVVGRMSGDTVLIKDAMGEKVGQFIQFMSTFIGGFVIAFTKGWLLTVVML 212
+FD+ GE+ R++ DT I+D +GEKVG ++TF+ GFVIAF + W T+++
Sbjct: 189 GYFDR-LGAGEITTRITTDTNFIQDGLGEKVGLVFFAIATFVSGFVIAFIRHWKFTLILS 247
Query: 213 SSIPLLILSGSMTSMVIAKASSTGQAAYSKSAGVVEQTIGSIRTVASFTGEKQATANYNR 272
S P + + I K + A ++S+ VE+ +IR +F + YN+
Sbjct: 248 SMFPAICGGIGLGVPFITKNTKGQIAVVAESSTFVEEVFSNIRNAFAFGTQDILAKLYNK 307
Query: 273 SLIKVYKTAVQEALASGVGFGTLFFVFICSYGLAVWFGGKMIIEKGYTGGDVMTVIFAVL 332
LI + + +A+A G+ G +FFV YGLA W GG+++ ++ FAVL
Sbjct: 308 YLITAQRFGINKAIAMGLMVGWMFFVAYGVYGLAFWEGGRLLHAGDLDVSKLIGCFFAVL 367
Query: 333 IGSTCLGQTSPSLSAFAAGQAAAFKMFETINRKPEIDAYDTSGKKLDDIRGDIELRDVCF 392
I S L SP + +F + +AA K+F+TI+R I+A+ +G + DI+G+IEL+++ F
Sbjct: 368 IASYSLANISPKMQSFVSCASAAKKIFDTIDRVSPINAFTPTGDVVKDIKGEIELKNIRF 427
Query: 393 SYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLK 452
YPTRP+ L+ + FSL PSG ALVG SGSGKST++ L+ERFYDP G+V +DG +L+
Sbjct: 428 VYPTRPEVLVLDNFSLVCPSGKITALVGASGSGKSTIIGLVERFYDPIGGQVFLDGKDLR 487
Query: 453 EFQLKWIRQKIGLVSQEPVLFTCSIKENIAYG-----KDCATDEEI--RV--AAELANAA 503
+ +R +I LV QEPVLF ++ ENI YG K + EE+ RV AA+LANA
Sbjct: 488 TLNVASLRNQISLVQQEPVLFATTVFENITYGLPDTIKGTLSKEELERRVYDAAKLANAY 547
Query: 504 KFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLDEATSALDAESERIVQ 563
FI LP+ T VG+ G +SGGQKQR+AIARA++ DP+ILLLDEATSALD++SE +VQ
Sbjct: 548 DFIMTLPEQFSTNVGQRGFLMSGGQKQRIAIARAVISDPKILLLDEATSALDSKSEVLVQ 607
Query: 564 EALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAELTNDPNGAYSQLIRL- 622
+AL+ +RTTIV+AHRLSTIRN D I V++ GKIVE+GSH EL D NGAY++L+
Sbjct: 608 KALDNASRSRTTIVIAHRLSTIRNADNIVVVNAGKIVEQGSHNELL-DLNGAYARLVEAQ 666
Query: 623 --------QEMKRSEQNDANDKNKPNSIVHSGRQSSQRSFSLRSISQGSAGNSGRHSFS- 673
QEM E DA + S + S +S + ++ + +
Sbjct: 667 KLSGGEKDQEMVEEELEDAPREIPITSFGDDDEDNDMASLEAPMMSHNTDTDTLNNKLNE 726
Query: 674 --------------ASYVAPTTD----GFLETEDGGPQASPSKNSSPPEVPLYRLAYF-- 713
AS + P G L E P+ S + E+ +F
Sbjct: 727 KDNVVFEDKTLQHVASEIVPNLPPADVGELNEE---PKKSKKSKKNNHEINSLTALWFIH 783
Query: 714 ----NKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYK-PADELRHDSKVWAIVFV 768
EI LL+G + +++ GA PV + ++ ++ F + + H V+A+ ++
Sbjct: 784 SFVRTMIEIICLLIGILASMICGAAYPVQAAVFARFLNIFTDLSSTDFLHKVNVFAVYWL 843
Query: 769 AVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLST 828
+A+ + A ++QRIR F ++ +V +FD E++ GA+ LST
Sbjct: 844 ILAIVQFFAYAISNFAMTYAMEAVLQRIRYHLFRTLLRQDVEFFDRSENTVGAITTSLST 903
Query: 829 DAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLK 888
S+ L G LG Q + II +++ W+L + L+ +P++ GY +V+ L
Sbjct: 904 KIQSLEGLSGPTLGTFFQILTNIISVTILSLATGWKLGLVTLSTSPVIITAGYYRVRALD 963
Query: 889 GFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGII--- 945
Y+E++ A ++ +IRTV+S EE V Y C+ IK G I
Sbjct: 964 QVQEKLSAAYKESAAFACESTSAIRTVASLNREENVFAEY---CDSLIKPGRESAIASLK 1020
Query: 946 SGLGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLV-- 1003
SGL F ++ + + ++A F+ G+ L+ G+ + F A+ G+ Q+G
Sbjct: 1021 SGLFFSAAQGVTFLINALTFWYGSTLMRKGEYNIVQFYTCFIAI---VFGIQQAGQFFGY 1077
Query: 1004 -PDSTNAKSAAASIFAILDQKSQIDSSDESGMTLEEVK-GDIEFNHVSFKYPTRLDVQIF 1061
D T AK+AA I + + K +ID+ G +E ++ IEF V F YPTR +++
Sbjct: 1078 SADVTKAKAAAGEIKYLSESKPKIDTWSTEGKKVESLQSAAIEFRQVEFSYPTRRHIKVL 1137
Query: 1062 NDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQM 1121
L L ++ G+ VA VG SG GKST I L++RFYD D+G + +DG+ ++ + R+Q+
Sbjct: 1138 RGLNLTVKPGQFVAFVGSSGCGKSTTIGLIERFYDCDNGAVLVDGVNVRDYNINDYRKQI 1197
Query: 1122 GLVSQEPILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGI 1181
LVSQEP L+ TVR NI G D +E E++ A + AN H+FI L GY+T+ G++G
Sbjct: 1198 ALVSQEPTLYQGTVRENIVLGASKDVSEEEMIEACKKANIHEFILGLPNGYNTLCGQKGS 1257
Query: 1182 QLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRL 1241
LSGGQKQR+AIARA+++NPKILLLDEATSALD+ SEKVVQ+AL+ RTT+ +AHRL
Sbjct: 1258 SLSGGQKQRIAIARALIRNPKILLLDEATSALDSHSEKVVQEALNAASQGRTTVAIAHRL 1317
Query: 1242 STIKGADLIAVVKNGVIAEKGKHEALLHKGGDYASLV 1278
S+I+ AD I V GVIAE G H L+ + G Y LV
