

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146792.8 - phase: 0
(266 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
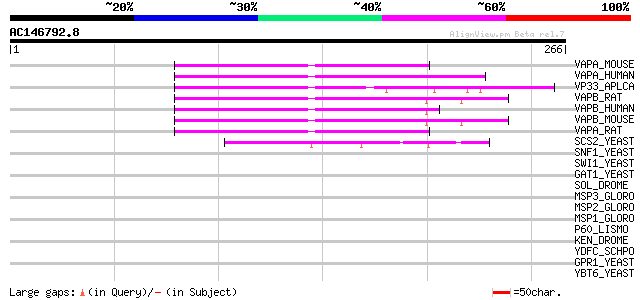
Score E
Sequences producing significant alignments: (bits) Value
VAPA_MOUSE (Q9WV55) Vesicle-associated membrane protein-associat... 55 2e-07
VAPA_HUMAN (Q9P0L0) Vesicle-associated membrane protein-associat... 54 3e-07
VP33_APLCA (Q16943) Vesicle-associated membrane protein/synaptob... 53 9e-07
VAPB_RAT (Q9Z269) Vesicle-associated membrane protein-associated... 53 9e-07
VAPB_HUMAN (O95292) Vesicle-associated membrane protein-associat... 52 1e-06
VAPB_MOUSE (Q9QY76) Vesicle-associated membrane protein-associat... 52 2e-06
VAPA_RAT (Q9Z270) Vesicle-associated membrane protein-associated... 52 2e-06
SCS2_YEAST (P40075) SCS2 protein 45 2e-04
SNF1_YEAST (P06782) Carbon catabolite derepressing protein kinas... 42 0.002
SWI1_YEAST (P09547) Transcription regulatory protein SWI1 (SWI/S... 41 0.003
GAT1_YEAST (P43574) Transcriptional regulatory protein GAT1 41 0.003
SOL_DROME (P27398) Small optic lobes protein (EC 3.4.22.-) 40 0.004
MSP3_GLORO (P53023) Major sperm protein 3 40 0.006
MSP2_GLORO (P53022) Major sperm protein 2 40 0.006
MSP1_GLORO (P53021) Major sperm protein 1 40 0.006
P60_LISMO (P21171) Protein p60 precursor (Invasion-associated pr... 39 0.010
KEN_DROME (O77459) Probable transcription factor Ken (Ken and Ba... 39 0.013
YDFC_SCHPO (Q10484) Hypothetical protein C17C9.12 in chromosome I 38 0.022
GPR1_YEAST (Q12361) G protein-coupled receptor GPR1 38 0.022
YBT6_YEAST (P38250) Hypothetical 105.9 kDa protein in AAC3-RFC5 ... 37 0.038
>VAPA_MOUSE (Q9WV55) Vesicle-associated membrane protein-associated
protein A (VAMP-associated protein A) (VAMP-A) (VAP-A)
(33 kDa Vamp-associated protein) (VAP-33)
Length = 242
Score = 54.7 bits (130), Expect = 2e-07
Identities = 37/122 (30%), Positives = 61/122 (49%), Gaps = 3/122 (2%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
L LDP + L F V + ++++N S V FK +TTAP+ +RP I+ PG +
Sbjct: 8 LVLDPPSDLKFKGPFTDVVTTNLKLQNPSDRKVCFKVKTTAPRRYCVRPNSGIIDPGSIV 67
Query: 140 IATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVFL 199
+V + QP + + EKS KF + ++ +I + ++ E K ++ LR VF
Sbjct: 68 TVSV---MLQPFDYDPNEKSKHKFMVQTIFAPPNISDMEAVWKEAKPDELMDSKLRCVFE 124
Query: 200 DP 201
P
Sbjct: 125 MP 126
>VAPA_HUMAN (Q9P0L0) Vesicle-associated membrane protein-associated
protein A (VAMP-associated protein A) (VAMP-A) (VAP-A)
(33 kDa Vamp-associated protein) (VAP-33)
Length = 242
Score = 54.3 bits (129), Expect = 3e-07
Identities = 38/149 (25%), Positives = 69/149 (45%), Gaps = 3/149 (2%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
L LDP L F V + ++++N S V FK +TTAP+ +RP I+ PG ++
Sbjct: 8 LVLDPPTDLKFKGPFTDVVTTNLKLRNPSDRKVCFKVKTTAPRRYCVRPNSGIIDPGSTV 67
Query: 140 IATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVFL 199
+V + QP + + EKS KF + ++ + + ++ E K ++ LR VF
Sbjct: 68 TVSV---MLQPFDYDPNEKSKHKFMVQTIFAPPNTSDMEAVWKEAKPDELMDSKLRCVFE 124
Query: 200 DPERPSPILEKLKRQLADADAALEERKKP 228
