

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146855.1 + phase: 0 /pseudo
(1356 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
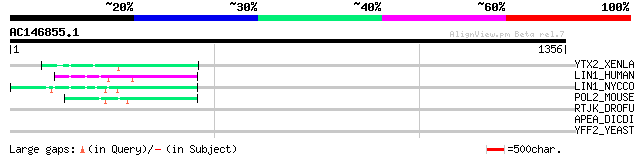
Score E
Sequences producing significant alignments: (bits) Value
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 80 3e-14
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 63 5e-09
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 58 2e-07
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 49 9e-05
RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile elem... 44 0.003
APEA_DICDI (P51173) DNA-(apurinic or apyrimidinic site) lyase (E... 33 4.1
YFF2_YEAST (P43551) Putative 62.1 kDa transcriptional regulatory... 33 6.9
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 80.5 bits (197), Expect = 3e-14
Identities = 93/405 (22%), Positives = 162/405 (39%), Gaps = 38/405 (9%)
Query: 77 HCSILNYSQHFINMSIQDPVKGPWRLTAFYGYPDHGRRRDSWELLRSLHSQSDDPWCIIG 136
H + + + M++ P GP R F + DS D+ I G
Sbjct: 94 HLRVRESGRTYNLMNVYAPTTGPERARFFESLSAYMETIDS-----------DEALIIGG 142
Query: 137 DFNDHLSPSDKRGGPDRPHWLIRGFQEAVSDCNLIDL----PLNGYQFTWFKSIGTSHAK 192
DFN L D R P + +E ++ +L+D+ FT+ + + H
Sbjct: 143 DFNYTLDARD-RNVPKKRDSSESVLRELIAHFSLVDVWREQNPETVAFTYVR-VRDGHVS 200
Query: 193 EARIDRALCTAPWLELFPHASLQTL-VAPMSDHTPLLLQLDPPPWRAPHNSFRFNNSWLI 251
++RIDR ++ L A T+ +AP SDH + L++ P + FNNS L
Sbjct: 201 QSRIDRIYISS---HLMSRAQSSTIRLAPFSDHNCVSLRMSIAPSLPKAAYWHFNNSLLE 257
Query: 252 EPELAHLVKDNW--------------SYYPSSNIVTKLSYCIEDMKYWSKANHPHFNQRK 297
+ A V+D W ++ + KL C E K S +
Sbjct: 258 DEGFAKSVRDTWRGWRAFQDEFATLNQWWDVGKVHLKL-LCQEYTKSVSGQRNAEIEALN 316
Query: 298 QQLKNQIDVMRNTSDASVDPRLIELQNSLANLILQEDVYWRQRSKIFWLKDGDKNSKFFH 357
++ + + + D ++ +E + +L N+ ++ RS++ L D D+ S+FF+
Sbjct: 317 GEVLDLEQRLSGSEDQALQCEYLERKEALRNMEQRQARGAFVRSRMQLLCDMDRGSRFFY 376
Query: 358 LTASSRRRKNTISKLRHPNGFWLTSQDDMSAHIHDYFSGLFQA--INGDQQPVITRIQAR 415
+ + I+ L +G L + + ++ LF I+ D +
Sbjct: 377 ALEKKKGNRKQITCLFAEDGTPLEDPEAIRDRARSFYQNLFSPDPISPDACEELWDGLPV 436
Query: 416 VTENDNATLTKPFSIDEFKEAVFSMHSDKSPGPDGLNPGFYQFFF 460
V+E L P ++DE +A+ M +KSPG DGL F+QFF+
Sbjct: 437 VSERRKERLETPITLDELSQALRLMPHNKSPGLDGLTIEFFQFFW 481
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 63.2 bits (152), Expect = 5e-09
