

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146819.13 + phase: 0
(97 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
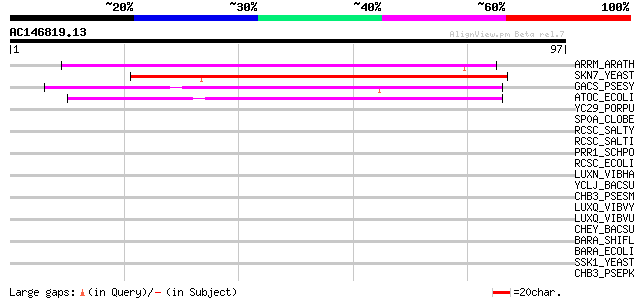
Score E
Sequences producing significant alignments: (bits) Value
ARRM_ARATH (Q9M8Y4) Two-component response regulator ARR22 55 3e-08
SKN7_YEAST (P38889) Putative transcription factor SKN7 (POS9 pro... 45 3e-05
GACS_PSESY (P48027) Sensor protein gacS (EC 2.7.3.-) 41 5e-04
ATOC_ECOLI (Q06065) Acetoacetate metabolism regulatory protein a... 40 8e-04
YC29_PORPU (P51343) Probable transcriptional regulator ycf29 40 0.001
SP0A_CLOBE (P52936) Stage 0 sporulation protein A homolog (Fragm... 39 0.002
RCSC_SALTY (P58662) Sensor protein rcsC (EC 2.7.3.-) (Capsular s... 38 0.004
RCSC_SALTI (Q56128) Sensor protein rcsC (EC 2.7.3.-) (Capsular s... 38 0.004
PRR1_SCHPO (O14283) Transcription factor prr1 (Pombe response re... 38 0.004
RCSC_ECOLI (P14376) Sensor protein rcsC (EC 2.7.3.-) (Capsular s... 38 0.005
LUXN_VIBHA (P54301) Autoinducer 1 sensor kinase/phosphatase luxN... 36 0.019
YCLJ_BACSU (P94413) Hypothetical sensory transduction protein yclJ 35 0.025
CHB3_PSESM (Q886S8) Chemotaxis response regulator protein-glutam... 35 0.025
LUXQ_VIBVY (Q7MD16) Autoinducer 2 sensor kinase/phosphatase luxQ... 35 0.033
LUXQ_VIBVU (Q8D5Z6) Autoinducer 2 sensor kinase/phosphatase luxQ... 35 0.033
CHEY_BACSU (P24072) Chemotaxis protein cheY homolog 35 0.033
BARA_SHIFL (P59342) Sensor protein barA (EC 2.7.3.-) 35 0.043
BARA_ECOLI (P26607) Sensor protein barA (EC 2.7.3.-) 35 0.043
SSK1_YEAST (Q07084) Osomolarity two-component system protein SSK1 34 0.056
CHB3_PSEPK (Q88MS5) Chemotaxis response regulator protein-glutam... 34 0.056
>ARRM_ARATH (Q9M8Y4) Two-component response regulator ARR22
Length = 142
Score = 55.1 bits (131), Expect = 3e-08
Identities = 32/77 (41%), Positives = 46/77 (59%), Gaps = 1/77 (1%)
Query: 10 IGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSR 69
IG + KNG+E++ L E FDLILM + MP DG++ TKKLR+++ IVGV+
Sbjct: 44 IGGISQTAKNGEEAVILHRDGEASFDLILMDKEMPERDGVSTTKKLREMKVTSMIVGVTS 103
Query: 70 HLTNDEYAE-FCNAGLD 85
+E + F AGL+
Sbjct: 104 VADQEEERKAFMEAGLN 120
>SKN7_YEAST (P38889) Putative transcription factor SKN7 (POS9
protein)
Length = 622
Score = 45.1 bits (105), Expect = 3e-05
Identities = 21/67 (31%), Positives = 41/67 (60%), Gaps = 1/67 (1%)
Query: 22 ESLDLISTVER-KFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSRHLTNDEYAEFC 80
+ L IST+E+ ++DL+LM +MP +DG TAT +R + I+ ++ ++ N + +
Sbjct: 408 DGLSAISTLEKYRYDLVLMDIVMPNLDGATATSIVRSFDNETPIIAMTGNIMNQDLITYL 467
Query: 81 NAGLDDL 87
G++D+
Sbjct: 468 QHGMNDI 474
>GACS_PSESY (P48027) Sensor protein gacS (EC 2.7.3.-)
Length = 907
Score = 41.2 bits (95), Expect = 5e-04
Identities = 28/85 (32%), Positives = 43/85 (49%), Gaps = 7/85 (8%)