Sbjct: 1318 SSIQDADCIFVFDGGVIAEAGTHAELVKQRGRYYELV 1354
Score = 346 bits (888), Expect = 2e-94
Identities = 218/625 (34%), Positives = 340/625 (53%), Gaps = 18/625 (2%)
Query: 13 SVQPVEDHDSNQDSEKSKDKDVTTKTV----PLYKLFSFADPSDRLL-MLMGTLGAIGNG 67
++ P + + N++ +KSK + L+ + SF ++ +L+G L ++ G
Sbjct: 747 NLPPADVGELNEEPKKSKKSKKNNHEINSLTALWFIHSFVRTMIEIICLLIGILASMICG 806
Query: 68 LSIPLMILIFGTMINAFGDSTNSKVVDEVSETTTYCDQVSLKFVYLAAGTFVASFLQLTC 127
+ P+ +F +N F D +++ + +V+ Y ++ LA F A +
Sbjct: 807 AAYPVQAAVFARFLNIFTDLSSTDFLHKVNVFAVY-------WLILAIVQFFAYAISNFA 859
Query: 128 WMITGERQSARIRGLYLKTILRQDVSFFDKETNT-GEVVGRMSGDTVLIKDAMGEKVGQF 186
E RIR +T+LRQDV FFD+ NT G + +S ++ G +G F
Sbjct: 860 MTYAMEAVLQRIRYHLFRTLLRQDVEFFDRSENTVGAITTSLSTKIQSLEGLSGPTLGTF 919
Query: 187 IQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKSAGV 246
Q ++ I +++ GW L +V LS+ P++I +G + + AAY +SA
Sbjct: 920 FQILTNIISVTILSLATGWKLGLVTLSTSPVIITAGYYRVRALDQVQEKLSAAYKESAAF 979
Query: 247 VEQTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGVGFGTLFFVFICSYGLA 306
++ +IRTVAS E+ A Y SLIK + + +L SG+ F V L
Sbjct: 980 ACESTSAIRTVASLNREENVFAEYCDSLIKPGRESAIASLKSGLFFSAAQGVTFLINALT 1039
Query: 307 VWFGGKMIIEKGYTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETINRKP 366
W+G ++ + Y T A++ G GQ + +AAA ++ KP
Sbjct: 1040 FWYGSTLMRKGEYNIVQFYTCFIAIVFGIQQAGQFFGYSADVTKAKAAAGEIKYLSESKP 1099
Query: 367 EIDAYDTSGKKLDDIRGD-IELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSG 425
+ID + T GKK++ ++ IE R V FSYPTR + G +L++ G A VG SG G
Sbjct: 1100 KIDTWSTEGKKVESLQSAAIEFRQVEFSYPTRRHIKVLRGLNLTVKPGQFVAFVGSSGCG 1159
Query: 426 KSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYG- 484
KST + LIERFYD +G VL+DG+N++++ + R++I LVSQEP L+ +++ENI G
Sbjct: 1160 KSTTIGLIERFYDCDNGAVLVDGVNVRDYNINDYRKQIALVSQEPTLYQGTVRENIVLGA 1219
Query: 485 -KDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPR 543
KD ++EE+ A + AN +FI LP G +T+ G+ G+ LSGGQKQR+AIARA++++P+
Sbjct: 1220 SKD-VSEEEMIEACKKANIHEFILGLPNGYNTLCGQKGSSLSGGQKQRIAIARALIRNPK 1278
Query: 544 ILLLDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERG 603
ILLLDEATSALD+ SE++VQEALN RTT+ +AHRLS+I++ D I V G I E G
Sbjct: 1279 ILLLDEATSALDSHSEKVVQEALNAASQGRTTVAIAHRLSSIQDADCIFVFDGGVIAEAG 1338
Query: 604 SHAELTNDPNGAYSQLIRLQEMKRS 628
+HAEL G Y +L+ Q + ++
Sbjct: 1339 THAELVKQ-RGRYYELVVEQGLNKA 1362
>MDR5_DROME (Q00748) Multidrug resistance protein homolog 65
(P-glycoprotein 65)
Length = 1302
Score = 760 bits (1962), Expect = 0.0
Identities = 478/1319 (36%), Positives = 726/1319 (54%), Gaps = 72/1319 (5%)
Query: 7 LEGDFVSVQPVEDHDSNQDSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGN 66
+E D VS E +Q+ + T+ + KLF F+ + + G +
Sbjct: 1 MERDEVSTSSSEG--KSQEEAPMAEGLEPTEPIAFLKLFRFSTYGEIGWLFFGFIMCCIK 58
Query: 67 GLSIPLMILI--------------FGTMINAF------GDSTNSKVVDEVSETTTYCDQV 106
L++P +++I FGT N G T + E + Y D +
Sbjct: 59 ALTLPAVVIIYSEFTSMLVDRAMQFGTSSNVHALPLFGGGKTLTNASREENNEALYDDSI 118
Query: 107 SLKFVYLAAGT--FVASFLQLTCWMITGERQSARIRGLYLKTILRQDVSFFDKETNTGEV 164
S + A F++ + + + RQ R+R +++RQD+ + D +
Sbjct: 119 SYGILLTIASVVMFISGIFSVDVFNMVALRQVTRMRIKLFSSVIRQDIGWHDLASKQN-F 177
Query: 165 VGRMSGDTVLIKDAMGEKVGQFIQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLILSGSM 224
M D I+D + EKVG F+ + FI I+F+ GW LT+ + S IPL+IL
Sbjct: 178 TQSMVDDVEKIRDGISEKVGHFVYLVVGFIITVAISFSYGWKLTLAVSSYIPLVILLNYY 237
Query: 225 TSMVIAKASSTGQAAYSKSAGVVEQTIGSIRTVASFTGEKQATANYNRSLIKVYKTAVQE 284
+ K ++ Q +Y+ + + E+ + SIRTV SF GEK Y L+ K + +
Sbjct: 238 VAKFQGKLTAREQESYAGAGNLAEEILSSIRTVVSFGGEKSEVQRYENFLVPARKASQWK 297
Query: 285 ALASGVGFGTLFFVFICSYGLAVWFGGKMIIE------KGYTGGDVMTVIFAVLIGSTCL 338
SG+ L + S A W+G +II+ K YT +M F +++G+ +
Sbjct: 298 GAFSGLSDAVLKSMLYLSCAGAFWYGVNLIIDDRNVENKEYTPAILMIAFFGIIVGADNI 357
Query: 339 GQTSPSLSAFAAGQAAAFKMFETINRKPEIDAYDTSGKKLD-DIRGDIELRDVCFSYPTR 397
+T+P L +FA + A +F+ I+ +ID T GK L+ +RGD+E +DV F YP+R
Sbjct: 358 ARTAPFLESFATARGCATNLFKVIDLTSKIDPLSTDGKLLNYGLRGDVEFQDVFFRYPSR 417
Query: 398 PDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLK 457