P + + + +A+ ++ P
Sbjct: 125 MPNENDKLNDMEPSKAVPLNASKQDGPMP 153
>VP33_APLCA (Q16943) Vesicle-associated membrane
protein/synaptobrevin binding protein (VAP-33)
Length = 260
Score = 52.8 bits (125), Expect = 9e-07
Identities = 53/194 (27%), Positives = 89/194 (45%), Gaps = 18/194 (9%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
L L+P+ +L F V + +++ N + + FK +TTAPK +RP IL P SI
Sbjct: 8 LILEPAGELRFKGPFTDVVTADLKLSNPTDRRICFKVKTTAPKRYCVRPNSGILEPKTSI 67
Query: 140 IATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPE----LFDEQKDQVAVEQILR 195
V + QP N + EK+ KF + S+ D+V E L+ + + ++ LR
Sbjct: 68 AVAV---MLQPFNYDPNEKNKHKFMVQSMYAP---DHVVESQELLWKDAPPESLMDTKLR 121
Query: 196 VVFLDPE--RPSPILEKLKRQLADA---DAALEE---RKKPAEDAGPKIIGEGLVIDEWK 247
VF P+ +P + + A A ++ALE+ + E PK +G E
Sbjct: 122 CVFEMPDGSHQAPASDASRATDAGAHFSESALEDPTVASRKTETQSPKRVGAVGSAGEDV 181
Query: 248 ERRERYLAKQQGDV 261
++ + L K Q ++
Sbjct: 182 KKLQHELKKAQSEI 195
>VAPB_RAT (Q9Z269) Vesicle-associated membrane protein-associated
protein B (VAMP-associated protein B) (VAMP-B) (VAP-B)
Length = 243
Score = 52.8 bits (125), Expect = 9e-07
Identities = 44/167 (26%), Positives = 78/167 (46%), Gaps = 10/167 (5%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
L L+P ++L F V + +++ N + NV FK +TTAP+ +RP ++ G S+
Sbjct: 8 LSLEPQHELKFRGPFTDVVTTNLKLGNPTDRNVCFKVKTTAPRRYCVRPNSGVIDAGASL 67
Query: 140 IATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVF- 198
+V + QP + + EKS KF + S+ + ++ E K + ++ LR VF
Sbjct: 68 NVSV---MLQPFDYDPNEKSKHKFMVQSMFAPPDTSDMEAVWKEAKPEDLMDSKLRCVFE 124
Query: 199 LDPERPSPILEKLKRQL------ADADAALEERKKPAEDAGPKIIGE 239
L E P ++ + + +A A + P +DA K + E
Sbjct: 125 LPAENAKPHDVEINKIMPTSASKTEAPVAAKPLTSPLDDAEVKKVME 171
>VAPB_HUMAN (O95292) Vesicle-associated membrane protein-associated
protein B/C (VAMP-associated protein B/C)
(VAMP-B/VAMP-C) (VAP-B/VAP-C) (UNQ484/PRO983)
Length = 243
Score = 52.4 bits (124), Expect = 1e-06
Identities = 39/128 (30%), Positives = 63/128 (48%), Gaps = 4/128 (3%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
L L+P ++L F V + +++ N + NV FK +TTAP+ +RP I+ G SI
Sbjct: 8 LSLEPQHELKFRGPFTDVVTTNLKLGNPTDRNVCFKVKTTAPRRYCVRPNSGIIDAGASI 67
Query: 140 IATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVF- 198
+V + QP + + EKS KF + S+ + ++ E K + ++ LR VF
Sbjct: 68 NVSV---MLQPFDYDPNEKSKHKFMVQSMFAPTDTSDMEAVWKEAKPEDLMDSKLRCVFE 124
Query: 199 LDPERPSP 206
L E P
Sbjct: 125 LPAENDKP 132
>VAPB_MOUSE (Q9QY76) Vesicle-associated membrane protein-associated
protein B (VAMP-associated protein B) (VAMP-associated
protein 33b) (VAMP-B) (VAP-B)
Length = 243
Score = 51.6 bits (122), Expect = 2e-06
Identities = 43/167 (25%), Positives = 77/167 (45%), Gaps = 10/167 (5%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
L L+P ++L F V + +++ N + NV FK +TT P+ +RP ++ G S+
Sbjct: 8 LSLEPQHELKFRGPFTDVVTTNLKLGNPTDRNVCFKVKTTVPRRYCVRPNSGVIDAGASL 67
Query: 140 IATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVF- 198
+V + QP + + EKS KF + S+ + ++ E K + ++ LR VF