Query: 7 LIDIGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQ--- 63
L D+G E AV+ G +++ + + FDL+LM MP MDG AT+ +R E
Sbjct: 676 LEDMGAEVVAVEGGYAAVNAVQ--QEAFDLVLMDVQMPGMDGRQATEAIRAWEAERNQSS 733
Query: 64 --IVGVSRHLTNDEYAEFCNAGLDD 86
IV ++ H +E +G+DD
Sbjct: 734 LPIVALTAHAMANEKRSLLQSGMDD 758
>ATOC_ECOLI (Q06065) Acetoacetate metabolism regulatory protein atoC
(Ornithine/arginine decarboxylase inhibitor) (Ornithine
decarboxylase antizyme)
Length = 461
Score = 40.4 bits (93), Expect = 8e-04
Identities = 24/76 (31%), Positives = 38/76 (49%), Gaps = 2/76 (2%)
Query: 11 GVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSRH 70
G ETH NG+ +L L + + D++LM MP MDGI A K++R E ++ ++ +
Sbjct: 28 GFETHCANNGRTALHLFADIHP--DVVLMDIRMPEMDGIKALKEMRSHETRTPVILMTAY 85
Query: 71 LTNDEYAEFCNAGLDD 86
+ E G D
Sbjct: 86 AEVETAVEALRCGAFD 101
>YC29_PORPU (P51343) Probable transcriptional regulator ycf29
Length = 209
Score = 40.0 bits (92), Expect = 0.001
Identities = 22/55 (40%), Positives = 32/55 (58%), Gaps = 2/55 (3%)
Query: 5 EWLIDIGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLE 59
E+LID G NG E+ L + + KFDLI+ IMP++DG +KLR ++
Sbjct: 20 EYLIDQGFNVDIALNGLEAFQLAN--KYKFDLIISDIIMPIIDGYELLEKLRKID 72
>SP0A_CLOBE (P52936) Stage 0 sporulation protein A homolog
(Fragment)
Length = 223
Score = 38.9 bits (89), Expect = 0.002
Identities = 21/48 (43%), Positives = 32/48 (65%), Gaps = 2/48 (4%)
Query: 12 VETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLE 59
V T K+G E+L+LI VERK DL+++ IMP +DG+ +KL ++
Sbjct: 32 VVTGIAKDGSEALELI--VERKPDLVILDIIMPHLDGLGVLEKLNTMQ 77
>RCSC_SALTY (P58662) Sensor protein rcsC (EC 2.7.3.-) (Capsular
synthesis regulator component C)
Length = 948
Score = 38.1 bits (87), Expect = 0.004
Identities = 22/84 (26%), Positives = 44/84 (52%), Gaps = 2/84 (2%)
Query: 2 ILFEWLIDIGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYP 61
+L + L +G + +G ++L+++S + D++L MP MDG T+++R L
Sbjct: 839 LLADQLGSLGYQCKTANDGVDALNVLS--KNAIDIVLSDVNMPNMDGYRLTQRIRQLGLT 896
Query: 62 GQIVGVSRHLTNDEYAEFCNAGLD 85
+VGV+ + +E +G+D
Sbjct: 897 LPVVGVTANALAEEKQRCLESGMD 920
>RCSC_SALTI (Q56128) Sensor protein rcsC (EC 2.7.3.-) (Capsular
synthesis regulator component C)
Length = 948
Score = 38.1 bits (87), Expect = 0.004
Identities = 22/84 (26%), Positives = 44/84 (52%), Gaps = 2/84 (2%)
Query: 2 ILFEWLIDIGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYP 61
+L + L +G + +G ++L+++S + D++L MP MDG T+++R L
Sbjct: 839 LLADQLGSLGYQCKTANDGVDALNVLS--KNAIDIVLSDVNMPNMDGYRLTQRIRQLGLT 896
Query: 62 GQIVGVSRHLTNDEYAEFCNAGLD 85
+VGV+ + +E +G+D
Sbjct: 897 LPVVGVTANALAEEKQRCLESGMD 920
>PRR1_SCHPO (O14283) Transcription factor prr1 (Pombe response
regulator 1)
Length = 539
Score = 38.1 bits (87), Expect = 0.004
Identities = 17/54 (31%), Positives = 34/54 (62%)
Query: 34 FDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSRHLTNDEYAEFCNAGLDDL 87