P+ ++ G ++ + +G T ALVG SG GKST V L++RFYDP G VL+D ++++++ ++
Sbjct: 418 PEVIVHRGLNIRIRAGQTVALVGSSGCGKSTCVQLLQRFYDPVFGSVLLDDLDIRKYNIQ 477
Query: 458 WIRQKIGLVSQEPVLFTCSIKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDTMV 517
W+R I +V QEPVLF +I +NI+YGK AT +EI AA A A +FI LP+ +M+
Sbjct: 478 WLRSNIAVVGQEPVLFLGTIAQNISYGKPGATQKEIEAAATQAGAHEFITNLPESYRSMI 537
Query: 518 GEHGTQLSGGQKQRVAIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTIV 577
GE G+QLSGGQKQR+AIARA++++P+ILLLDEATSALD +SE+ VQ+AL+ RTTIV
Sbjct: 538 GERGSQLSGGQKQRIAIARALIQNPKILLLDEATSALDYQSEKQVQQALDLASKGRTTIV 597
Query: 578 VAHRLSTIRNVDTIAVIHQGKIVERGSHAELTNDPNGAYSQLIRLQEMKRSEQNDANDKN 637
V+HRLS IR D I IH GK++E GSH +L GAY ++R ++ ++ + D
Sbjct: 598 VSHRLSAIRGADKIVFIHDGKVLEEGSHDDLM-ALEGAYYNMVRAGDINMPDEVEKEDS- 655
Query: 638 KPNSIVHSGRQSSQRSFSLRSISQGSAGNSGRHSFSASYVAPTTDGFLETEDGGPQASPS 697
+ +Q S F +S S + P + +D Q++
Sbjct: 656 -----IEDTKQKSLALFE-KSFETSPLNFEKGQKNSVQFEEPIIKALI--KDTNAQSA-- 705
Query: 698 KNSSPPEVPLY-----RLAYFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYK- 751
+PPE P + R+ K E L++GTI+AV G + P ++ + + +
Sbjct: 706 --EAPPEKPNFFRTFSRILQLAKQEWCYLILGTISAVAVGFLYPAFAVIFGEFYAALAEK 763
Query: 752 -PADELRHDSKV-WAIVFVAVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEV 809
P D LR + + WA + +A + L+ + Y F AG L R+R + F +V+ EV
Sbjct: 764 DPEDALRRTAVLSWAC--LGLAFLTGLVCFLQTYLFNYAGIWLTTRMRAMTFNAMVNQEV 821
Query: 810 SWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIV 869
WFDD +S GAL ARLS +A ++ +G L ++Q ++ I + +A +W+LA +
Sbjct: 822 GWFDDENNSVGALSARLSGEAVDIQGAIGYPLSGMIQALSNFISSVSVAMYYNWKLALLC 881
Query: 870 LALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYK 929
LA P++ + ++ K++ K++ EEA ++A +++ +IRTV+ E V+ Y
Sbjct: 882 LANCPIIVGSVILEAKMMSNAVVREKQVIEEACRIATESITNIRTVAGLRREADVIREYT 941
Query: 930 ---QKCEGPIKKGVR-RGIISGLGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLV 985
Q+ E I++ +R RG+++ S+FF YAV C G LV +G+ F D+ V
Sbjct: 942 EEIQRVEVLIRQKLRWRGVLNSTMQASAFF-AYAVALCY---GGVLVSEGQLPFQDIIKV 997
Query: 986 FFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDS-----SDESGMTLEEVK 1040
L +M ++QS P + A A +F ILD+K +I S + L +
Sbjct: 998 SETLLYGSMMLAQSLAFTPAFSAALIAGHRLFQILDRKPKIQSPMGTIKNTLAKQLNLFE 1057
Query: 1041 GDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSG 1100
G + + + F+YPTR D +I N L L + G+TVALVG SG GKST + LLQR+YDPD G
Sbjct: 1058 G-VRYRGIQFRYPTRPDAKILNGLDLEVLKGQTVALVGHSGCGKSTCVQLLQRYYDPDEG 1116
Query: 1101 HITLDGIEIQR-MQVKWLRQQMGLVSQEPILFNDTVRANIAYGKG-GDATEAEIVAAAEL 1158
I +D +IQ + + +R ++G+VSQEP LF ++ NIAYG + EI+AAA+
Sbjct: 1117 TIHIDHDDIQHDLTLDGVRTKLGIVSQEPTLFERSIAENIAYGDNRRSVSMVEIIAAAKS 1176
Query: 1159 ANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESE 1218
ANAH FI SL GYDT +G RG QLSGGQKQR+AIARA+V+NPKILLLDEATSALD +SE
Sbjct: 1177 ANAHSFIISLPNGYDTRMGARGTQLSGGQKQRIAIARALVRNPKILLLDEATSALDLQSE 1236
Query: 1219 KVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLHKGGDYASL 1277
++VQ ALD RT I++AHRLST++ AD+I V++NG + E+G H L+ +GG YA L
Sbjct: 1237 QLVQQALDTACSGRTCIVIAHRLSTVQNADVICVIQNGQVVEQGNHMQLISQGGIYAKL 1295
Score = 338 bits (866), Expect = 7e-92
Identities = 197/532 (37%), Positives = 299/532 (56%), Gaps = 10/532 (1%)
Query: 754 DELRHDSKVWAIVFVAVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFD 813
+ L DS + I+ +V + F + + + R+R F V+ ++ W D
Sbjct: 111 EALYDDSISYGILLTIASVVMFISGIFSVDVFNMVALRQVTRMRIKLFSSVIRQDIGWHD 170
Query: 814 DVEHSSGALGARLSTDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALA 873
+ + D +R + + +G V + I+ + I+F W+L V +
Sbjct: 171 LASKQN--FTQSMVDDVEKIRDGISEKVGHFVYLVVGFIITVAISFSYGWKLTLAVSSYI 228
Query: 874 PLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCE 933
PL+ L Y K +A ++ Y A +A + + SIRTV SF E+ ++ Y+
Sbjct: 229 PLVILLNYYVAKFQGKLTAREQESYAGAGNLAEEILSSIRTVVSFGGEKSEVQRYENFLV 288
Query: 934 GPIKKGVRRGIISGLGFGSSFFMLYAVDACVFYAGARLVEDGKS------TFSDVFLVFF 987
K +G SGL MLY A F+ G L+ D ++ T + + + FF