Sbjct: 68 NVSV---MLQPFDYDPNEKSKHKFMVQSMFAPPDTSDMEAVWKEAKPEDLMDSKLRCVFE 124
Query: 199 LDPERPSPILEKLKRQL------ADADAALEERKKPAEDAGPKIIGE 239
L E P ++ + + +A AA + P +D K + E
Sbjct: 125 LPAENAKPHDVEINKIIPTSASKTEAPAAAKSLTSPLDDTEVKKVME 171
>VAPA_RAT (Q9Z270) Vesicle-associated membrane protein-associated
protein A (VAMP-associated protein A) (VAMP-A) (VAP-A)
(33 kDa Vamp-associated protein) (VAP-33)
Length = 242
Score = 51.6 bits (122), Expect = 2e-06
Identities = 35/122 (28%), Positives = 60/122 (48%), Gaps = 3/122 (2%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
L LDP + L F V + ++++N S V FK +TTAP+ +RP ++ PG +
Sbjct: 8 LVLDPPSDLKFKGPFTDVVTTNLKLQNPSDRKVCFKVKTTAPRRYCVRPNSGVIDPGSIV 67
Query: 140 IATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYVPELFDEQKDQVAVEQILRVVFL 199
+V + QP + + EKS KF + ++ +I + ++ E K ++ R VF
Sbjct: 68 TVSV---MLQPFDYDPNEKSKHKFMVQTIFAPPNISDMEAVWKEAKPDELMDSKPRCVFE 124
Query: 200 DP 201
P
Sbjct: 125 MP 126
>SCS2_YEAST (P40075) SCS2 protein
Length = 244
Score = 44.7 bits (104), Expect = 2e-04
Identities = 42/147 (28%), Positives = 67/147 (45%), Gaps = 23/147 (15%)
Query: 104 IKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESIIATV--FKFVEQPENNEKPEKSGL 161
I N S +AFK +TTAPK +RP A++APGE+I V E+P + K L
Sbjct: 27 ISNNSDQTIAFKVKTTAPKFYCVRPNAAVVAPGETIQVQVIFLGLTEEPAADFKCRDKFL 86
Query: 162 KFKIMS---LKVKGSIDYVPELFDEQKDQVAVEQILRVVFL---------------DPER 203
+ S L K D +L E K Q A+ + ++V +L + E
Sbjct: 87 VITLPSPYDLNGKAVADVWSDLEAEFKQQ-AISKKIKVKYLISPDVHPAQNQNIQENKET 145
Query: 204 PSPILEKLKRQLADADAALEERKKPAE 230
P+++ + + + A + E++ PAE
Sbjct: 146 VEPVVQDSEPK--EVPAVVNEKEVPAE 170
>SNF1_YEAST (P06782) Carbon catabolite derepressing protein kinase
(EC 2.7.1.-)
Length = 633
Score = 41.6 bits (96), Expect = 0.002
Identities = 18/46 (39%), Positives = 30/46 (65%)
Query: 25 SNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSVA 70
S+ ++TNT ++ N++ HHHHHH+ + + GS S +N SS+A
Sbjct: 2 SSNNNTNTAPANANSSHHHHHHHHHHHHHGHGGSNSTLNNPKSSLA 47
>SWI1_YEAST (P09547) Transcription regulatory protein SWI1 (SWI/SNF
complex component SWI1) (Transcription regulatory
protein ADR6) (Regulatory protein GAM3)
Length = 1314
Score = 41.2 bits (95), Expect = 0.003
Identities = 32/120 (26%), Positives = 57/120 (46%), Gaps = 5/120 (4%)
Query: 16 NFFKLPFRNSN---TSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSVARS 72
+FF L N+N T++T TT+++ NN + ++++NP +NT ST H+SN+ ++ +
Sbjct: 2 DFFNLNNNNNNNNTTTTTTTTNNNNTNNNNTNNNNNPANNTNNNNSTGHSSNTNNNTNNN 61
Query: 73 LLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAI 132
T D N +F +P Q + + S +N T A + F P +
Sbjct: 62 NTNTGASGVDDFQN--FFDPKPFDQNLDSNNNNSNSNNNDNNNSNTVASSTNFTSPTAVV 119
>GAT1_YEAST (P43574) Transcriptional regulatory protein GAT1
Length = 510
Score = 41.2 bits (95), Expect = 0.003
Identities = 31/113 (27%), Positives = 53/113 (46%), Gaps = 19/113 (16%)
Query: 20 LPFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSVARSLLPTRRR 79
LPF N+N S+NNN H+ HN NS + + ++T+ + S+ + P RR
Sbjct: 209 LPFLNNN---------SINNNHSHNSSHNNNSPSIANNTNANTNTNTSASTNTNSPLLRR 259