FDLILM I+P +DG++ T +R ++ I+ ++ +++ ++ + N G+ DL
Sbjct: 412 FDLILMDFILPNLDGLSVTCLIRQYDHNTPILAITSNISMNDAVTYFNHGVTDL 465
>RCSC_ECOLI (P14376) Sensor protein rcsC (EC 2.7.3.-) (Capsular
synthesis regulator component C)
Length = 949
Score = 37.7 bits (86), Expect = 0.005
Identities = 21/84 (25%), Positives = 44/84 (52%), Gaps = 2/84 (2%)
Query: 2 ILFEWLIDIGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYP 61
+L + L +G + +G ++L+++S + D++L MP MDG T+++R L
Sbjct: 839 LLADQLGSLGYQCKTANDGVDALNVLS--KNHIDIVLSDVNMPNMDGYRLTQRIRQLGLT 896
Query: 62 GQIVGVSRHLTNDEYAEFCNAGLD 85
++GV+ + +E +G+D
Sbjct: 897 LPVIGVTANALAEEKQRCLESGMD 920
>LUXN_VIBHA (P54301) Autoinducer 1 sensor kinase/phosphatase luxN
(EC 2.7.3.-) (EC 3.1.3.-)
Length = 849
Score = 35.8 bits (81), Expect = 0.019
Identities = 20/63 (31%), Positives = 36/63 (56%), Gaps = 2/63 (3%)
Query: 6 WLIDIGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIV 65
+L +GV + NG+ ++++ DLILM MPVM+G A++++++L IV
Sbjct: 739 YLNQLGVNSLQANNGENAVEVFKA--NHVDLILMDVQMPVMNGFDASQRIKELSPQTPIV 796
Query: 66 GVS 68
+S
Sbjct: 797 ALS 799
>YCLJ_BACSU (P94413) Hypothetical sensory transduction protein yclJ
Length = 227
Score = 35.4 bits (80), Expect = 0.025
Identities = 24/87 (27%), Positives = 42/87 (47%), Gaps = 3/87 (3%)
Query: 11 GVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSRH 70
G E V +G E + E +DLI++ ++P MDG+T +K+R+ I+ ++
Sbjct: 24 GFEAEFVHDGLEGYQRFT--EENWDLIILDIMLPSMDGVTICRKIRETSTVPIIMLTAKD 81
Query: 71 LTNDEYAEFCNAGLDDLHHEDGIPLAI 97
+D+ F G DD + PL +
Sbjct: 82 TESDQVIGF-EMGADDYVTKPFSPLTL 107
>CHB3_PSESM (Q886S8) Chemotaxis response regulator
protein-glutamate methylesterase group 3 operon (EC
3.1.1.61)
Length = 336
Score = 35.4 bits (80), Expect = 0.025
Identities = 22/50 (44%), Positives = 33/50 (66%), Gaps = 3/50 (6%)
Query: 19 NGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVS 68
NG +++ ++E+ DLILM IMPVMDG+ AT+++ E P IV V+
Sbjct: 34 NGADAVQ--RSIEQTPDLILMDLIMPVMDGVEATRRIM-AETPCAIVIVT 80
>LUXQ_VIBVY (Q7MD16) Autoinducer 2 sensor kinase/phosphatase luxQ
(EC 2.7.3.-) (EC 3.1.3.-)
Length = 857
Score = 35.0 bits (79), Expect = 0.033
Identities = 23/68 (33%), Positives = 37/68 (53%), Gaps = 3/68 (4%)
Query: 17 VKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRD-LEYPGQIVGVSRHLTNDE 75
V++G ++L+ + E FDLILM +P M GI AT+++R+ L+ I +
Sbjct: 763 VQDGTQALEKLK--EHAFDLILMDNQLPKMGGIEATREIRETLKLGTPIYACTADAQEST 820
Query: 76 YAEFCNAG 83
EF +AG
Sbjct: 821 KQEFLSAG 828
>LUXQ_VIBVU (Q8D5Z6) Autoinducer 2 sensor kinase/phosphatase luxQ
(EC 2.7.3.-) (EC 3.1.3.-)
Length = 857
Score = 35.0 bits (79), Expect = 0.033
Identities = 23/68 (33%), Positives = 37/68 (53%), Gaps = 3/68 (4%)
Query: 17 VKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRD-LEYPGQIVGVSRHLTNDE 75
V++G ++L+ + E FDLILM +P M GI AT+++R+ L+ I +
Sbjct: 763 VQDGTQALEKLK--EHAFDLILMDNQLPKMGGIEATREIRETLKLGTPIYACTADAQEST 820
