

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146818.6 + phase: 0 /pseudo
(642 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
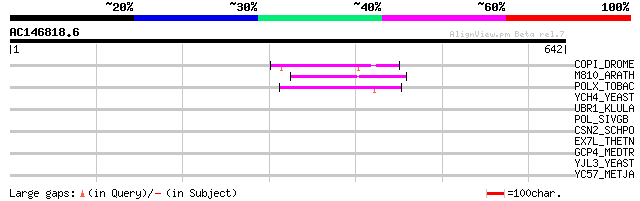
Score E
Sequences producing significant alignments: (bits) Value
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 78 6e-14
M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810... 74 9e-13
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 62 6e-09
YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein 44 0.001
UBR1_KLULA (O60014) N-end-recognizing protein (Ubiquitin-protein... 34 1.4
POL_SIVGB (P22382) Pol polyprotein [Contains: Protease (Retropep... 34 1.4
CSN2_SCHPO (Q9HFR0) COP9 signalosome complex subunit 2 (CSN comp... 34 1.4
EX7L_THETN (Q8RAC9) Probable exodeoxyribonuclease VII large subu... 33 2.3
GCP4_MEDTR (Q9SC88) Gamma-tubulin complex component 4 homolog 32 3.9
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 32 6.7
YC57_METJA (Q58654) Hypothetical protein MJ1257 31 8.8
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 78.2 bits (191), Expect = 6e-14
Identities = 44/158 (27%), Positives = 88/158 (54%), Gaps = 14/158 (8%)
Query: 302 DFKRGQVDTTLF---RGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMG 358
+F VD ++ +G + + I V+ +YVDD++ + + + F + + ++F + +
Sbjct: 1060 EFVNSSVDRCIYILDKGNINENIYVL-LYVDDVVIATGDMTRMNNFKRYLMEKFRMTDLN 1118
Query: 359 ELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPM----HPTCT*SKEDIG 414
E+K F+GI+I ++D +Y+ Q+ Y K++L KF +E+C ++TP+ + S ED
Sbjct: 1119 EIKHFIGIRIEMQEDKIYLSQSAYVKKILSKFNMENCNAVSTPLPSKINYELLNSDEDCN 1178
Query: 415 TKVDQKLYIGMIGSLLY-LTASRPDILFSVCLCARFQS 451
T +IG L+Y + +RPD+ +V + +R+ S
Sbjct: 1179 TPCR-----SLIGCLMYIMLCTRPDLTTAVNILSRYSS 1211
>M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810
(ORF240b)
Length = 240
Score = 74.3 bits (181), Expect = 9e-13
Identities = 41/134 (30%), Positives = 69/134 (50%), Gaps = 1/134 (0%)
Query: 326 IYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRKDGVYVHQTKYTKE 385
+YVDDI+ ++ +L + F +G + +FLGIQI G+++ QTKY ++
Sbjct: 5 LYVDDILLTGSSNTLLNMLIFQLSSTFSMKDLGPVHYFLGIQIKTHPSGLFLSQTKYAEQ 64
Query: 386 LLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGMIGSLLYLTASRPDILFSVCL 445
+L + DCK M+TP+ P S D + ++G+L YLT +RPDI ++V +
Sbjct: 65 ILNNAGMLDCKPMSTPL-PLKLNSSVSTAKYPDPSDFRSIVGALQYLTLTRPDISYAVNI 123
Query: 446 CARFQSNPKRISFN 459
+ P F+
Sbjct: 124 VCQRMHEPTLADFD 137
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 61.6 bits (148), Expect = 6e-09
Identities = 38/150 (25%), Positives = 79/150 (52%), Gaps = 9/150 (6%)
Query: 313 FRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRK 372
