

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.18 - phase: 0 /pseudo
(155 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
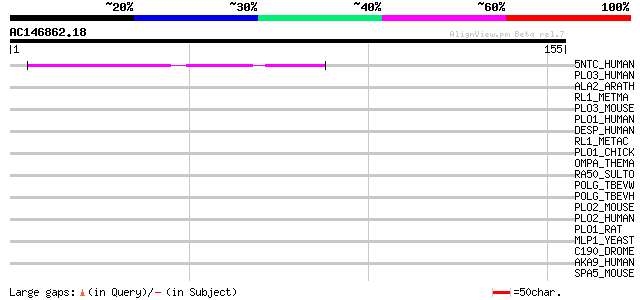
Score E
Sequences producing significant alignments: (bits) Value
5NTC_HUMAN (P49902) Cytosolic purine 5'-nucleotidase (EC 3.1.3.5... 48 1e-05
PLO3_HUMAN (O60568) Procollagen-lysine,2-oxoglutarate 5-dioxygen... 33 0.33
ALA2_ARATH (P98205) Potential phospholipid-transporting ATPase 2... 31 1.3
RL1_METMA (Q8PY52) 50S ribosomal protein L1P 30 1.6
PLO3_MOUSE (Q9R0E1) Procollagen-lysine,2-oxoglutarate 5-dioxygen... 30 2.1
PLO1_HUMAN (Q02809) Procollagen-lysine,2-oxoglutarate 5-dioxygen... 30 2.1
DESP_HUMAN (P15924) Desmoplakin (DP) (250/210 kDa paraneoplastic... 30 2.1
RL1_METAC (Q8TI81) 50S ribosomal protein L1P 30 2.8
PLO1_CHICK (P24802) Procollagen-lysine,2-oxoglutarate 5-dioxygen... 30 2.8
OMPA_THEMA (Q01969) Outer membrane protein alpha precursor 30 2.8
RA50_SULTO (Q96YR5) DNA double-strand break repair rad50 ATPase 29 4.8
POLG_TBEVW (P14336) Genome polyprotein [Contains: Capsid protein... 29 4.8
POLG_TBEVH (Q01299) Genome polyprotein [Contains: Capsid protein... 29 4.8
PLO2_MOUSE (Q9R0B9) Procollagen-lysine,2-oxoglutarate 5-dioxygen... 29 4.8
PLO2_HUMAN (O00469) Procollagen-lysine,2-oxoglutarate 5-dioxygen... 29 4.8
PLO1_RAT (Q63321) Procollagen-lysine,2-oxoglutarate 5-dioxygenas... 29 4.8
MLP1_YEAST (Q02455) MLP1 protein (Myosin-like protein 1) 29 4.8
C190_DROME (Q9VJE5) Restin homolog (Cytoplasmic linker protein 1... 29 4.8
AKA9_HUMAN (Q99996) A-kinase anchor protein 9 (Protein kinase A ... 29 4.8
SPA5_MOUSE (Q7TME2) Sperm associated antigen 5 (Mastrin) (Mitoti... 28 6.2
>5NTC_HUMAN (P49902) Cytosolic purine 5'-nucleotidase (EC 3.1.3.5)
(5'-nucleotidase cytosolic II)
Length = 561
Score = 47.8 bits (112), Expect = 1e-05
Identities = 29/83 (34%), Positives = 42/83 (49%), Gaps = 7/83 (8%)
Query: 6 LGIHGDEILYVGDHIYTDVSQSKVHLRWRTALICRELEDEYSALIRCRGDRESLVELINQ 65
LG G +ILY+GDHI+ D+ +SK WRT L+ EL E + D+ SL E +
Sbjct: 339 LGAKGKDILYIGDHIFGDILKSKKRQGWRTFLVIPELAQE----LHVWTDKSSLFEELQS 394
Query: 66 KEVVGDLFNQLRLALQRRSKDRP 88
++ +L L S +RP
Sbjct: 395 LDI---FLAELYKHLDSSSNERP 414
>PLO3_HUMAN (O60568) Procollagen-lysine,2-oxoglutarate 5-dioxygenase
3 precursor (EC 1.14.11.4) (Lysyl hydroxylase 3) (LH3)
Length = 738
Score = 32.7 bits (73), Expect = 0.33
Identities = 15/44 (34%), Positives = 29/44 (65%)
Query: 96 NMDDEDLTESMQKLLIVMQRLDEKIAPMLEADGELFNSRWGFLS 139
++D + + ++Q L I+++ + IAPML G+L+++ WG LS