Identities = 77/372 (20%), Positives = 152/372 (40%), Gaps = 29/372 (7%)
Query: 109 PDHGRRRDSWELLRSLHSQSDDPWCIIGDFNDHLSPSDKRGGPDRPHWLIRGFQEAVSDC 168
P+ G R ++L L D I+GDFN LS D R + + I+ A+
Sbjct: 116 PNTGAPRFIKQVLSDLQRDLDSHTIIMGDFNTPLSTLD-RSTRQKINKDIQELNSALHQA 174
Query: 169 NLID----LPLNGYQFTWFKSIGTSHAKEARIDRALCTAPWLELFPHASLQTLVAPMSDH 224
+LID L ++T+F + H ++ D L + L + T +SDH
Sbjct: 175 DLIDIYRTLHPKSTEYTFFSA---PHHTYSKTDHILGSKTLLSKCKRTEIITNC--LSDH 229
Query: 225 TPLLLQLDPPPWRAPH------NSFRFNNSWL---IEPELAHLVKDNWSYYPSSNIVTKL 275
+ + L+L H N+ N+ W+ ++ E+ + N + + +
Sbjct: 230 SAIKLELRIKKLTQNHSTTWKLNNLLLNDYWVHNEMKAEIKKFFETNENKDTTYQNLWDT 289
Query: 276 SYCIEDMKYWSKANHPHFNQRKQ------QLKNQIDVMRNTSDASVDPRLIELQNSLANL 329
+ + K+ + H +R + QLK + S AS +I+++ L +
Sbjct: 290 AKAVCRGKFIALNAHKRKQERSKIDTLISQLKELEKQEQTNSKASRRQEIIKIRAELKEI 349
Query: 330 ILQEDVYWRQRSKIFWLKDGDKNSKFFHLTASSRRRKNTISKLRHPNGFWLTSQDDMSAH 389
Q+ + S+ ++ + +K + +R KN I +++ G T ++
Sbjct: 350 ETQKTLQKINESRSWFFEKINKIDRPLARLIKKKREKNQIDTIKNDRGDITTDPTEIQTT 409
Query: 390 IHDYFSGLF----QAINGDQQPVITRIQARVTENDNATLTKPFSIDEFKEAVFSMHSDKS 445
I +Y+ L+ + + + + T R+ + + +L +P + E + + S+ + KS
Sbjct: 410 IREYYKHLYANKLENLEEMDKFLDTYTLPRLNQEEVESLNRPITSSEIEAIINSLPNKKS 469
Query: 446 PGPDGLNPGFYQ 457
PGP+G FYQ
Sbjct: 470 PGPEGFTAEFYQ 481
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 58.2 bits (139), Expect = 2e-07
Identities = 96/494 (19%), Positives = 192/494 (38%), Gaps = 52/494 (10%)
Query: 1 MNVIAWNCRGLGNVKAVPCIKDLVRVYKPDIVILIETLCNNNKISGLKYAIGFDYHFSVD 60
+++ + N GL + D ++ KPDI + E+ LK G+ F +
Sbjct: 7 LSIFSINVNGLNCPLKRHRLADWIQKLKPDICCIQESHLTLKDKYRLKVK-GWSSIFQAN 65
Query: 61 CIGRSGGIAVLWRNSAHCSILNYSQ----HFINMSIQDPVKGPWR-----LTAFYGYPDH 111
+ GIA+L+ ++ + HFI VKG + + Y P+H
Sbjct: 66 GKQKKAGIAILFADAIGFKPTKIRKDKDGHFIF------VKGNTQYDEISIINIYA-PNH 118
Query: 112 GRRRDSWELLRSLHSQSDDPWCIIGDFNDHLSPSDKRGGPDRPHWLIRGFQEAVSDCNLI 171
+ E L + + ++GDFN L+ D R + I + +L
Sbjct: 119 NAPQFIRETLTDMSNLISSTSIVVGDFNTPLAVLD-RSSKKKLSKEILDLNSTIQHLDLT 177
Query: 172 DLPL----NGYQFTWFKSIGTSHAKEARIDRALCTAPWLELFPHASLQTLVAPMSDHTPL 227
D+ N ++T+F S +H ++ID L L F ++ + SDH +
Sbjct: 178 DIYRTFHPNKTEYTFFSS---AHGTYSKIDHILGHKSNLSKFK--KIEIIPCIFSDHHGI 232
Query: 228 LLQLD--------PPPWRAPHNSFRFNNSWLIEP---ELAHLVKDN-------WSYYPSS 269
