

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146819.12 + phase: 0 /pseudo
(1226 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
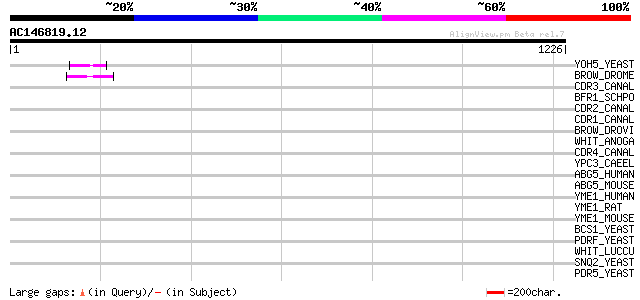
Score E
Sequences producing significant alignments: (bits) Value
YOH5_YEAST (Q08234) Probable ATP-dependent transporter YOL074C/Y... 47 2e-04
BROW_DROME (P12428) Brown protein 46 7e-04
CDR3_CANAL (O42690) Opaque-specific ABC transporter CDR3 44 0.004
BFR1_SCHPO (P41820) Brefeldin A resistance protein 43 0.006
CDR2_CANAL (P78595) Multidrug resistance protein CDR2 42 0.008
CDR1_CANAL (P43071) Multidrug resistance protein CDR1 42 0.008
BROW_DROVI (Q24739) Brown protein 42 0.008
WHIT_ANOGA (Q27256) White protein 40 0.039
CDR4_CANAL (O74676) ABC transporter CDR4 40 0.039
YPC3_CAEEL (Q11180) Putative ABC transporter C05D10.3 in chromos... 40 0.051
ABG5_HUMAN (Q9H222) ATP-binding cassette, sub-family G, member 5... 40 0.051
ABG5_MOUSE (Q99PE8) ATP-binding cassette, sub-family G, member 5... 39 0.11
YME1_HUMAN (Q96TA2) ATP-dependent metalloprotease YME1L1 (EC 3.4... 38 0.15
YME1_RAT (Q925S8) ATP-dependent metalloprotease YME1L1 (EC 3.4.2... 38 0.19
YME1_MOUSE (O88967) ATP-dependent metalloprotease YME1L1 (EC 3.4... 38 0.19
BCS1_YEAST (P32839) Mitochondrial chaperone BCS1 38 0.19
PDRF_YEAST (Q04182) ATP-dependent permease PDR15 37 0.25
WHIT_LUCCU (Q05360) White protein 37 0.33
SNQ2_YEAST (P32568) SNQ2 protein 37 0.33
PDR5_YEAST (P33302) Suppressor of toxicity of sporidesmin 37 0.33
>YOH5_YEAST (Q08234) Probable ATP-dependent transporter
YOL074C/YOL075C
Length = 1294
Score = 47.4 bits (111), Expect = 2e-04
Identities = 25/82 (30%), Positives = 46/82 (55%), Gaps = 6/82 (7%)
Query: 132 ILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFAGKVSYNG 191
ILQ V+ I K + ++GP SGK+ LL ++G+L ++ + +G + +N
Sbjct: 709 ILQSVNAIFKPGMINAIMGPSGSGKSSLLNLISGRLKSSVFAKFDT------SGSIMFND 762
Query: 192 HEMNEFVPQRTAAYVSQNDTHL 213
+++E + + +YVSQ+D HL
Sbjct: 763 IQVSELMFKNVCSYVSQDDDHL 784
>BROW_DROME (P12428) Brown protein
Length = 675
Score = 45.8 bits (107), Expect = 7e-04
Identities = 32/105 (30%), Positives = 50/105 (47%), Gaps = 12/105 (11%)
Query: 125 RRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFA 184
R+K++L ILQD SG +K L +LG +GKT LL A++ +L NL
Sbjct: 41 RKKRELRILQDASGHMKTGDLIAILGGSGAGKTTLLAAISQRLRGNL------------T 88
Query: 185 GKVSYNGHEMNEFVPQRTAAYVSQNDTHLGN*LSEKPWPFQQEFK 229
G V NG M R ++++ Q + ++ + + F FK
Sbjct: 89 GDVVLNGMAMERHQMTRISSFLPQFEINVKTFTAYEHLYFMSHFK 133
>CDR3_CANAL (O42690) Opaque-specific ABC transporter CDR3
Length = 1501
Score = 43.5 bits (101), Expect = 0.004
Identities = 31/112 (27%), Positives = 54/112 (47%), Gaps = 11/112 (9%)
Query: 102 HVGKRALHTITNYMLDLVEYILKRRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLL 161