Sbjct: 390 SLDADAVLTNLQTLRILIEENRKVIAPMLSRHGKLWSNFWGALS 433
>ALA2_ARATH (P98205) Potential phospholipid-transporting ATPase 2
(EC 3.6.3.1) (Aminophospholipid flippase 2)
Length = 1107
Score = 30.8 bits (68), Expect = 1.3
Identities = 29/108 (26%), Positives = 49/108 (44%), Gaps = 14/108 (12%)
Query: 21 YTDVSQSKVHLRWRTALICRELE-DEYSALIRCRGDR------ESLVEL----INQKEVV 69
+ + S V WR A +C+ LE D Y + DR E++ L IN +
Sbjct: 577 FKEASSLLVDREWRIAEVCQRLEHDLYILGVTAIEDRLQDGVPETIETLRKAGINFWMLT 636
Query: 70 GDLFN---QLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQ 114
GD N Q+ L+ S + Q L +ED++ S++++L+ M+
Sbjct: 637 GDKQNTAIQIALSCNFISPEPKGQLLMIDGKTEEDVSRSLERVLLTMR 684
>RL1_METMA (Q8PY52) 50S ribosomal protein L1P
Length = 213
Score = 30.4 bits (67), Expect = 1.6
Identities = 16/53 (30%), Positives = 34/53 (63%), Gaps = 2/53 (3%)
Query: 69 VGDLFNQLRLALQRRSKDRPAQTLAA--TNMDDEDLTESMQKLLIVMQRLDEK 119
+G+L + A++ RSKDR +A NM+ +DL+E+++ ++ ++R+ +K
Sbjct: 140 IGELIQSKQNAVKLRSKDRLTFHVAVGRRNMNPDDLSENIETIMSRLERVLDK 192
>PLO3_MOUSE (Q9R0E1) Procollagen-lysine,2-oxoglutarate 5-dioxygenase
3 precursor (EC 1.14.11.4) (Lysyl hydroxylase 3) (LH3)
Length = 741
Score = 30.0 bits (66), Expect = 2.1
Identities = 13/44 (29%), Positives = 29/44 (65%)
Query: 96 NMDDEDLTESMQKLLIVMQRLDEKIAPMLEADGELFNSRWGFLS 139
++D + + + + L +++++ + IAPML G+L+++ WG LS
Sbjct: 393 SLDADAVLTNPETLRVLIEQNRKVIAPMLSRHGKLWSNFWGALS 436
>PLO1_HUMAN (Q02809) Procollagen-lysine,2-oxoglutarate 5-dioxygenase
1 precursor (EC 1.14.11.4) (Lysyl hydroxylase 1) (LH1)
Length = 727
Score = 30.0 bits (66), Expect = 2.1
Identities = 17/49 (34%), Positives = 28/49 (56%), Gaps = 1/49 (2%)
Query: 99 DEDLTESMQKLLIVMQRLDEKIAPMLEADGELFNSRWGFLSRAGLWDKS 147
D LTE L +++Q+ IAP++ G L+++ WG LS G + +S
Sbjct: 384 DVALTEP-NSLRLLIQQNKNVIAPLMTRHGRLWSNFWGALSADGYYARS 431
>DESP_HUMAN (P15924) Desmoplakin (DP) (250/210 kDa paraneoplastic
pemphigus antigen)
Length = 2871
Score = 30.0 bits (66), Expect = 2.1
Identities = 24/86 (27%), Positives = 46/86 (52%), Gaps = 15/86 (17%)
Query: 42 LEDEYSALIRCRGDRE----SLVELINQKEVVGDLFNQLRLALQRRSKDRPAQTLAATNM 97
LEDE + L R + D + S E IN+ +V +LR+ +R S++R +
Sbjct: 1500 LEDENARLQRVQYDLQKANSSATETINKLKVQEQELTRLRIDYERVSQER--------TV 1551
Query: 98 DDEDLT---ESMQKLLIVMQRLDEKI 120
D+D+T S+++L + Q+++E++
Sbjct: 1552 KDQDITRFQNSLKELQLQKQKVEEEL 1577
>RL1_METAC (Q8TI81) 50S ribosomal protein L1P
Length = 213
Score = 29.6 bits (65), Expect = 2.8
Identities = 17/53 (32%), Positives = 33/53 (62%), Gaps = 2/53 (3%)
Query: 69 VGDLFNQLRLALQRRSKDRPAQTLAA--TNMDDEDLTESMQKLLIVMQRLDEK 119