Query: 76 YAEFCNAG 83
EF +AG
Sbjct: 821 KQEFLSAG 828
>CHEY_BACSU (P24072) Chemotaxis protein cheY homolog
Length = 119
Score = 35.0 bits (79), Expect = 0.033
Identities = 25/87 (28%), Positives = 43/87 (48%), Gaps = 3/87 (3%)
Query: 1 MILFEWLIDIGVETHA-VKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLE 59
M++ + L+ G E A +NG ++++ E DL+ M MP MDGITA K+++ ++
Sbjct: 15 MMIKDILVKNGFEVVAEAENGAQAVEKYK--EHSPDLVTMDITMPEMDGITALKEIKQID 72
Query: 60 YPGQIVGVSRHLTNDEYAEFCNAGLDD 86
+I+ S + AG D
Sbjct: 73 AQARIIMCSAMGQQSMVIDAIQAGAKD 99
>BARA_SHIFL (P59342) Sensor protein barA (EC 2.7.3.-)
Length = 918
Score = 34.7 bits (78), Expect = 0.043
Identities = 20/55 (36%), Positives = 29/55 (52%), Gaps = 2/55 (3%)
Query: 34 FDLILMARIMPVMDGITATKKLRDLEYPGQ--IVGVSRHLTNDEYAEFCNAGLDD 86
FDLILM MP MDGI A + + L + Q ++ V+ H + + AG+ D
Sbjct: 712 FDLILMDIQMPDMDGIRACELIHQLPHQRQTPVIAVTAHAMAGQKEKLLGAGMSD 766
>BARA_ECOLI (P26607) Sensor protein barA (EC 2.7.3.-)
Length = 918
Score = 34.7 bits (78), Expect = 0.043
Identities = 20/55 (36%), Positives = 29/55 (52%), Gaps = 2/55 (3%)
Query: 34 FDLILMARIMPVMDGITATKKLRDLEYPGQ--IVGVSRHLTNDEYAEFCNAGLDD 86
FDLILM MP MDGI A + + L + Q ++ V+ H + + AG+ D
Sbjct: 712 FDLILMDIQMPDMDGIRACELIHQLPHQQQTPVIAVTAHAMAGQKEKLLGAGMSD 766
>SSK1_YEAST (Q07084) Osomolarity two-component system protein SSK1
Length = 712
Score = 34.3 bits (77), Expect = 0.056
Identities = 21/56 (37%), Positives = 32/56 (56%), Gaps = 3/56 (5%)
Query: 18 KNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSRHLTN 73
KNG+E++++ E LI M +PV+ GI A K++RD E I G+ + L N
Sbjct: 534 KNGQEAVNIWK--EGGLHLIFMDLQLPVLSGIEAAKQIRDFEKQNGI-GIQKSLNN 586
>CHB3_PSEPK (Q88MS5) Chemotaxis response regulator
protein-glutamate methylesterase group 3 operon (EC
3.1.1.61)
Length = 337
Score = 34.3 bits (77), Expect = 0.056
Identities = 23/50 (46%), Positives = 32/50 (64%), Gaps = 3/50 (6%)
Query: 19 NGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVS 68
NG E++ + E+ DLILM IMPVMDG+ AT+++ E P IV V+
Sbjct: 34 NGAEAVQRCT--EQLPDLILMDLIMPVMDGVEATRRIM-AETPCAIVIVT 80
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.142 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,952,781
Number of Sequences: 164201
Number of extensions: 423656
Number of successful extensions: 1089
Number of sequences better than 10.0: 116
Number of HSP's better than 10.0 without gapping: 49
Number of HSP's successfully gapped in prelim test: 67
Number of HSP's that attempted gapping in prelim test: 1038
Number of HSP's gapped (non-prelim): 116
length of query: 97
length of database: 59,974,054
effective HSP length: 73
effective length of query: 24
effective length of database: 47,987,381
effective search space: 1151697144
effective search space used: 1151697144
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146819.13