++ ++ L T + + Y +K + ++ IL ++ G +K +T L+G +GKT LL
Sbjct: 828 YMDRKLLDTSNIFHWRNLTYTVKIKSEERVILNNIDGWVKPGEVTALMGASGAGKTTLLN 887
Query: 162 ALAGKLDPNLKIANEVQFYEQFAGKVSYNGHEMNEFVPQRTAAYVSQNDTHL 213
AL+ +L + +G NG E++ QR+ YV Q D HL
Sbjct: 888 ALSERLTTGVIT----------SGTRMVNGGELDSSF-QRSIGYVQQQDLHL 928
>BFR1_SCHPO (P41820) Brefeldin A resistance protein
Length = 1530
Score = 42.7 bits (99), Expect = 0.006
Identities = 27/96 (28%), Positives = 45/96 (46%), Gaps = 12/96 (12%)
Query: 119 VEYILKRRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQ 178
+ Y ++ + + +L V G + +LT L+G +GKT LL LA ++D +
Sbjct: 887 LNYDIQIKGEHRRLLNGVQGFVVPGKLTALMGESGAGKTTLLNVLAQRVDTGV------- 939
Query: 179 FYEQFAGKVSYNGHEMNEFVPQRTAAYVSQNDTHLG 214
G + NG ++ +RT YV Q D H+G
Sbjct: 940 ----VTGDMLVNGRGLDSTFQRRT-GYVQQQDVHIG 970
>CDR2_CANAL (P78595) Multidrug resistance protein CDR2
Length = 1499
Score = 42.4 bits (98), Expect = 0.008
Identities = 29/93 (31%), Positives = 45/93 (48%), Gaps = 11/93 (11%)
Query: 121 YILKRRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFY 180
Y +K +K+ IL V G +K ++T L+G +GKT LL L+ ++ +
Sbjct: 864 YQVKIKKEDRVILDHVDGWVKPGQITALMGASGAGKTTLLNCLSERVTTGIIT------- 916
Query: 181 EQFAGKVSYNGHEMNEFVPQRTAAYVSQNDTHL 213
G+ NGH ++ QR+ YV Q D HL
Sbjct: 917 ---DGERLVNGHALDSSF-QRSIGYVQQQDVHL 945
>CDR1_CANAL (P43071) Multidrug resistance protein CDR1
Length = 1501
Score = 42.4 bits (98), Expect = 0.008
Identities = 29/93 (31%), Positives = 45/93 (48%), Gaps = 11/93 (11%)
Query: 121 YILKRRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFY 180
Y +K +K+ IL V G +K ++T L+G +GKT LL L+ ++ +
Sbjct: 866 YQVKIKKEDRVILDHVDGWVKPGQITALMGASGAGKTTLLNCLSERVTTGIIT------- 918
Query: 181 EQFAGKVSYNGHEMNEFVPQRTAAYVSQNDTHL 213
G+ NGH ++ QR+ YV Q D HL
Sbjct: 919 ---DGERLVNGHALDSSF-QRSIGYVQQQDVHL 947
>BROW_DROVI (Q24739) Brown protein
Length = 668
Score = 42.4 bits (98), Expect = 0.008
Identities = 31/105 (29%), Positives = 49/105 (46%), Gaps = 12/105 (11%)
Query: 125 RRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFA 184
R++++L IL DVSG LK L +LG +GKT LL A++ +L NL
Sbjct: 38 RKQRELGILHDVSGHLKTGDLIAILGGSGAGKTTLLAAISQRLRGNL------------T 85
Query: 185 GKVSYNGHEMNEFVPQRTAAYVSQNDTHLGN*LSEKPWPFQQEFK 229
G V NG M R ++++ + + ++ + F FK
Sbjct: 86 GDVVLNGMAMERDQMTRISSFLREFEINVKTFTAYDDLYFMSHFK 130
>WHIT_ANOGA (Q27256) White protein
Length = 695
Score = 40.0 bits (92), Expect = 0.039
Identities = 27/81 (33%), Positives = 44/81 (53%), Gaps = 10/81 (12%)
Query: 131 NILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIA-NEVQFYEQFAGKVSY 189
++L++V+G+ K L ++G +GKT LL ALA + P +KI+ N V+ +
Sbjct: 114 HLLKNVTGVAKSGELLAVMGSSGAGKTTLLNALAFRSPPGVKISPNAVR---------AL 164