Sbjct: 289 PARKASQWKGAFSGLSDAVLKSMLYLSCAGAFWYGVNLIIDDRNVENKEYTPAILMIAFF 348
Query: 988 ALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDSSDESGMTLEE-VKGDIEFN 1046
+ + A ++++ + A+ A ++F ++D S+ID G L ++GD+EF
Sbjct: 349 GIIVGADNIARTAPFLESFATARGCATNLFKVIDLTSKIDPLSTDGKLLNYGLRGDVEFQ 408
Query: 1047 HVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDG 1106
V F+YP+R +V + L + IR+G+TVALVG SG GKST + LLQRFYDP G + LD
Sbjct: 409 DVFFRYPSRPEVIVHRGLNIRIRAGQTVALVGSSGCGKSTCVQLLQRFYDPVFGSVLLDD 468
Query: 1107 IEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIG 1166
++I++ ++WLR + +V QEP+LF T+ NI+YGK G AT+ EI AAA A AH+FI
Sbjct: 469 LDIRKYNIQWLRSNIAVVGQEPVLFLGTIAQNISYGKPG-ATQKEIEAAATQAGAHEFIT 527
Query: 1167 SLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALD 1226
+L + Y +++GERG QLSGGQKQR+AIARA+++NPKILLLDEATSALD +SEK VQ ALD
Sbjct: 528 NLPESYRSMIGERGSQLSGGQKQRIAIARALIQNPKILLLDEATSALDYQSEKQVQQALD 587
Query: 1227 RVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLHKGGDYASLV 1278
RTTI+V+HRLS I+GAD I + +G + E+G H+ L+ G Y ++V
Sbjct: 588 LASKGRTTIVVSHRLSAIRGADKIVFIHDGKVLEEGSHDDLMALEGAYYNMV 639
>MDR4_DROME (Q00449) Multidrug resistance protein homolog 49
(P-glycoprotein 49)
Length = 1302
Score = 754 bits (1947), Expect = 0.0
Identities = 458/1283 (35%), Positives = 703/1283 (54%), Gaps = 55/1283 (4%)
Query: 36 TKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILIFGTMINA------------ 83
T+ + LF ++ +R L+++ L A IP ++I+G +
Sbjct: 26 TRKYSYFDLFRYSTRCERFLLVVSLLVATAASAFIPYFMIIYGEFTSLLVDRTVGVGTSS 85
Query: 84 -------FGDSTNSKVVDEVSETTTYCDQVSLKFVYLAAGTFVASFLQLTCWMITGER-- 134
FG + D + + G+ VA FL +T + R
Sbjct: 86 PAFALPMFGGGQQLTNASKEENNQAIIDDATAFGIGSLVGS-VAMFLLITLAIDLANRIA 144
Query: 135 --QSARIRGLYLKTILRQDVSFFDKETNTGEVVGRMSGDTVLIKDAMGEKVGQFIQFMST 192
Q RIR L+L+ +LRQD++++D + + +M+ D +K+ +GEK+ + + T
Sbjct: 145 LNQIDRIRKLFLEAMLRQDIAWYDTSSGSN-FASKMTEDLDKLKEGIGEKIVIVVFLIMT 203
Query: 193 FIGGFVIAFTKGWLLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKSAGVVEQTIG 252
F+ G V AF GW LT+V+LS +P +I + S+ + + + +YS +A VVE+
Sbjct: 204 FVIGIVSAFVYGWKLTLVVLSCVPFIIAATSVVARLQGSLAEKELKSYSDAANVVEEVFS 263
Query: 253 SIRTVASFTGEKQATANYNRSLIKVYKTAVQEALASGVGFGTLFFVFICSYGLAVWFGGK 312
IRTV +F+G+++ + + LI T ++ L SG+G + + LA+W+G
Sbjct: 264 GIRTVFAFSGQEKEKERFGKLLIPAENTGRKKGLYSGMGNALSWLIIYLCMALAIWYGVT 323
Query: 313 MIIEKG------YTGGDVMTVIFAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETINRKP 366
+I+++ YT ++ V+FAV++G+ LG SP + A A AA +F I+R
Sbjct: 324 LILDERDLPDRVYTPAVLVIVLFAVIMGAQNLGFASPHVEAIAVATAAGQTLFNIIDRPS 383
Query: 367 EIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGK 426
++D D G + ++ G I + F YP RPD I G ++ + G T A VG SG GK
Sbjct: 384 QVDPMDEKGNRPENTAGHIRFEGIRFRYPARPDVEILKGLTVDVLPGQTVAFVGASGCGK 443
Query: 427 STVVSLIERFYDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKD 486
ST++ L++RFYDP G V +DG +L+ + W+R +IG+V QEPVLF +I ENI YG+
Sbjct: 444 STLIQLMQRFYDPEAGSVKLDGRDLRTLNVGWLRSQIGVVGQEPVLFATTIGENIRYGRP 503
Query: 487 CATDEEIRVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILL 546
AT +I AA AN FI +LP+G DT VGE G Q+SGGQKQR+AIARA+++ P++LL
Sbjct: 504 SATQADIEKAARAANCHDFITRLPKGYDTQVGEKGAQISGGQKQRIAIARALVRQPQVLL 563
Query: 547 LDEATSALDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHA 606
LDEATSALD SE+ VQ AL TT+VVAHRLSTI N D I + G + E+G+H
Sbjct: 564 LDEATSALDPTSEKRVQSALELASQGPTTLVVAHRLSTITNADKIVFLKDGVVAEQGTHE 623
Query: 607 ELTNDPNGAYSQLIRLQEMKRSEQNDANDKNKPNSIVHSGRQSSQRSFSLRSISQGSAGN 666
EL + G Y +L+ + + K + + D ++ Q SQ +
Sbjct: 624 ELM-ERRGLYCELVSITQRKEATEADEG------AVAGRPLQKSQNLSDEETDDDEEDEE 676
Query: 667 SGRHSFSASYVAPTTDGFLETEDGGPQASPSKNSSPPEV----PLYRLAYFNKPEIPVLL 722
+ + GF + ++ K EV +L N PE ++
Sbjct: 677 EDEEPELQTSGSSRDSGFRASTRRKRRSQRRKKKKDKEVVSKVSFTQLMKLNSPEWRFIV 736
Query: 723 MGTITAVLHGAIMPVIGLLVSKMISTFYKPADEL-RHDSKVWAIVFVAVAVASLLIIPCR 781
+G I +V+HGA P+ GL D++ R + +++FV + + + L +
Sbjct: 737 VGGIASVMHGATFPLWGLFFGDFFGILSDGDDDVVRAEVLKISMIFVGIGLMAGLGNMLQ 796