++L+ W+ N+ ++W+I+ E+ ++ N + + ++
Sbjct: 233 KVELNNNRNLHTHTKTWKL--NNLMLKDTWVIDEIKKEITKFLEQNNNQDTNYQNLWDTA 290
Query: 270 NIVTKLSYCIEDMKYWSKANHPHFNQRKQQLKNQIDVMRNTSDASVDPRLIELQNSLANL 329
V + + I + K N LK + S + +++ L +
Sbjct: 291 KAVLRGKF-IALQAFLKKTEREEVNNLMGHLKQLEKEEHSNPKPSRRKEITKIRAELNEI 349
Query: 330 ILQEDVYWRQRSKIFWLKDGDKNSKFFHLTASSRRRKNTISKLRHPNGFWLTSQDDMSAH 389
+ + +SK ++ + +K K +R K+ IS +R+ N T ++
Sbjct: 350 ENKRIIQQINKSKSWFFEKINKIDKPLANLTRKKRVKSLISSIRNGNDEITTDPSEIQKI 409
Query: 390 IHDYFSGLFQAINGDQQPVITRIQA----RVTENDNATLTKPFSIDEFKEAVFSMHSDKS 445
+++Y+ L+ + + + ++A R+++ + L +P S E + ++ KS
Sbjct: 410 LNEYYKKLYSHKYENLKEIDQYLEACHLPRLSQKEVEMLNRPISSSEIASTIQNLPKKKS 469
Query: 446 PGPDGLNPGFYQFF 459
PGPDG FYQ F
Sbjct: 470 PGPDGFTSEFYQTF 483
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 48.9 bits (115), Expect = 9e-05
Identities = 72/349 (20%), Positives = 133/349 (37%), Gaps = 29/349 (8%)
Query: 134 IIGDFNDHLSPSDKRGGPDRPHWLIRGFQEAVSDCNLIDLPLNGYQFT-WFKSIGTSHAK 192
I+GDFN LS D+ ++ E + +L D+ Y T + H
Sbjct: 168 IVGDFNTPLSSKDRSWKQKLNRDTVK-LTEVMKQMDLTDIYRTFYPKTKGYTFFSAPHGT 226
Query: 193 EARIDRALCTAPWLELFPHASLQTLVAPMSDHTPLLLQLD------PPPWRAPHNSFRFN 246
++ID + L + + + + +SDH L L + P + N+ N
Sbjct: 227 FSKIDHIIGHKTGLNRYKNIEIVPCI--LSDHHGLRLIFNNNINNGKPTFTWKLNNTLLN 284
Query: 247 NSWLIEPELAHLVKDNWSYYPSSNIVTKLSYCIEDMKYW------------SKANHPHFN 294
++ L++ + +KD + + N T + MK + K H +
Sbjct: 285 DT-LVKEGIKKEIKDFLEF--NENEATTYPNLWDTMKAFLRGKLIALSASKKKRETAHTS 341
Query: 295 QRKQQLKNQIDVMRNTSDASVDPRLIELQNSLANLILQEDVYWRQRSKIFWLKDGDKNSK 354
LK N+ S +I+L+ + + + + +++ ++ + +K K
Sbjct: 342 SLTTHLKALEKKEANSPKRSRRQEIIKLRGEINQVETRRTIQRINQTRSWFFEKINKIDK 401
Query: 355 FFHLTASSRRRKNTISKLRHPNGFWLTSQDDMSAHIHDYFSGLFQAI--NGDQQP-VITR 411
R K I+K+R+ G T +++ I ++ L+ N D+ + R
Sbjct: 402 PLARLTKGHRDKILINKIRNEKGDITTDPEEIQNTIRSFYKRLYSTKLENLDEMDKFLDR 461
Query: 412 IQARVTENDNAT-LTKPFSIDEFKEAVFSMHSDKSPGPDGLNPGFYQFF 459
Q D L P S E + + S+ + KSPGPDG + FYQ F
Sbjct: 462 YQVPKLNQDQVDHLNSPISPKEIEAVINSLPTKKSPGPDGFSAEFYQTF 510
>RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 43.9 bits (102), Expect = 0.003
Identities = 62/267 (23%), Positives = 106/267 (39%), Gaps = 34/267 (12%)
Query: 213 SLQTLVAPMSDHTPLLLQLDPPPWRAPHNSFRFNNSWLIEPELAHLVKD---NWSYYPSS 269
++QTL SDHTPLL L P P S IE A+L + + +