Query: 190 NGHEMNEFVPQRTAAYVSQND 210
NG +N + AYV Q+D
Sbjct: 165 NGVPVNAEQLRARCAYVQQDD 185
>CDR4_CANAL (O74676) ABC transporter CDR4
Length = 1490
Score = 40.0 bits (92), Expect = 0.039
Identities = 30/93 (32%), Positives = 44/93 (47%), Gaps = 11/93 (11%)
Query: 121 YILKRRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFY 180
Y +K + + IL VSG +K ++T L+G +GKT LL AL+ +L +
Sbjct: 853 YQVKIKSEDRVILDHVSGWVKPGQVTALMGASGAGKTTLLNALSDRLTTGVVT------- 905
Query: 181 EQFAGKVSYNGHEMNEFVPQRTAAYVSQNDTHL 213
G NG ++ QR+ YV Q D HL
Sbjct: 906 ---EGIRLVNGRPLDSSF-QRSIGYVQQQDLHL 934
Score = 35.4 bits (80), Expect = 0.96
Identities = 28/92 (30%), Positives = 46/92 (49%), Gaps = 10/92 (10%)
Query: 120 EYILKRRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGK-----LDPNLKIA 174
EYIL+ +IL+ + G++K LT++LG P +G + L +A + +D + I
Sbjct: 160 EYILRHTGPTFDILKPMDGLIKPGELTVVLGRPGAGCSTFLKTIASQTYGYHIDKDSVIR 219
Query: 175 -NEVQFYE---QFAGKVSYNGHEMNEFVPQRT 202
N + +E + G+V Y N F PQ T
Sbjct: 220 YNSLTPHEIKKHYRGEVVYCAETENHF-PQLT 250
>YPC3_CAEEL (Q11180) Putative ABC transporter C05D10.3 in chromosome
III
Length = 598
Score = 39.7 bits (91), Expect = 0.051
Identities = 26/91 (28%), Positives = 46/91 (49%), Gaps = 12/91 (13%)
Query: 124 KRRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQF 183
KRR ++ IL +VSG+ + +L +LG +GKT L+ L + NL +
Sbjct: 3 KRRVKE--ILHNVSGMAESGKLLAILGSSGAGKTTLMNVLTSRNLTNLDV---------- 50
Query: 184 AGKVSYNGHEMNEFVPQRTAAYVSQNDTHLG 214
G + +G N++ + +A+V Q+D +G
Sbjct: 51 QGSILIDGRRANKWKIREMSAFVQQHDMFVG 81
>ABG5_HUMAN (Q9H222) ATP-binding cassette, sub-family G, member 5
(Sterolin-1)
Length = 651
Score = 39.7 bits (91), Expect = 0.051
Identities = 29/91 (31%), Positives = 45/91 (48%), Gaps = 10/91 (10%)
Query: 125 RRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFA 184
R++ IL+DVS ++ ++ +LG SGKT LL A++G+L F
Sbjct: 61 RQQWTRQILKDVSLYVESGQIMCILGSSGSGKTTLLDAMSGRLGR----------AGTFL 110
Query: 185 GKVSYNGHEMNEFVPQRTAAYVSQNDTHLGN 215
G+V NG + Q +YV Q+DT L +
Sbjct: 111 GEVYVNGRALRREQFQDCFSYVLQSDTLLSS 141
>ABG5_MOUSE (Q99PE8) ATP-binding cassette, sub-family G, member 5
(Sterolin-1)
Length = 652
Score = 38.5 bits (88), Expect = 0.11
Identities = 28/91 (30%), Positives = 45/91 (48%), Gaps = 10/91 (10%)
Query: 125 RRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFA 184
++K IL+DVS ++ ++ +LG SGKT LL A++G+L +
Sbjct: 62 QQKWDRQILKDVSLYIESGQIMCILGSSGSGKTTLLDAISGRL----------RRTGTLE 111
Query: 185 GKVSYNGHEMNEFVPQRTAAYVSQNDTHLGN 215
G+V NG E+ Q +YV Q+D L +
Sbjct: 112 GEVFVNGCELRRDQFQDCFSYVLQSDVFLSS 142
>YME1_HUMAN (Q96TA2) ATP-dependent metalloprotease YME1L1 (EC
3.4.24.-) (YME1-like protein 1) (ATP-dependent
metalloprotease FtsH1) (Meg-4) (Presenilin-associated
metalloprotease) (PAMP) (UNQ1868/PRO4304)