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTS----KSNVAFKFQTTAPKSCFMRP 128
+PS + +PG + S++R K + KS+ + + T P + P
Sbjct: 260 ---NPSPSI---VKPGSRRNSSVRKKKPALKKIKSSTSVQSSATPPSNTSSNP 306
>SOL_DROME (P27398) Small optic lobes protein (EC 3.4.22.-)
Length = 1594
Score = 40.4 bits (93), Expect = 0.004
Identities = 18/45 (40%), Positives = 31/45 (68%)
Query: 28 SSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSVARS 72
S +SSSV+++ +HHHHH+ NSN+ G+++ +N+ SS + S
Sbjct: 416 SEPMASSSSVSSSSNHHHHHHSNSNSNSSGNSNIINNNSSSSSGS 460
Score = 35.8 bits (81), Expect = 0.11
Identities = 16/43 (37%), Positives = 27/43 (62%)
Query: 31 NTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSVARSL 73
+++S S ++N HHHHH N NSN+ + + ++S SS + L
Sbjct: 421 SSSSVSSSSNHHHHHHSNSNSNSSGNSNIINNNSSSSSGSNKL 463
>MSP3_GLORO (P53023) Major sperm protein 3
Length = 125
Score = 40.0 bits (92), Expect = 0.006
Identities = 19/56 (33%), Positives = 27/56 (47%)
Query: 84 PSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
P+ K+ F + +RI N + F F+TT PK M PP +L P ES+
Sbjct: 12 PAQKVVFNAPFDNKATYYVRIINPGTKRIGFAFKTTKPKRINMNPPNGVLGPKESV 67
>MSP2_GLORO (P53022) Major sperm protein 2
Length = 125
Score = 40.0 bits (92), Expect = 0.006
Identities = 18/56 (32%), Positives = 28/56 (49%)
Query: 84 PSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
P+ K+ F + +R+ N + + F F+TT PK M PP +L P ES+
Sbjct: 12 PNQKVVFNAPFDNKATYYVRVINPGTNRIGFAFKTTKPKRINMNPPNGVLGPKESV 67
>MSP1_GLORO (P53021) Major sperm protein 1
Length = 125
Score = 40.0 bits (92), Expect = 0.006
Identities = 19/56 (33%), Positives = 27/56 (47%)
Query: 84 PSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
P+ K+ F + +RI N + F F+TT PK M PP +L P ES+
Sbjct: 12 PAQKVVFNAPFDNKATYYVRIINPGTKRIGFAFKTTKPKRINMNPPNGVLGPKESV 67
>P60_LISMO (P21171) Protein p60 precursor (Invasion-associated
protein)
Length = 484
Score = 39.3 bits (90), Expect = 0.010
Identities = 20/46 (43%), Positives = 32/46 (69%), Gaps = 1/46 (2%)
Query: 24 NSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPL-EGSTSHTSNSVSS 68
N+NT++TNT + S N N + + + N NSNT +GS+++ SNS +S
Sbjct: 325 NTNTNNTNTNTPSKNTNTNSNTNTNTNSNTNANQGSSNNNSNSSAS 370
Score = 30.8 bits (68), Expect = 3.6
Identities = 14/45 (31%), Positives = 27/45 (59%), Gaps = 1/45 (2%)
Query: 21 PFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNS 65
P N+N + TNT +++ N N ++ + + P+ NT +T+ +NS
Sbjct: 309 PSTNTNANKTNTNTNT-NTNTNNTNTNTPSKNTNTNSNTNTNTNS 352
>KEN_DROME (O77459) Probable transcription factor Ken (Ken and
Barbie protein)
Length = 601
Score = 38.9 bits (89), Expect = 0.013
Identities = 17/53 (32%), Positives = 31/53 (58%)
Query: 21 PFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSVARSL 73
P S ++S +TSS+ HHHHHH+ ++N + + +S++++ V SL
Sbjct: 229 PSGTSGSNSNISTSSNHQQQQHHHHHHHNHNNNNNNNNNNSSSSTINPVNLSL 281
Score = 37.0 bits (84), Expect = 0.050
Identities = 19/76 (25%), Positives = 37/76 (48%)
Query: 13 KVWNFFKLPFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSVARS 72
++ N P ++ S++N ++SS + HHHHH+ N N + +++S+S +