F+ + +++ +YVDD++ + L + + F+ +G + LG++I + +
Sbjct: 994 FKRFSENNFIILLLYVDDMLIVGKDKGLIAKLKGDLSKSFDMKDLGPAQQILGMKIVRER 1053
Query: 373 DG--VYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQK------LYIG 424
+++ Q KY + +L++F +++ K ++TP+ SK+ T V++K Y
Sbjct: 1054 TSRKLWLSQEKYIERVLERFNMKNAKPVSTPLAGHLKLSKKMCPTTVEEKGNMAKVPYSS 1113
Query: 425 MIGSLLY-LTASRPDILFSVCLCARFQSNP 453
+GSL+Y + +RPDI +V + +RF NP
Sbjct: 1114 AVGSLMYAMVCTRPDIAHAVGVVSRFLENP 1143
>YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein
Length = 308
Score = 44.3 bits (103), Expect = 0.001
Identities = 30/137 (21%), Positives = 63/137 (45%), Gaps = 2/137 (1%)
Query: 322 LVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRKDG-VYVHQT 380
+ + +YVDD++ + + + + + + +G++ FLG+ I+Q +G + +
Sbjct: 82 IYIGVYVDDLLVAAPSPKIYDRVKQELTKLYSMKDLGKVDKFLGLNIHQSTNGDITLSLQ 141
Query: 381 KYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGMIGSLLY-LTASRPDI 439
Y + + ++ K+ TP+ + + D Y ++G LL+ RPDI
Sbjct: 142 DYIAKAASESEINTFKLTQTPLCNSKPLFETTSPHLKDITPYQSIVGQLLFCANTGRPDI 201
Query: 440 LFSVCLCARFQSNPKRI 456
+ V L +RF P+ I
Sbjct: 202 SYPVSLLSRFLREPRAI 218
>UBR1_KLULA (O60014) N-end-recognizing protein (Ubiquitin-protein
ligase E3 component) (N-recognin)
Length = 1945
Score = 33.9 bits (76), Expect = 1.4
Identities = 22/70 (31%), Positives = 35/70 (49%), Gaps = 1/70 (1%)
Query: 335 STNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRKDGVYVHQTKYTKELLKKFKLED 394
S N + + S+ +D F + GE + F +Q + VY TK+T+E + KF ED
Sbjct: 1384 SNNICMLEMLSRLNKDPFGTLLSGEEQKFKTLQNILKSLAVYTRLTKHTEENVLKF-YED 1442
Query: 395 CKVMNTPMHP 404
+ N P +P
Sbjct: 1443 IRSSNLPSYP 1452
>POL_SIVGB (P22382) Pol polyprotein [Contains: Protease
(Retropepsin) (EC 3.4.23.-); Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)]
Length = 1009
Score = 33.9 bits (76), Expect = 1.4
Identities = 16/36 (44%), Positives = 23/36 (63%), Gaps = 1/36 (2%)
Query: 365 GIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNT 400
GI N + GV ++ KY KEL++K + EDCK + T
Sbjct: 864 GIPYNPQSQGVVENKNKYLKELIEKIR-EDCKELKT 898
>CSN2_SCHPO (Q9HFR0) COP9 signalosome complex subunit 2 (CSN complex
subunit 2) (SGN2)
Length = 437
Score = 33.9 bits (76), Expect = 1.4
Identities = 23/81 (28%), Positives = 38/81 (46%)
Query: 106 KTFQNLIRKLNLKKVQKHNPLQNLNEPQKHKMKMLLKKLTMTLSK*FNPKVHSSTSLHIL 165
K QNL + + KV H L + HK K LL+++ LS N + H+L
Sbjct: 138 KALQNLNNERLMLKVLMHVARFLLTQKNYHKFKYLLRQMHELLSDENNSVADQNRGTHLL 197
Query: 166 RI*SLETRIVLEEQDHTSNKK 186
+ SLE ++ + +D+ K+
Sbjct: 198 ELYSLEIQMYSDIEDNKRLKE 218
>EX7L_THETN (Q8RAC9) Probable exodeoxyribonuclease VII large subunit
(EC 3.1.11.6) (Exonuclease VII large subunit)
Length = 404