Length = 773
Score = 38.1 bits (87), Expect = 0.15
Identities = 31/108 (28%), Positives = 48/108 (43%), Gaps = 17/108 (15%)
Query: 78 IDRAGVDIPTIEVRFEHLNVQAQVHVGKRALHTITNYMLDLVEYILKRRKQQLNILQDVS 137
+D A + V FEH+ V K+ L + ++ + Q+ IL
Sbjct: 324 LDSAVDPVQMKNVTFEHVK---GVEEAKQELQEVVEFL---------KNPQKFTILGG-- 369
Query: 138 GILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFAG 185
K + LL+GPP +GKT+L A+AG+ D A+ +F E F G
Sbjct: 370 ---KLPKGILLVGPPGTGKTLLARAVAGEADVPFYYASGSEFDEMFVG 414
>YME1_RAT (Q925S8) ATP-dependent metalloprotease YME1L1 (EC
3.4.24.-) (YME1-like protein 1) (ATP-dependent
metalloprotease FtsH1) (Meg-4)
Length = 715
Score = 37.7 bits (86), Expect = 0.19
Identities = 18/39 (46%), Positives = 25/39 (63%)
Query: 147 LLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFAG 185
LL+GPP +GKT+L A+AG+ D A+ +F E F G
Sbjct: 318 LLVGPPGTGKTLLARAVAGEADVPFYYASGSEFDEMFVG 356
>YME1_MOUSE (O88967) ATP-dependent metalloprotease YME1L1 (EC
3.4.24.-) (YME1-like protein 1) (ATP-dependent
metalloprotease FtsH1)
Length = 715
Score = 37.7 bits (86), Expect = 0.19
Identities = 18/39 (46%), Positives = 25/39 (63%)
Query: 147 LLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFAG 185
LL+GPP +GKT+L A+AG+ D A+ +F E F G
Sbjct: 318 LLVGPPGTGKTLLARAVAGEADVPFYYASGSEFDEMFVG 356
>BCS1_YEAST (P32839) Mitochondrial chaperone BCS1
Length = 456
Score = 37.7 bits (86), Expect = 0.19
Identities = 19/36 (52%), Positives = 24/36 (65%)
Query: 140 LKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIAN 175
+ + R LL GPP SGKT + ALAG+LD N+ I N
Sbjct: 257 IPYRRGYLLYGPPGSGKTSFIQALAGELDYNICILN 292
>PDRF_YEAST (Q04182) ATP-dependent permease PDR15
Length = 1529
Score = 37.4 bits (85), Expect = 0.25
Identities = 27/85 (31%), Positives = 38/85 (43%), Gaps = 12/85 (14%)
Query: 129 QLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFAGKVS 188
Q IL +V G +K LT L+G +GKT LL LA ++ + G +
Sbjct: 899 QRRILNNVDGWVKPGTLTALMGASGAGKTTLLDCLAERVTMGV-----------ITGNIF 947
Query: 189 YNGHEMNEFVPQRTAAYVSQNDTHL 213
+G +E P R+ Y Q D HL
Sbjct: 948 VDGRLRDESFP-RSIGYCQQQDLHL 971
Score = 32.3 bits (72), Expect = 8.1
Identities = 28/96 (29%), Positives = 45/96 (46%), Gaps = 13/96 (13%)
Query: 122 ILKRRKQQ--LNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQF 179
+LK K++ IL+ + G L L ++LG P SG T LL +++ KIA +
Sbjct: 173 LLKPSKEEDTFQILKPMDGCLNPGELLVVLGRPGSGCTTLLKSISSN-SHGFKIAKD--- 228
Query: 180 YEQFAGKVSYNGHEMNEFVP--QRTAAYVSQNDTHL 213
VSYNG ++ + Y +++D HL
Sbjct: 229 -----SIVSYNGLSSSDIRKHYRGEVVYNAESDIHL 259
>WHIT_LUCCU (Q05360) White protein
Length = 677
Score = 37.0 bits (84), Expect = 0.33
Identities = 28/99 (28%), Positives = 47/99 (47%), Gaps = 8/99 (8%)
Query: 127 KQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFAGK 186