VG+L + A++ RSKDR +A +M+ EDL E+++ ++ ++R+ +K
Sbjct: 140 VGELIQSKQNAVKLRSKDRLTFHVAVGRRDMNPEDLAENIETIMSRLERVLDK 192
>PLO1_CHICK (P24802) Procollagen-lysine,2-oxoglutarate 5-dioxygenase
1 precursor (EC 1.14.11.4) (Lysyl hydroxylase 1) (LH1)
Length = 730
Score = 29.6 bits (65), Expect = 2.8
Identities = 14/52 (26%), Positives = 32/52 (60%)
Query: 96 NMDDEDLTESMQKLLIVMQRLDEKIAPMLEADGELFNSRWGFLSRAGLWDKS 147
++D E + ++ + L I++++ IAP++ +L+++ WG LS G + +S
Sbjct: 383 SLDAEVVLKNTETLRILIEQNKSVIAPLVSRHEKLWSNFWGALSPDGYYARS 434
>OMPA_THEMA (Q01969) Outer membrane protein alpha precursor
Length = 400
Score = 29.6 bits (65), Expect = 2.8
Identities = 29/135 (21%), Positives = 63/135 (46%), Gaps = 22/135 (16%)
Query: 2 VENSLGIHGDEILYVGDHIYTDVSQSKVHLRWRTALICREL---EDEYSALIRCRGDRES 58
+E + +H +I+ +IY +S L + A +L ++E SA++
Sbjct: 204 LEGIVNLHEKDII----NIYNKISSVNEELNNKIAATEEKLSRKDEEISAMVE------- 252
Query: 59 LVELINQKEVVGDLFNQ---LRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQR 115
+++K+++ +L+N+ L L ++ D A+ A ++ L KLL +
Sbjct: 253 ----LHEKDII-NLYNKVAALNEDLNKKILDTKAELSAKIESQEKTLNMVYTKLLDTESK 307
Query: 116 LDEKIAPMLEADGEL 130
L+++I+ + E D E+
Sbjct: 308 LNDEISALKEKDAEI 322
>RA50_SULTO (Q96YR5) DNA double-strand break repair rad50 ATPase
Length = 879
Score = 28.9 bits (63), Expect = 4.8
Identities = 19/70 (27%), Positives = 35/70 (49%), Gaps = 3/70 (4%)
Query: 40 RELEDEYSALIRCRGDRESLVELINQKEVVGDLFNQLRLALQRRSKDRPAQTLAATNMDD 99
RELE++Y I +G+ E L E + + D L++ L S+ + +++
Sbjct: 331 RELEEDYKKYIEIKGELEELEEKERKFNSLSDRLKSLKIKL---SEIESKISNRKISINI 387
Query: 100 EDLTESMQKL 109
E+L + +QKL
Sbjct: 388 EELDKELQKL 397
>POLG_TBEVW (P14336) Genome polyprotein [Contains: Capsid protein C
(Core protein); Matrix protein (Envelope protein M);
Major envelope protein E; Nonstructural proteins NS1,
NS2A, NS4A and NS4B; Flavivirin (EC 3.4.21.91) (NS2B/NS3
proteinase); RNA-direct
Length = 3414
Score = 28.9 bits (63), Expect = 4.8
Identities = 34/155 (21%), Positives = 58/155 (36%), Gaps = 23/155 (14%)
Query: 7 GIHGDEILYVGDHI-----------YTDVSQSKVHLRWRTALICRELEDEYSALIRCRGD 55
G+ G + Y+G H+ Y D + W T + +LEDE L G+
Sbjct: 3018 GVEGISLNYLGWHLKKLSTLNGGLFYADDTAG-----WDTKVTNADLEDEEQILRYMEGE 3072
Query: 56 RESLVELINQKEVVGDLFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQR 115
+ L I QK ++ + + R S+D T D + + L +
Sbjct: 3073 HKQLATTIMQK-----AYHAKVVKVARPSRDGGCIMDVITRRDQRGSGQVVTYALNTLTN 3127
Query: 116 LDEKIAPMLEADG--ELFNSRWGFLSRAGLWDKSH 148
+ ++ M+E +G E ++ L R W K H
Sbjct: 3128 IKVQLIRMMEGEGVIEAADAHNPRLLRVERWLKEH 3162