Query: 782 FYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDAL 841
Y F AG K+ R+RK F ++ ++++FDD +S GAL +RL++D ++V+ G +
Sbjct: 797 TYMFTTAGVKMTTRLRKRAFGTIIGQDIAYFDDERNSVGALCSRLASDCSNVQGATGARV 856
Query: 842 GLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEA 901
G ++Q +AT++VGMV+ F SWQ + L PL+ L+ Y++ + + + AK EEA
Sbjct: 857 GTMLQAVATLVVGMVVGFVFSWQQTLLTLVTLPLVCLSVYLEGRFIMKSAQKAKASIEEA 916
Query: 902 SQVANDAVGSIRTVSSFCAEEKVMELYKQKCEG---PIKKGVR-RGIISGLGFGSSFFML 957
SQVA +A+ +IRTV+ C E +V++ Y Q+ + ++ VR RG++ LG + F +
Sbjct: 917 SQVAVEAITNIRTVNGLCLERQVLDQYVQQIDRVDVACRRKVRFRGLVFALGQAAPF-LA 975
Query: 958 YAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIF 1017
Y + Y G LV + + + D+ V AL + + Q+ P+ +A +A +
Sbjct: 976 YGIS---MYYGGILVAEERMNYEDIIKVAEALIFGSWMLGQALAYAPNVNDAILSAGRLM 1032
Query: 1018 AILDQKSQIDSSDESGM-TLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVAL 1076
+ + S + +S T+E+ +GDI + +V F+YPTR I L L I+ TVAL
Sbjct: 1033 DLFKRTSTQPNPPQSPYNTVEKSEGDIVYENVGFEYPTRKGTPILQGLNLTIKKSTTVAL 1092
Query: 1077 VGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVR 1136
VG SGSGKST + LL R+YDP SG + L G+ + LR ++GLVSQEP+LF+ T+
Sbjct: 1093 VGPSGSGKSTCVQLLLRYYDPVSGSVNLSGVPSTEFPLDTLRSKLGLVSQEPVLFDRTIA 1152
Query: 1137 ANIAYGKG--GDATEAEIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIA 1194
NIAYG D + EI+ AA+ +N H FI +L +GYDT +G+ QLSGGQKQR+AIA
Sbjct: 1153 ENIAYGNNFRDDVSMQEIIEAAKKSNIHNFISALPQGYDTRLGKTS-QLSGGQKQRIAIA 1211
Query: 1195 RAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVAHRLSTIKGADLIAVVK 1254
RA+V+NPKIL+LDEATSALD ESEKVVQ ALD RT + +AHRL+T++ ADLI V+K
Sbjct: 1212 RALVRNPKILILDEATSALDLESEKVVQQALDEARSGRTCLTIAHRLTTVRNADLICVLK 1271
Query: 1255 NGVIAEKGKHEALLHKGGDYASL 1277
GV+ E G H+ L+ YA+L
Sbjct: 1272 RGVVVEHGTHDELMALNKIYANL 1294
Score = 365 bits (937), Expect = e-100
Identities = 206/528 (39%), Positives = 308/528 (58%), Gaps = 9/528 (1%)
Query: 759 DSKVWAIVFVAVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHS 818
D+ + I + +VA L+I I RIRKL E ++ +++W+D S
Sbjct: 114 DATAFGIGSLVGSVAMFLLITLAIDLANRIALNQIDRIRKLFLEAMLRQDIAWYDTSSGS 173
Query: 819 SGALGARLSTDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGL 878
+ A ++++ D ++ +G+ + ++V I T ++G+V AF W+L +VL+ P +
Sbjct: 174 NFA--SKMTEDLDKLKEGIGEKIVIVVFLIMTFVIGIVSAFVYGWKLTLVVLSCVPFIIA 231
Query: 879 NGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKK 938
V ++ + K Y +A+ V + IRTV +F +EK E + +
Sbjct: 232 ATSVVARLQGSLAEKELKSYSDAANVVEEVFSGIRTVFAFSGQEKEKERFGKLLIPAENT 291
Query: 939 GVRRGIISGLGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVF------LVFFALSMA 992
G ++G+ SG+G S+ ++Y A + G L+ D + V+ +V FA+ M
Sbjct: 292 GRKKGLYSGMGNALSWLIIYLCMALAIWYGVTLILDERDLPDRVYTPAVLVIVLFAVIMG 351
Query: 993 AMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKY 1052
A + + V A +A ++F I+D+ SQ+D DE G E G I F + F+Y
Sbjct: 352 AQNLGFASPHVEAIAVATAAGQTLFNIIDRPSQVDPMDEKGNRPENTAGHIRFEGIRFRY 411
Query: 1053 PTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRM 1112
P R DV+I L +++ G+TVA VG SG GKST+I L+QRFYDP++G + LDG +++ +
Sbjct: 412 PARPDVEILKGLTVDVLPGQTVAFVGASGCGKSTLIQLMQRFYDPEAGSVKLDGRDLRTL 471
Query: 1113 QVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQKGY 1172
V WLR Q+G+V QEP+LF T+ NI YG+ AT+A+I AA AN H FI L KGY
Sbjct: 472 NVGWLRSQIGVVGQEPVLFATTIGENIRYGRPS-ATQADIEKAARAANCHDFITRLPKGY 530
Query: 1173 DTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVER 1232
DT VGE+G Q+SGGQKQR+AIARA+V+ P++LLLDEATSALD SEK VQ AL+
Sbjct: 531 DTQVGEKGAQISGGQKQRIAIARALVRQPQVLLLDEATSALDPTSEKRVQSALELASQGP 590
Query: 1233 TTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALLHKGGDYASLVAL 1280
TT++VAHRLSTI AD I +K+GV+AE+G HE L+ + G Y LV++
Sbjct: 591 TTLVVAHRLSTITNADKIVFLKDGVVAEQGTHEELMERRGLYCELVSI 638
Score = 350 bits (897), Expect = 2e-95
Identities = 218/612 (35%), Positives = 343/612 (55%), Gaps = 24/612 (3%)
Query: 23 NQDSEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILIFGTMIN 82