Score = 33.1 bits (74), Expect = 2.3
Identities = 25/68 (36%), Positives = 36/68 (52%), Gaps = 7/68 (10%)
Query: 88 LEVKLQSKMKVLQIYRSLKTFQNLIRKLNLKKVQKHNPLQNLNEPQKHKMKMLLKKLTMT 147
L+ K+ + MK Q+ S K F+ L R L L +NP++ NE K ++K L K LT
Sbjct: 274 LKTKIMNLMKA-QVLHSKKEFEGLKRALYL-----NNPIKK-NEVLKQRVKNLKKSLTKE 326
Query: 148 LSK*FNPK 155
+ FN K
Sbjct: 327 MLSIFNQK 334
>GCP4_MEDTR (Q9SC88) Gamma-tubulin complex component 4 homolog
Length = 739
Score = 32.3 bits (72), Expect = 3.9
Identities = 25/84 (29%), Positives = 40/84 (46%), Gaps = 7/84 (8%)
Query: 102 YRSLKTFQNLIRKLNLKKVQKHNPLQNLNEPQKHKMKMLLKKLTMTLSK*FNPKVHSSTS 161
YR L+ F R LN + + NPL+N +P ++ + L + L+ V+SS+
Sbjct: 64 YRELERFSAKSRNLNWIRSENANPLENKEKPSVYR-RALANGIVEILA------VYSSSI 116
Query: 162 LHILRI*SLETRIVLEEQDHTSNK 185
LHI ++ ET +L NK
Sbjct: 117 LHIEQLLLSETMPILATVTQGLNK 140
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 31.6 bits (70), Expect = 6.7
Identities = 35/150 (23%), Positives = 65/150 (43%), Gaps = 18/150 (12%)
Query: 322 LVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGEL------KFFLGIQI--NQRKD 373
L++ +YVDD + ++N EF ++ FE + G L LG+ + N+R
Sbjct: 1456 LMIAVYVDDCVIAASNEQRLDEFINKLKSNFELKITGTLIDDVLDTDILGMDLVYNKRLG 1515
Query: 374 GVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*S---KEDIGTKVDQKLYIG------ 424
+ + + + KK+ E K+ + + T K+D+ +++ G
Sbjct: 1516 TIDLTLKSFINRMDKKYNEELKKIRKSSIPHMSTYKIDPKKDVLQMSEEEFRQGVLKLQQ 1575
Query: 425 MIGSLLYLT-ASRPDILFSVCLCARFQSNP 453
++G L Y+ R DI F+V AR + P
Sbjct: 1576 LLGELNYVRHKCRYDIEFAVKKVARLVNYP 1605
>YC57_METJA (Q58654) Hypothetical protein MJ1257
Length = 349
Score = 31.2 bits (69), Expect = 8.8
Identities = 23/74 (31%), Positives = 38/74 (51%), Gaps = 8/74 (10%)
Query: 45 NLMPRPKEESF*VTLKGQRHTQCIIQRYYVLKNQCTLNLMTKSLEVKLQSKMKVLQIYRS 104
NL P +ES +TLK +H + Y++ NL TK + L+ +K+L YR+
Sbjct: 274 NLYP---DESLFLTLKFAKHLKT---NGYIIHTLKARNLKTKKED--LEKVLKILSYYRN 325
Query: 105 LKTFQNLIRKLNLK 118
+K F+ + + N K
Sbjct: 326 IKIFKIINLRANTK 339
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.352 0.155 0.517
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 61,599,241
Number of Sequences: 164201
Number of extensions: 2236678
Number of successful extensions: 7148
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 7139
Number of HSP's gapped (non-prelim): 13
length of query: 642
length of database: 59,974,054
effective HSP length: 117
effective length of query: 525
effective length of database: 40,762,537
effective search space: 21400331925
effective search space used: 21400331925
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (22.0 bits)
S2: 69 (31.2 bits)
Medicago: description of AC146818.6