>POLG_TBEVH (Q01299) Genome polyprotein [Contains: Capsid protein C
(Core protein); Matrix protein (Envelope protein M);
Major envelope protein E; Nonstructural proteins NS1,
NS2A, NS4A and NS4B; Flavivirin (EC 3.4.21.91) (NS2B/NS3
proteinase); RNA-direct
Length = 3414
Score = 28.9 bits (63), Expect = 4.8
Identities = 34/155 (21%), Positives = 58/155 (36%), Gaps = 23/155 (14%)
Query: 7 GIHGDEILYVGDHI-----------YTDVSQSKVHLRWRTALICRELEDEYSALIRCRGD 55
G+ G + Y+G H+ Y D + W T + +LEDE L G+
Sbjct: 3018 GVEGISLNYLGWHLKKLSTLNGGLFYADDTAG-----WDTKVTNADLEDEEQILRYMEGE 3072
Query: 56 RESLVELINQKEVVGDLFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVMQR 115
+ L I QK ++ + + R S+D T D + + L +
Sbjct: 3073 HKQLATTIMQK-----AYHAKVVKVARPSRDGGCIMDVITRRDQRGSGQVVTYALNTLTN 3127
Query: 116 LDEKIAPMLEADG--ELFNSRWGFLSRAGLWDKSH 148
+ ++ M+E +G E ++ L R W K H
Sbjct: 3128 IKVQLIRMMEGEGVIEAADAHNPRLLRVERWLKEH 3162
>PLO2_MOUSE (Q9R0B9) Procollagen-lysine,2-oxoglutarate 5-dioxygenase
2 precursor (EC 1.14.11.4) (Lysyl hydroxylase 2) (LH2)
Length = 737
Score = 28.9 bits (63), Expect = 4.8
Identities = 14/52 (26%), Positives = 33/52 (62%)
Query: 96 NMDDEDLTESMQKLLIVMQRLDEKIAPMLEADGELFNSRWGFLSRAGLWDKS 147
++D + + + + L I++++ + IAP++ G+L+++ WG LS G + +S
Sbjct: 390 SVDADVVLTNPRTLKILIEQNRKIIAPLVTRHGKLWSNFWGALSPDGYYARS 441
>PLO2_HUMAN (O00469) Procollagen-lysine,2-oxoglutarate 5-dioxygenase
2 precursor (EC 1.14.11.4) (Lysyl hydroxylase 2) (LH2)
Length = 737
Score = 28.9 bits (63), Expect = 4.8
Identities = 14/52 (26%), Positives = 33/52 (62%)
Query: 96 NMDDEDLTESMQKLLIVMQRLDEKIAPMLEADGELFNSRWGFLSRAGLWDKS 147
++D + + + + L I++++ + IAP++ G+L+++ WG LS G + +S
Sbjct: 390 SVDADVVLTNPRTLKILIEQNRKIIAPLVTRHGKLWSNFWGALSPDGYYARS 441
>PLO1_RAT (Q63321) Procollagen-lysine,2-oxoglutarate 5-dioxygenase 1
precursor (EC 1.14.11.4) (Lysyl hydroxylase 1) (LH1)
Length = 728
Score = 28.9 bits (63), Expect = 4.8
Identities = 17/49 (34%), Positives = 27/49 (54%), Gaps = 1/49 (2%)
Query: 99 DEDLTESMQKLLIVMQRLDEKIAPMLEADGELFNSRWGFLSRAGLWDKS 147
D LTE L++ Q + IAP++ G L+++ WG LS G + +S
Sbjct: 385 DVALTEPNSLRLLIEQNKNV-IAPLMTRHGRLWSNFWGALSADGYYARS 432
>MLP1_YEAST (Q02455) MLP1 protein (Myosin-like protein 1)
Length = 1875
Score = 28.9 bits (63), Expect = 4.8
Identities = 22/79 (27%), Positives = 37/79 (45%), Gaps = 2/79 (2%)
Query: 31 LRWRTALICRELEDEYSALIRCRGDRESLVELINQKEVVGDLFNQLRLALQRRSKDRPAQ 90
LR + +L+ EL+D + L R ++E+ +I Q + + + NQL L R S
Sbjct: 1171 LRQKISLMDVELQDARTKLDNSRVEKENHSSIIQQHDDIMEKLNQLNLL--RESNITLRN 1228