+Q +K KDK+V +K V +L P R +++ G + ++ +G + PL L FG
Sbjct: 704 SQRRKKKKDKEVVSK-VSFTQLMKLNSPEWRFIVV-GGIASVMHGATFPLWGLFFGDFFG 761
Query: 83 AFGDSTNSKVVDEVSETTTYCDQVSLKFVYLAAGTFVASFLQLTCWMITGERQSARIRGL 142
D + V EV + +S+ FV + + + LQ + G + + R+R
Sbjct: 762 ILSDGDDDVVRAEVLK-------ISMIFVGIGLMAGLGNMLQTYMFTTAGVKMTTRLRKR 814
Query: 143 YLKTILRQDVSFFDKETNT-GEVVGRMSGDTVLIKDAMGEKVGQFIQFMSTFIGGFVIAF 201
TI+ QD+++FD E N+ G + R++ D ++ A G +VG +Q ++T + G V+ F
Sbjct: 815 AFGTIIGQDIAYFDDERNSVGALCSRLASDCSNVQGATGARVGTMLQAVATLVVGMVVGF 874
Query: 202 TKGWLLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKSAGVVEQTIGSIRTVASFT 261
W T++ L ++PL+ LS + I K++ +A+ +++ V + I +IRTV
Sbjct: 875 VFSWQQTLLTLVTLPLVCLSVYLEGRFIMKSAQKAKASIEEASQVAVEAITNIRTVNGLC 934
Query: 262 GEKQATANYNRSLIKVYKTAVQEALASGVGFGTLFFVFICSYGLAVWFGGKMIIEKGYTG 321
E+Q Y + + +V ++ G+ F +YG+++++GG ++ E+
Sbjct: 935 LERQVLDQYVQQIDRVDVACRRKVRFRGLVFALGQAAPFLAYGISMYYGGILVAEERMNY 994
Query: 322 GDVMTVIFAVLIGSTCLGQT---SPSLSAFAAGQAAAFKMFETINRKPEI--DAYDTSGK 376
D++ V A++ GS LGQ +P+++ +F+ + +P Y+T K
Sbjct: 995 EDIIKVAEALIFGSWMLGQALAYAPNVNDAILSAGRLMDLFKRTSTQPNPPQSPYNTVEK 1054
Query: 377 KLDDIRGDIELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIERF 436
GDI +V F YPTR I G +L++ TT ALVG SGSGKST V L+ R+
Sbjct: 1055 S----EGDIVYENVGFEYPTRKGTPILQGLNLTIKKSTTVALVGPSGSGKSTCVQLLLRY 1110
Query: 437 YDPTDGEVLIDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKDCATD---EEI 493
YDP G V + G+ EF L +R K+GLVSQEPVLF +I ENIAYG + D +EI
Sbjct: 1111 YDPVSGSVNLSGVPSTEFPLDTLRSKLGLVSQEPVLFDRTIAENIAYGNNFRDDVSMQEI 1170
Query: 494 RVAAELANAAKFIDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLDEATSA 553
AA+ +N FI LPQG DT +G+ +QLSGGQKQR+AIARA++++P+IL+LDEATSA
Sbjct: 1171 IEAAKKSNIHNFISALPQGYDTRLGKT-SQLSGGQKQRIAIARALVRNPKILILDEATSA 1229
Query: 554 LDAESERIVQEALNRIMINRTTIVVAHRLSTIRNVDTIAVIHQGKIVERGSHAELTNDPN 613
LD ESE++VQ+AL+ RT + +AHRL+T+RN D I V+ +G +VE G+H EL N
Sbjct: 1230 LDLESEKVVQQALDEARSGRTCLTIAHRLTTVRNADLICVLKRGVVVEHGTHDELM-ALN 1288
Query: 614 GAYSQLIRLQEM 625
Y+ L +Q++
Sbjct: 1289 KIYANLYLMQQV 1300
>MDR3_CAEEL (P34713) Multidrug resistance protein 3 (P-glycoprotein C)
Length = 1268
Score = 739 bits (1909), Expect = 0.0
Identities = 444/1245 (35%), Positives = 686/1245 (54%), Gaps = 11/1245 (0%)
Query: 42 YKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILIFGTMINAF--GDSTNSKVVDEVSET 99
+ +F AD D +L G + + NG +P LIF + NA G+S +
Sbjct: 32 FDVFRDADYKDYILFSGGLILSAVNGALVPFNSLIFEGIANALMEGESQYQNGTINMPWF 91
Query: 100 TTYCDQVSLKFVYLAAGTFVASFLQLTCWMITGERQSARIRGLYLKTILRQDVSFFDKET 159
++ L++ YL F+ S+ +C ER+ IR YLK++LRQD +FD ET
Sbjct: 92 SSEIKMFCLRYFYLGVALFLCSYFANSCLYTLCERRLHCIRKKYLKSVLRQDAKWFD-ET 150
Query: 160 NTGEVVGRMSGDTVLIKDAMGEKVGQFIQFMSTFIGGFVIAFTKGWLLTVVMLSSIPLLI 219
G + +MS IKD +G+KVG + ++TFI G I F W LT+VM+ ++PL +
Sbjct: 151 TIGGLTQKMSSGIEKIKDGIGDKVGVLVGGVATFISGVSIGFYMCWQLTLVMMITVPLQL 210
Query: 220 LSGSMTSMVIAKASSTGQAAYSKSAGVVEQTIGSIRTVASFTGEKQATANYNRSLIKVYK 279
S +++ + +A+ +AYS + G+ + I IRTV +F + Y L + +
Sbjct: 211 GSMYLSAKHLNRATKNEMSAYSNAGGMANEVIAGIRTVMAFNAQPFEINRYAHQLNEARR 270
Query: 280 TAVQEALASGVGFGTLFFVFICSYGLAVWFGGKMIIEKGYTGGDVMTVIFAVLIGSTCLG 339
+++A+ + + +A W+G + + G V V +AVLIG+ LG
Sbjct: 271 MGIRKAIILAICTAFPLMLMFTCMAVAFWYGATLAAAGAVSSGAVFAVFWAVLIGTRRLG 330
Query: 340 QTSPSLSAFAAGQAAAFKMFETINRKPEIDAYDTSGKKLDDIRGDIELRDVCFSYPTRPD 399
+ +P L A + A +F+ I+ +PEI + GK + I+G + + F+YPTRP+
Sbjct: 331 EAAPHLGAITGARLAIHDIFKVIDHEPEIKCTSSEGKIPEKIQGKLTFDGIEFTYPTRPE 390
Query: 400 ELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVLIDGINLKEFQLKWI 459
I G S + G T ALVG SG GKST + L+ RFY+ G + +DGI ++E+ ++W+
Sbjct: 391 LKILKGVSFEVNPGETVALVGHSGCGKSTSIGLLMRFYNQCAGMIKLDGIPIQEYNIRWL 450
Query: 460 RQKIGLVSQEPVLFTCSIKENIAYGKDCATDEEIRVAAELANAAKFIDKLPQGLDTMVGE 519
R IG+V QEP++F ++ ENI G TD++I A ++ANA +FI KL DT++G
Sbjct: 451 RSTIGIVQQEPIIFVATVAENIRMGDVLITDQDIEEACKMANAHEFICKLSDRYDTVIGA 510