Sbjct: 220 EIVNLSNAPPSGTSGSNSNISTSSNHQQQQHHHHHHHNHNNNNNNNNNNSSSSTINPVNL 279
Query: 73 LLPTRRRLKLDPSNKL 88
L R + + S L
Sbjct: 280 SLDLRTKSENSASRTL 295
>YDFC_SCHPO (Q10484) Hypothetical protein C17C9.12 in chromosome I
Length = 319
Score = 38.1 bits (87), Expect = 0.022
Identities = 25/94 (26%), Positives = 47/94 (49%), Gaps = 5/94 (5%)
Query: 80 LKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPGAILAPGESI 139
+ L+ + + FP + V+ + ++NT+ + FK +TTAPK +RP G + ++
Sbjct: 1 MALECDSTIVFPRPLTRLVKCDLELRNTAPYPIGFKVKTTAPKQYCVRPNGGRIEANSAV 60
Query: 140 IATVFKFVEQPENNEKP--EKSGLKFKIMSLKVK 171
V + QP ++E K KF + S ++K
Sbjct: 61 SVEV---ILQPLDHEPAPGTKCRDKFLVQSTELK 91
>GPR1_YEAST (Q12361) G protein-coupled receptor GPR1
Length = 961
Score = 38.1 bits (87), Expect = 0.022
Identities = 27/106 (25%), Positives = 54/106 (50%), Gaps = 7/106 (6%)
Query: 24 NSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSVARSLLPTRRRLKLD 83
N+N + N ++S NNN ++++++N N+N + ++ +N+ S+ ++ + D
Sbjct: 502 NNNNDNDNDNNNSNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNSNNIKNNVDNNNTNPAD 561
Query: 84 P----SNKLYFPYEPGKQVR---SAIRIKNTSKSNVAFKFQTTAPK 122
SN+ + P + Q R +A R +N+S +NV FQ K
Sbjct: 562 NIPTLSNEAFTPSQQFSQERVNNNADRCENSSFTNVQQHFQAQTYK 607
>YBT6_YEAST (P38250) Hypothetical 105.9 kDa protein in AAC3-RFC5
intergenic region
Length = 946
Score = 37.4 bits (85), Expect = 0.038
Identities = 17/56 (30%), Positives = 30/56 (53%)
Query: 24 NSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSVARSLLPTRRR 79
N+NT++T TT ++ ++ HHHH H + ++ T +S SS A P ++
Sbjct: 881 NNNTTTTTTTDATQPHHHHHHHRHRDAGVKNVTNNSKTTESSSSSSAAKEKPKHKK 936
Score = 35.8 bits (81), Expect = 0.11
Identities = 19/66 (28%), Positives = 35/66 (52%), Gaps = 1/66 (1%)
Query: 23 RNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSVARSLLPTRRRLKL 82
R SN ++T TT++ HHHHHH + + ++ T+++ + SS + S + + K
Sbjct: 878 RPSNNNTTTTTTTDATQPHHHHHHHR-HRDAGVKNVTNNSKTTESSSSSSAAKEKPKHKK 936
Query: 83 DPSNKL 88
+KL
Sbjct: 937 GLLHKL 942
Score = 34.3 bits (77), Expect = 0.32
Identities = 15/51 (29%), Positives = 27/51 (52%)
Query: 21 PFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSVAR 71
P N+ T++T T ++ +++ HHH H + S + S+S SS A+
Sbjct: 879 PSNNNTTTTTTTDATQPHHHHHHHRHRDAGVKNVTNNSKTTESSSSSSAAK 929
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.130 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,144,587
Number of Sequences: 164201
Number of extensions: 1359420
Number of successful extensions: 8165
Number of sequences better than 10.0: 186
Number of HSP's better than 10.0 without gapping: 109
Number of HSP's successfully gapped in prelim test: 80
Number of HSP's that attempted gapping in prelim test: 6618
Number of HSP's gapped (non-prelim): 763
length of query: 266
length of database: 59,974,054
effective HSP length: 108
effective length of query: 158
effective length of database: 42,240,346
effective search space: 6673974668
effective search space used: 6673974668
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 65 (29.6 bits)
Medicago: description of AC146792.8