Query: 91 TLAATNMDDEDLTESMQKL 109
L N ++L + KL
Sbjct: 1229 ELENNNNKKKELQSELDKL 1247
>C190_DROME (Q9VJE5) Restin homolog (Cytoplasmic linker protein 190)
(Microtubule binding protein 190) (d-CLIP-190)
Length = 1690
Score = 28.9 bits (63), Expect = 4.8
Identities = 22/74 (29%), Positives = 34/74 (45%), Gaps = 6/74 (8%)
Query: 54 GDRESLVELINQKEVVGDLFNQLRLALQRRSKDRPAQTLAATNMDDEDLTESMQKLLIVM 113
G RESL + + +++R+ Q+R P + + AT SMQ LL
Sbjct: 321 GSRESLTSIGTMNSIATTATSRMRMNAQQRKSSTPVKPILATPKSQ----FSMQDLLREK 376
Query: 114 QRLDEKIAPMLEAD 127
Q+ EK+ M+E D
Sbjct: 377 QQHVEKL--MVERD 388
>AKA9_HUMAN (Q99996) A-kinase anchor protein 9 (Protein kinase A
anchoring protein 9) (PRKA9) (A-kinase anchor protein
450 kDa) (AKAP 450) (A-kinase anchor protein 350 kDa)
(AKAP 350) (hgAKAP 350) (AKAP 120 like protein)
(Hyperion protein) (Yotiao protein
Length = 3911
Score = 28.9 bits (63), Expect = 4.8
Identities = 25/112 (22%), Positives = 54/112 (47%), Gaps = 21/112 (18%)
Query: 40 RELEDEYSALIRCRGDRE-----------SLVELINQK---EVVGDLFNQLRLALQRRSK 85
+EL++EY+ L++ + D E S ++ +N++ + + +++ ++ K
Sbjct: 874 QELQEEYACLLKVKDDLEDSKNKQELEYKSKLKALNEELHLQRINPTTVKMKSSVFDEDK 933
Query: 86 DRPAQTLAATNMDDEDLTESMQKL-------LIVMQRLDEKIAPMLEADGEL 130
A+TL + ++D TE M+KL L + QRL + + + GE+
Sbjct: 934 TFVAETLEMGEVVEKDTTELMEKLEVTKREKLELSQRLSDLSEQLKQKHGEI 985
>SPA5_MOUSE (Q7TME2) Sperm associated antigen 5 (Mastrin) (Mitotic
spindle associated protein p126) (MAP126)
Length = 1165
Score = 28.5 bits (62), Expect = 6.2
Identities = 27/106 (25%), Positives = 51/106 (47%), Gaps = 13/106 (12%)
Query: 41 ELEDEYSALIRCRGDRESLVELINQ--KEVVGDLFNQ---LRLALQRRSKDRPAQTLAAT 95
+LE + + + +SL L+ + +E G L + L LQ + ++ + L A
Sbjct: 937 DLEKSLAEMSTVLQELKSLCSLLQESKEEATGVLQREICELHSRLQAQEEEHQ-EALKAK 995
Query: 96 NMDDEDLTESM-------QKLLIVMQRLDEKIAPMLEADGELFNSR 134
D E L +++ ++LL V+Q+ +EKI ++ G+L N R
Sbjct: 996 EADMEKLNQALCLLRKNEKELLEVIQKQNEKILGQIDKSGQLINLR 1041
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.136 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,812,451
Number of Sequences: 164201
Number of extensions: 601655
Number of successful extensions: 1693
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 1682
Number of HSP's gapped (non-prelim): 28
length of query: 155
length of database: 59,974,054
effective HSP length: 101
effective length of query: 54
effective length of database: 43,389,753
effective search space: 2343046662
effective search space used: 2343046662
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146862.18