Sbjct: 203 NVQTLHELSSDHTPLLADLHAMPINKPPRSCLLARGADIERFKAYLTQHIDLSVGIQGTD 262
Query: 270 NIVTKLSYCIEDMKYWSKANHPHFNQRKQ---------------QLKNQIDVMRNTSDAS 314
+I + ++ +K + + P Q + +LK ++ R +
Sbjct: 263 DIDNAIDSFMDILKRAAIRSAPSHQQNVESSRQLQLPPIVASLIRLKRKV---RREYART 319
Query: 315 VDPRLIELQNSLANLILQEDVYWRQRSKIFWL-----KDGDKNSKFFHLTASSRRRKNTI 369
D R+ ++ + LAN L + + R++S+I L DG N + +T + +
Sbjct: 320 GDARIQQIHSRLANR-LHKVLNRRKQSQIDNLLENLDTDGSTNFSLWRITKRYKTQATPN 378
Query: 370 SKLRHPNGFWLTSQDD----MSAHIHDYFSGLFQAINGDQQPVITRIQARVTENDNATLT 425
S +R+P G W + + + H+ F L A V +Q T A
Sbjct: 379 SAIRNPAGGWCRTSREKTEVFANHLEQRFKALAFAPESHSLMVAESLQ---TPFQMALPA 435
Query: 426 KPFSIDEFKEAVFSMHSDKSPGPDGLN 452
P +++E KE V + K+PG D L+
Sbjct: 436 DPVTLEEVKELVSKLKPKKAPGEDLLD 462
>APEA_DICDI (P51173) DNA-(apurinic or apyrimidinic site) lyase (EC
4.2.99.18) (Class II
apurinic/apyrimidinic(AP)-endonuclease)
Length = 361
Score = 33.5 bits (75), Expect = 4.1
Identities = 18/57 (31%), Positives = 29/57 (50%), Gaps = 1/57 (1%)
Query: 1 MNVIAWNCRGLGNVKAVPCIKDLVRVYKPDIVILIETLCNNNKISGLKYAIGFDYHF 57
M +I+WN G +V + + V PD++ L ET N + I + G++YHF
Sbjct: 105 MKIISWNVAGFKSVLSKG-FTECVEKENPDVLCLQETKINPSNIKKDQMPKGYEYHF 160
>YFF2_YEAST (P43551) Putative 62.1 kDa transcriptional regulatory
protein in DAK3-ALR2 intergenic region
Length = 465
Score = 32.7 bits (73), Expect = 6.9
Identities = 17/55 (30%), Positives = 28/55 (50%), Gaps = 2/55 (3%)
Query: 761 VRNLSLTISKTVFGKSVNLGVLNLCREQVKRCSSNRLLKQFLHIVWELFSFQPLY 815
VRNL VF KS + G++ + + + +C RL L+++W L + LY
Sbjct: 50 VRNLKKVDDVQVFSKSSSGGIMKVPKALIDQCL--RLYNDKLYVIWPLLCYDDLY 102
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.338 0.147 0.484
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 147,560,996
Number of Sequences: 164201
Number of extensions: 5931266
Number of successful extensions: 17534
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 17520
Number of HSP's gapped (non-prelim): 11
length of query: 1356
length of database: 59,974,054
effective HSP length: 122
effective length of query: 1234
effective length of database: 39,941,532
effective search space: 49287850488
effective search space used: 49287850488
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 72 (32.3 bits)
Medicago: description of AC146855.1