K + +++++V G+ L ++G +GKT LL ALA + A VQ
Sbjct: 96 KPRKHLIKNVCGVAYPGELLAVMGSSGAGKTTLLNALA------FRSARGVQISPSSVRM 149
Query: 187 VSYNGHEMNEFVPQRTAAYVSQNDTHLGN*LSEKPWPFQ 225
+ NGH ++ Q AYV Q+D +G+ + + FQ
Sbjct: 150 L--NGHPVDAKEMQARCAYVQQDDLFIGSLTAREHLIFQ 186
>SNQ2_YEAST (P32568) SNQ2 protein
Length = 1501
Score = 37.0 bits (84), Expect = 0.33
Identities = 43/177 (24%), Positives = 77/177 (43%), Gaps = 28/177 (15%)
Query: 51 LLQRLLRNNTVEVDND-------HSFLLKLMRDRIDRAGVDIPTIEVRFEHLNVQAQVHV 103
L + R+ +D+D H+ +RD D G+ I V E ++ + V
Sbjct: 85 LSRHTTRSGAFNMDSDSDDGFDAHAIFESFVRDA-DEQGIHIRKAGVTIEDVSAKG---V 140
Query: 104 GKRALH--TITNYM---LDLVEYILKRRKQQLN-ILQDVSGILKHSRLTLLLGPPNSGKT 157
AL T N + L + + I +R Q++ I+ +V+ + + + L+LG P +G +
Sbjct: 141 DASALEGATFGNILCLPLTIFKGIKAKRHQKMRQIISNVNALAEAGEMILVLGRPGAGCS 200
Query: 158 ILLLALAGKLDPNLKIANEVQFYEQFAGKVSYNGHEMNEFVPQRTA--AYVSQNDTH 212
L AG++D QF +G+V+Y+G E + + A Y + D H
Sbjct: 201 SFLKVTAGEID---------QFAGGVSGEVAYDGIPQEEMMKRYKADVIYNGELDVH 248
Score = 33.1 bits (74), Expect = 4.7
Identities = 26/82 (31%), Positives = 38/82 (45%), Gaps = 13/82 (15%)
Query: 132 ILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFAGKVSYNG 191
+L +VSG +T L+G +GKT LL LA + N+ I G + NG
Sbjct: 871 LLDNVSGYCIPGTMTALMGESGAGKTTLLNTLAQR---NVGI---------ITGDMLVNG 918
Query: 192 HEMNEFVPQRTAAYVSQNDTHL 213
++ +RT YV Q D H+
Sbjct: 919 RPIDASFERRT-GYVQQQDIHI 939
>PDR5_YEAST (P33302) Suppressor of toxicity of sporidesmin
Length = 1511
Score = 37.0 bits (84), Expect = 0.33
Identities = 27/93 (29%), Positives = 42/93 (45%), Gaps = 12/93 (12%)
Query: 121 YILKRRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFY 180
Y ++ + + IL +V G +K LT L+G +GKT LL LA ++ +
Sbjct: 876 YEVQIKAETRRILNNVDGWVKPGTLTALMGASGAGKTTLLDCLAERVTMGV--------- 926
Query: 181 EQFAGKVSYNGHEMNEFVPQRTAAYVSQNDTHL 213
G + NG ++ P R+ Y Q D HL
Sbjct: 927 --ITGDILVNGIPRDKSFP-RSIGYCQQQDLHL 956
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.352 0.155 0.576
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 123,884,833
Number of Sequences: 164201
Number of extensions: 4652804
Number of successful extensions: 23120
Number of sequences better than 10.0: 137
Number of HSP's better than 10.0 without gapping: 55
Number of HSP's successfully gapped in prelim test: 82
Number of HSP's that attempted gapping in prelim test: 23008
Number of HSP's gapped (non-prelim): 173
length of query: 1226
length of database: 59,974,054
effective HSP length: 122
effective length of query: 1104
effective length of database: 39,941,532
effective search space: 44095451328
effective search space used: 44095451328
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 72 (32.3 bits)
Medicago: description of AC146819.12