Query: 520 HGTQLSGGQKQRVAIARAILKDPRILLLDEATSALDAESERIVQEALNRIMINRTTIVVA 579
QLSGGQKQRVAIARAI++ P+ILLLDEATSALD ESER+VQ AL++ RTT+ +A
Sbjct: 511 GAVQLSGGQKQRVAIARAIVRKPQILLLDEATSALDTESERMVQTALDKASEGRTTLCIA 570
Query: 580 HRLSTIRNVDTIAVIHQGKIVERGSHAELTNDPNGAYSQLIRLQEMKRSEQNDANDKNKP 639
HRLSTIRN I V QG I ERG+H EL + +G Y+ +++ QE++R++++ D +
Sbjct: 571 HRLSTIRNASKILVFDQGLIAERGTHDELISKDDGIYASMVKAQEIERAKEDTTLDDEED 630
Query: 640 NSIVHSGRQSSQRSFSLRSISQGSAGNSGRHSFSASYVAPTTD-GFLETEDGGPQASPSK 698
S + S S R + Q A +S R S ++ TT E E+ +
Sbjct: 631 EKTHRSFHRDSVTSDEERELQQSLARDSTR--LRQSMISTTTQVPEWEIENAREEMI--- 685
Query: 699 NSSPPEVPLYRLAYFNKPEIPVLLMGTITAVLHGAIMPVIGLLVSKMISTFYKPADELRH 758
E L+ + + PE+ +++ + ++ G P ++ ++ D++
Sbjct: 686 EEGAMEASLFDIFKYASPEMRNIIISLVFTLIRGFTWPAFSIVYGQLFKILSAGGDDVSI 745
Query: 759 DSKVWAIVFVAVAVASLLIIPCRFYFFGVAGGKLIQRIRKLCFEKVVHMEVSWFDDVEHS 818
+ + ++ F+ +A + G AG + R+R F ++ + S+FDD H+
Sbjct: 746 KALLNSLWFILLAFTGGISTLISGSLLGKAGETMSGRLRMDVFRNIMQQDASYFDDSRHN 805
Query: 819 SGALGARLSTDAASVRALVGDALGLLVQNIATIIVGMVIAFQASWQLAFIVLALAPLLGL 878
G+L +RL+TDA +V+A + L ++ I ++ G+ +AF W +A I LA A LL +
Sbjct: 806 VGSLTSRLATDAPNVQAAIDQRLAEVLTGIVSLFCGVGVAFYYGWNMAPIGLATALLLVV 865
Query: 879 NGYVQVKVLKGFSADAKKLYEEASQVANDAVGSIRTVSSFCAEEKVMELYKQKCEGPIKK 938
+ LK EAS++ +++ + +TV + +E + + + + P ++
Sbjct: 866 VQSSVAQYLKFRGQRDMDSAIEASRLVTESISNWKTVQALTKQEYMYDAFTAASKSPHRR 925
Query: 939 GVRRGIISGLGFGSSFFMLYAVDACVFYAGARLVEDGKSTFSDVFLVFFALSMAAMGVSQ 998
+ RG+ L F + + A + G L+ + ST VF V AL+MA+M V
Sbjct: 926 AIVRGLWQSLSFALAGSFVMWNFAIAYMFGLWLISNNWSTPYTVFQVIEALNMASMSVML 985
Query: 999 SGTLVPDSTNAKSAAASIFAILDQKSQIDSSDESGMTLEEVKGDIEFNHVSFKYPTRLDV 1058
+ + P+ A+ +A +F ++ QKS ID+ +G T +KG+I V F YP R
Sbjct: 986 AASYFPEYVRARISAGIMFTMIRQKSVIDNRGLTGDT-PTIKGNINMRGVYFAYPNRRRQ 1044
Query: 1059 QIFNDLCLNIRSGKTVALVGESGSGKSTVISLLQRFYDPDSGHITLDGIEIQRMQVKWLR 1118
+ + ++ G+TVALVG SG GKST I L++R+YD G + +D +I+ + VK LR
Sbjct: 1045 LVLDGFNMSANFGQTVALVGPSGCGKSTTIQLIERYYDALCGSVKIDDSDIRDLSVKHLR 1104
Query: 1119 QQMGLVSQEPILFNDTVRANIAYGKGGDATEAEIVAAAELANAHQFIGSLQKGYDTIVGE 1178
+ LV QEP LFN T+R NI YG + T+ ++ AA LAN H F+ L GYDT VG
Sbjct: 1105 DNIALVGQEPTLFNLTIRENITYGL-ENITQDQVEKAATLANIHTFVMGLPDGYDTSVGA 1163
Query: 1179 RGIQLSGGQKQRVAIARAIVKNPKILLLDEATSALDAESEKVVQDALDRVMVERTTIIVA 1238
G +LSGGQKQRVAIARAIV++PKILLLDEATSALD ESEK+VQ+ALD+ + RT +++A
Sbjct: 1164 SGGRLSGGQKQRVAIARAIVRDPKILLLDEATSALDTESEKIVQEALDKARLGRTCVVIA 1223
Query: 1239 HRLSTIKGADLIAVVKNGVIAEKGKHEALLHKGGDYASLVALHTS 1283
HRLSTI+ AD I V +NG E+G H+ LL + G Y LV +S
Sbjct: 1224 HRLSTIQNADKIIVCRNGKAIEEGTHQTLLARRGLYYRLVEKQSS 1268
>MDR1_LEIEN (Q06034) Multidrug resistance protein 1 (P-glycoprotein 1)
Length = 1280
Score = 667 bits (1722), Expect = 0.0
Identities = 416/1275 (32%), Positives = 693/1275 (53%), Gaps = 56/1275 (4%)
Query: 26 SEKSKDKDVTTKTVPLYKLFSFADPSDRLLMLMGTLGAIGNGLSIPLMILIFGTMINAFG 85
+E+ V +TV ++F +AD +DR+LM+ GT A+ G +P+ IFG +
Sbjct: 42 AEEEVKGTVVRETVGPIEIFRYADATDRVLMIAGTAFAVACGAGMPVFSFIFGRIA---- 97
Query: 86 DSTNSKVVDEVSETTTYCDQVSLKFVYLAAGTFVASFLQLTCWMITGERQSARIRGLYLK 145
++ V + SL VY+ +A + CW + RQ ARIR L+ +
Sbjct: 98 ----MDLMSGVGSAEEKAAKTSLIMVYVGIAMLIACAGHVMCWTVAACRQVARIRLLFFR 153
Query: 146 TILRQDVSFFDKETNTGEVVGRMSGDTVLIKDAMGEKVGQFIQFMSTFIGGFVIAFTKGW 205
+LRQD+ + D E + G + RM+GDT +I++ + +K+ Q I S + G++ F W
Sbjct: 154 AVLRQDIGWHD-EHSPGALTARMTGDTRVIQNGINDKLSQGIMNGSMGVIGYIAGFVFSW 212
Query: 206 LLTVVMLSSIPLLILSGSMTSMVIAKASSTGQAAYSKSAGVVEQTIGSIRTVASFTGEKQ 265
LT++M+ +P +I+ ++ +++K + + + ++K+ + + + +IRTV +F E
Sbjct: 213 ELTLMMIGMMPFIIVMAAIIGSIVSKITESSRKYFAKAGSLATEVMENIRTVQAFGREDY 272
Query: 266 ATANYNRSLIKVYKTAVQEALASGVGFGTLFFVFICSYGLAVWFGGKMIIEKGYTGGDVM 325
+ ++++ +++ LAS + + + SY +A +FG ++ D++
Sbjct: 273 ELERFTKAVLYAQGRGIRKELASNLSAAVIMALMYVSYTVAFFFGSYLVEWGRRDMADII 332
Query: 326 TVIFAVLIGSTCLGQTSPSLSAFAAGQAAAFKMFETINRKPEIDAYDTSGKKLDDIRGDI 385
+ AVL+GS LG +PS +AF +AAA+++F+ I+R P +D D G + + I
Sbjct: 333 STFLAVLMGSFGLGFVAPSRTAFTESRAAAYEIFKAIDRVPPVDI-DAGGVPVPGFKESI 391
Query: 386 ELRDVCFSYPTRPDELIFNGFSLSLPSGTTAALVGQSGSGKSTVVSLIERFYDPTDGEVL 445
E R+V F+YPTRP ++F SL + G A G SG GKS+V+ LI+RFYDP G VL
Sbjct: 392 EFRNVRFAYPTRPGMILFRDLSLKIKCGQKVAFSGASGCGKSSVIGLIQRFYDPIGGAVL 451
Query: 446 IDGINLKEFQLKWIRQKIGLVSQEPVLFTCSIKENIAYGKDCATDEEIRVAAELANAAKF 505
+DG+ ++E L+ R +IG+VSQEP LF ++ EN+ GK ATDEE+ A AN
Sbjct: 452 VDGVRMRELCLREWRDQIGIVSQEPNLFAGTMMENVRMGKPNATDEEVVEACRQANIHDT 511
Query: 506 IDKLPQGLDTMVGEHGTQLSGGQKQRVAIARAILKDPRILLLDEATSALDAESERIVQEA 565
I LP DT VG G+ LSGGQKQR+AIARA++K P ILLLDEATSALD +SE VQ A
Sbjct: 512 IMALPDRYDTPVGPVGSLLSGGQKQRIAIARALVKRPPILLLDEATSALDRKSEMEVQAA 571
Query: 566 LNRIMI--NRTTIVVAHRLSTIRNVDTIAVI-HQG----KIVERGSHAELTNDPNGAYSQ 618
L++++ T +V+AHRL+TIR++D I + H G +I E G+ EL + +G ++
Sbjct: 572 LDQLIQRGGTTVVVIAHRLATIRDMDRIYYVKHDGAEGSRITESGTFDELL-ELDGEFAA 630
Query: 619 LIRLQEMKRSEQNDANDKNKPNSIVHSGRQSSQRSFSLRSISQGSAGNSGRHSFSASYVA 678
+ ++Q + + K + V +++S G+ G A
Sbjct: 631 VAKMQGVLAGDA-------KSGASVRDAKKAS--------------GHLGVILDEADLAQ 669
Query: 679 PTTDGFLETEDGGPQASPSK-NSSPPEVPLYRLAYFNKPEIPVLLMGTITAVLHGAIMPV 737
D P +K +V RL NK + + +G +++V+ G+ P
Sbjct: 670 LDEDVPRTARQNVPIDELAKWEVKHAKVGFLRLMRMNKDKAWAVALGILSSVVIGSARPA 729
Query: 738 IGLLVSKMISTF-----YKPADELRHDSKVWAIVFVAVAVASL--LIIPCRFYFFGVAGG 790
+++ M+ K + LR + ++A +F+ AVA+ I+ F+G AG
Sbjct: 730 SSIVMGHMLRVLGEYSATKDVEALRSGTNLYAPLFIVFAVANFSGWIL---HGFYGYAGE 786
Query: 791 KLIQRIRKLCFEKVVHMEVSWFDDVEHSSGALGARLSTDAASVRALVGDALGLLVQNIAT 850
L +IR L F +++ ++++FD +G L LS D +V L G ++GL VQ +
Sbjct: 787 HLTTKIRVLLFRQIMRQDINFFDIPGRDAGTLAGMLSGDCEAVHQLWGPSIGLKVQTMCI 846
Query: 851 IIVGMVIAFQASWQLAFIVLALAPLLGLNGYVQVKVLKGFSADAKKLYEEASQVANDAVG 910
I G+V+ F W+LA + LA PL+ + ++ G++ + + + +A+
Sbjct: 847 IASGLVVGFIYQWKLALVALACMPLMIGCSLTRRLMINGYTKSREG--DTDDTIVTEALS 904
Query: 911 SIRTVSSFCAEEKVMELYKQKCEGPIKKGVRRGIISGLGFGSSFFMLYAVDACVFYAGAR 970
++RTV+S +E +E ++ + VR+GII+G +G + F+ Y V A F+ G++
Sbjct: 905 NVRTVTSLNMKEDCVEAFQAALREEAPRSVRKGIIAGGIYGITQFIFYGVYALCFWYGSK 964
Query: 971 LVEDGKSTFSDVFLVFFALSMAAMGVSQSGTLVPDSTNAKSAAASIFAILDQKSQIDSSD 1030
L++ G++ F DV + ++ A ++G +A+++A +F+++D+ +D
Sbjct: 965 LIDKGEAEFKDVMIASMSILFGAQNAGEAGAFATKLADAEASAKRVFSVIDRVPDVDIEQ 1024
Query: 1031 ESGMTLEEVKGDIEFNHVSFKYPTRLDVQIFNDLCLNIRSGKTVALVGESGSGKSTVISL 1090
L E DIE+ +V F Y R + + + + L+G++G GKSTVI +
Sbjct: 1025 AGNKDLGE-GCDIEYRNVQFIYSARPKQVVLASVNMRFGDATSNGLIGQTGCGKSTVIQM 1083
Query: 1091 LQRFYDPDSGHITLDGIEIQRMQVKWLRQQMGLVSQEPILFNDTVRANIAYGKGGDATEA 1150
L RFY+ SG I+++G ++ + + R+ + +V QEP LF+ TVR NI Y + G AT+
Sbjct: 1084 LARFYERRSGLISVNGRDLSSLDIAEWRRNISIVLQEPNLFSGTVRENIRYAREG-ATDE 1142
Query: 1151 EIVAAAELANAHQFIGSLQKGYDTIVGERGIQLSGGQKQRVAIARAIVKNPKILLLDEAT 1210
E+ AA LA+ H I GYDT VG +G LSGGQKQR+AIAR +++ P++LLLDEAT
Sbjct: 1143 EVEEAARLAHIHHEIIKWTDGYDTEVGYKGRALSGGQKQRIAIARGLLRRPRLLLLDEAT 1202
Query: 1211 SALDAESEKVVQDALD--RVMVERTTIIVAHRLSTIKGADLIAVVKNGVIAEKGKHEALL 1268
SALD+ +E VQ+ ++ + + TT+ +AHRL+TI+ D I ++ +G I E+G HE L+
Sbjct: 1203 SALDSVTEAKVQEGIEAFQAKYKVTTVSIAHRLTTIRHCDQIILLDSGCIIEQGSHEELM 1262
Query: 1269 HKGGDYASLVALHTS 1283
GG+Y + L+ S
Sbjct: 1263 ALGGEYKTRYDLYMS 1277
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.135 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 139,282,392
Number of Sequences: 164201
Number of extensions: 5843977
Number of successful extensions: 26787
Number of sequences better than 10.0: 1019
Number of HSP's better than 10.0 without gapping: 876
Number of HSP's successfully gapped in prelim test: 143
Number of HSP's that attempted gapping in prelim test: 21137
Number of HSP's gapped (non-prelim): 3085
length of query: 1287
length of database: 59,974,054
effective HSP length: 122
effective length of query: 1165
effective length of database: 39,941,532
effective search space: 46531884780
effective search space used: 46531884780
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 72 (32.3 bits)
Medicago: description of AC146862.19


